Recognition: 1 theorem link
· Lean TheoremAutomated Detection of Abnormalities in Zebrafish Development
Pith reviewed 2026-05-12 05:06 UTC · model grok-4.3
The pith
A new zebrafish embryo image dataset and spatiotemporal transformer model automate detection of developmental abnormalities at 98% fertility and 92% toxicity accuracy.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
The paper establishes a large-scale dataset of zebrafish embryonic development image sequences under control and 3,4-dichloroaniline exposure conditions, annotated at fine-grained temporal levels for two tasks: fertility classification on 130,368 images and toxicity assessment on 55,296 images. It further presents a spatiotemporal transformer baseline model that achieves 98% accuracy in fertility classification and 92% accuracy in toxicity assessment, demonstrating the feasibility of automated early-stage prediction of developmental abnormalities.
What carries the argument
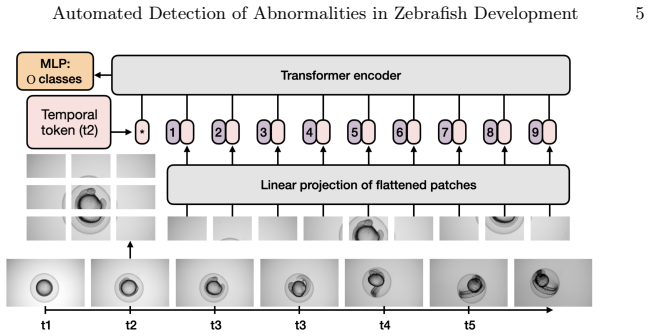
The spatiotemporal transformer baseline model that integrates spatial and temporal features from high-resolution microscopic image sequences to classify fertility and detect toxicity-induced malformations.
If this is right
- Manual inspection can be replaced by automated systems to increase throughput in zebrafish-based drug discovery.
- Developmental abnormalities can be flagged at early stages using video sequences rather than later visual checks.
- The released dataset serves as a benchmark for other models to improve toxicity assessment over time.
- Fine-grained temporal annotations allow precise tracking of when malformations appear after exposure.
Where Pith is reading between the lines
- If the dataset expands to more compounds it could become a standard test set for computer vision methods in developmental biology.
- Similar video-based approaches might transfer to other transparent embryos such as those of fruit flies or nematodes.
- Real-time lab monitoring hardware could incorporate the model to reduce the need for constant human oversight.
Load-bearing premise
The reported accuracies will hold on new compounds, new imaging setups, and new fish strains.
What would settle it
Testing the model on image sequences from a different compound or zebrafish strain and measuring whether accuracy falls well below the reported 92 percent.
Figures



read the original abstract
Zebrafish embryos are a valuable model for drug discovery due to their optical transparency and genetic similarity to humans. However, current evaluations rely on manual inspection, which is costly and labor-intensive. While machine learning offers automation potential, progress is limited by the lack of comprehensive datasets. To address this, we introduce a large-scale dataset of high-resolution microscopic image sequences capturing zebrafish embryonic development under both control conditions and exposure to compounds (3,4-dichloroaniline). This dataset, with expert annotations at fine-grained temporal levels, supports two benchmarking tasks: (1) fertility classification, assessing zebrafish egg viability (130,368 images), and (2) toxicity assessment, detecting malformations induced by toxic exposure over time (55,296 images). Alongside the dataset, we present the first transformer-based baseline model that integrates spatiotemporal features to predict developmental abnormalities at early stages. Experimental results present the model's effectiveness, achieving 98% accuracy in fertility classification and 92% in toxicity assessment. These findings underscore the potential of automated approaches to enhance zebrafish-based toxicity analysis.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper introduces a new large-scale dataset of high-resolution microscopic image sequences of zebrafish embryonic development under control conditions and exposure to 3,4-dichloroaniline, with expert annotations supporting two benchmarking tasks: fertility classification (130,368 images) and toxicity assessment (55,296 images). It also presents a transformer-based baseline model that integrates spatiotemporal features and reports 98% accuracy on fertility classification and 92% accuracy on toxicity assessment.
Significance. If the accuracies are shown to be robust, the dataset would be a significant contribution as a benchmark resource for automated analysis in zebrafish-based drug discovery and toxicity screening, addressing the current reliance on manual inspection. The spatiotemporal transformer baseline would demonstrate the applicability of modern sequence models to early developmental abnormality detection.
major comments (2)
- [Abstract] Abstract: The headline accuracies (98% fertility, 92% toxicity) are reported without any description of train/test split strategy, cross-validation (e.g., embryo-ID stratified k-fold), handling of temporal sequences, class imbalance, or statistical significance. This directly affects the ability to judge whether the numbers support the central claim that the model predicts abnormalities at early stages rather than memorizing batch-specific or compound-specific patterns.
- [Experimental results] Toxicity assessment results: The 92% accuracy is obtained exclusively on embryos exposed to a single compound (3,4-dichloroaniline). Without evidence of generalization testing across additional compounds, concentrations, or imaging sessions, the result does not yet establish the model's effectiveness for the broader drug-discovery use cases claimed.
minor comments (1)
- [Abstract] The sentence 'Experimental results present the model's effectiveness' in the abstract is grammatically awkward and should be revised for clarity (e.g., 'Experimental results demonstrate the model's effectiveness').
Simulated Author's Rebuttal
We thank the referee for the constructive comments on our manuscript. We provide point-by-point responses below and indicate the revisions we will incorporate.
read point-by-point responses
-
Referee: [Abstract] Abstract: The headline accuracies (98% fertility, 92% toxicity) are reported without any description of train/test split strategy, cross-validation (e.g., embryo-ID stratified k-fold), handling of temporal sequences, class imbalance, or statistical significance. This directly affects the ability to judge whether the numbers support the central claim that the model predicts abnormalities at early stages rather than memorizing batch-specific or compound-specific patterns.
Authors: We agree that the abstract omits key evaluation details. The manuscript body specifies an embryo-ID stratified 5-fold cross-validation to avoid leakage across temporal sequences of the same embryo, a spatiotemporal transformer architecture that processes image sequences, weighted loss to address class imbalance, and mean accuracy with standard deviation across runs. We will revise the abstract to concisely note the stratified cross-validation and temporal sequence handling. This will clarify that the reported accuracies reflect generalization across embryos rather than batch-specific memorization. revision: yes
-
Referee: [Experimental results] Toxicity assessment results: The 92% accuracy is obtained exclusively on embryos exposed to a single compound (3,4-dichloroaniline). Without evidence of generalization testing across additional compounds, concentrations, or imaging sessions, the result does not yet establish the model's effectiveness for the broader drug-discovery use cases claimed.
Authors: We acknowledge the limitation: the toxicity task uses only 3,4-dichloroaniline as the exposure condition, consistent with the dataset construction. No additional compounds or concentrations are present in the current data release, so we cannot provide empirical generalization results. We will revise the text to explicitly state the single-compound scope, frame the 92% accuracy as a benchmark for this established toxicant, and note broader applicability as future work. The baseline still demonstrates the spatiotemporal transformer's utility for early malformation detection in this setting. revision: partial
- The dataset contains only control and 3,4-dichloroaniline conditions, so we cannot supply evidence of generalization to other compounds or concentrations.
Circularity Check
No circularity: empirical accuracies on newly collected dataset
full rationale
The paper introduces a new image-sequence dataset of zebrafish embryos (control and 3,4-dichloroaniline-exposed) together with expert annotations and then trains a spatiotemporal transformer baseline, reporting direct empirical accuracies (98% fertility, 92% toxicity) on that data. These numbers are computed from model outputs versus held-out labels; they are not obtained by fitting a parameter to a subset and renaming the fit as a prediction, nor by any self-definitional equation, nor by a load-bearing self-citation chain. No derivation step reduces to its own inputs by construction. The central claims therefore remain independent empirical measurements.
Axiom & Free-Parameter Ledger
axioms (2)
- domain assumption Expert annotations at fine-grained temporal levels constitute reliable ground truth for both tasks.
- domain assumption The collected image sequences are representative of zebrafish development under control and toxic conditions.
Lean theorems connected to this paper
-
IndisputableMonolith/Foundation/AbsoluteFloorClosure.leanreality_from_one_distinction unclearWe employ a Vision Transformer (ViT) architecture adapted for spatiotemporal processing... patch embeddings... temporal positional embedding... 12 transformer blocks... MLP classification head
Reference graph
Works this paper leans on
-
[1]
Alshut, R., Legradi, J., Liebel, U., Yang, L., van Wezel, J., Strähle, U., Mikut, R., Reischl, M.: Methods for automated high-throughput toxicity testing using zebrafish embryos. In: KI 2010: Advances in Artificial Intelligence: 33rd Annual German Conference on AI, Karlsruhe, Germany, September 21-24, 2010. Proceed- ings 33. pp. 219–226. Springer (2010)
work page 2010
-
[2]
Cachat, J., Stewart, A., Utterback, E., Hart, P., Gaikwad, S., Wong, K., Kyzar, E., Wu, N., Kalueff, A.V.: Three-dimensional neurophenotyping of adult zebrafish behavior. PloS one6(3), e17597 (2011)
work page 2011
-
[3]
The Journal of physiology589(15), 3703–3708 (2011)
Cario, C.L., Farrell, T.C., Milanese, C., Burton, E.A.: Automated measurement of zebrafish larval movement. The Journal of physiology589(15), 3703–3708 (2011)
work page 2011
-
[4]
In: 2015 IEEE 12th international symposium on biomedical imaging (ISBI)
Dong, B., Shao, L., Da Costa, M., Bandmann, O., Frangi, A.F.: Deep learning for automatic cell detection in wide-field microscopy zebrafish images. In: 2015 IEEE 12th international symposium on biomedical imaging (ISBI). pp. 772–776. IEEE (2015)
work page 2015
-
[5]
An Image is Worth 16x16 Words: Transformers for Image Recognition at Scale
Dosovitskiy, A.: An image is worth 16x16 words: Transformers for image recogni- tion at scale. arXiv preprint arXiv:2010.11929 (2020)
work page internal anchor Pith review Pith/arXiv arXiv 2010
-
[6]
Toxicological Sciences163(1), 5–12 (2018)
Horzmann, K.A., Freeman, J.L.: Making waves: New developments in toxicology with the zebrafish. Toxicological Sciences163(1), 5–12 (2018)
work page 2018
-
[7]
IEEE transactions on medical imaging (2018)
Iakovidis, D.K., Georgakopoulos, S.V., Vasilakakis, M., Koulaouzidis, A., Pla- gianakos, V.P.: Detecting and locating gastrointestinal anomalies using deep learn- ing and iterative cluster unification. IEEE transactions on medical imaging (2018)
work page 2018
-
[8]
In: International Conference on Data Science and Artificial Intelli- gence
Javanmardi, S., Tang, X., Jahanbanifard, M., Verbeek, F.J.: Unsupervised seg- mentation of high-throughput zebrafish images using deep neural networks and transformers. In: International Conference on Data Science and Artificial Intelli- gence. pp. 213–227. Springer (2023) 10 S. Sivaprasad et al
work page 2023
-
[9]
In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2014)
Karpathy,A.,Toderici,G.,Shetty,S.,Leung,T.,Sukthankar,R.,Fei-Fei,L.:Large- scale video classification with convolutional neural networks. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2014)
work page 2014
-
[10]
Nature chemical biology6(3), 231–237 (2010)
Kokel, D., Bryan, J., Laggner, C., White, R., Cheung, C.Y.J., Mateus, R., Healey, D., Kim, S., Werdich, A.A., Haggarty, S.J., et al.: Rapid behavior-based identifica- tion of neuroactive small molecules in the zebrafish. Nature chemical biology6(3), 231–237 (2010)
work page 2010
-
[11]
Nature Reviews Genetics8(5), 353–367 (2007)
Lieschke, G.J., Currie, P.D.: Animal models of human disease: zebrafish swim into view. Nature Reviews Genetics8(5), 353–367 (2007)
work page 2007
-
[12]
Nature reviews Drug discovery14(10), 721–731 (2015)
MacRae, C.A., Peterson, R.T.: Zebrafish as tools for drug discovery. Nature reviews Drug discovery14(10), 721–731 (2015)
work page 2015
-
[13]
Zebrafish10(3), 401–421 (2013)
Mikut, R., Dickmeis, T., Driever, W., Geurts, P., Hamprecht, F.A., Kausler, B.X., Ledesma-Carbayo, M.J., Marée, R., Mikula, K., Pantazis, P., et al.: Automated processing of zebrafish imaging data: a survey. Zebrafish10(3), 401–421 (2013)
work page 2013
-
[14]
In: The eleventh international conference on learning representations (2022)
Mirzaei, H., Salehi, M., Shahabi, S., Gavves, E., Snoek, C.G., Sabokrou, M., Ro- hban, M.H.: Fake it until you make it: Towards accurate near-distribution novelty detection. In: The eleventh international conference on learning representations (2022)
work page 2022
-
[15]
In: 2011 IEEE International Sym- posium on Biomedical Imaging: From Nano to Macro
Ohn, J., Liebling, M.: In vivo, high-throughput imaging for functional charac- terization of the embryonic zebrafish heart. In: 2011 IEEE International Sym- posium on Biomedical Imaging: From Nano to Macro. pp. 1549–1552 (2011). https://doi.org/10.1109/ISBI.2011.5872696
-
[16]
Reproductive toxicology33(2), 174–187 (2012)
Padilla, S., Corum, D., Padnos, B., Hunter, D., Beam, A., Houck, K., Sipes, N., Kleinstreuer, N., Knudsen, T., Dix, D., et al.: Zebrafish developmental screening of the toxcast™phase i chemical library. Reproductive toxicology33(2), 174–187 (2012)
work page 2012
-
[17]
Current opinion in chemical biology24, 58–70 (2015)
Rennekamp, A.J., Peterson, R.T.: 15 years of zebrafish chemical screening. Current opinion in chemical biology24, 58–70 (2015)
work page 2015
-
[18]
Science327(5963), 348– 351 (2010)
Rihel,J.,Prober,D.A.,Arvanites,A.,Lam,K.,Zimmerman,S.,Jang,S.,Haggarty, S.J., Kokel, D., Rubin, L.L., Peterson, R.T., et al.: Zebrafish behavioral profiling links drugs to biological targets and rest/wake regulation. Science327(5963), 348– 351 (2010)
work page 2010
-
[19]
Environmental Science & Technology58(50), 21942–21953 (2024)
Schunck, F., Kodritsch, B., Krauss, M., Busch, W., Focks, A.: Integrating time- resolved nrf2 gene-expression data into a full guts model as a proxy for toxicody- namic damage in zebrafish embryo. Environmental Science & Technology58(50), 21942–21953 (2024)
work page 2024
-
[20]
Journal of laboratory automation17(6), 435–442 (2012)
Spomer, W., Pfriem, A., Alshut, R., Just, S., Pylatiuk, C.: High-throughput screen- ing of zebrafish embryos using automated heart detection and imaging. Journal of laboratory automation17(6), 435–442 (2012)
work page 2012
-
[21]
Toxicological sciences 137(1), 212–233 (2014)
Truong, L., Reif, D.M., St Mary, L., Geier, M.C., Truong, H.D., Tanguay, R.L.: Multidimensional in vivo hazard assessment using zebrafish. Toxicological sciences 137(1), 212–233 (2014)
work page 2014
-
[22]
Tyagi, G., Patel, N., Sethi, I.: A fine-tuned convolution neural network based ap- proachforphenotypeclassificationofzebrafishembryo.ProcediaComputerScience 126, 1138–1144 (2018)
work page 2018
-
[23]
Vaswani, A., Shazeer, N., Parmar, N., Uszkoreit, J., Jones, L., Gomez, A.N., Kaiser, L., Polosukhin, I.: Attention is all you need.(nips), 2017. arXiv preprint arXiv:1706.0376210, S0140525X16001837 (2017)
work page internal anchor Pith review Pith/arXiv arXiv 2017
-
[24]
Xu, L., Wang, S.: Technical report: Masked skeleton sequence modeling for learn- ing larval zebrafish behavior latent embeddings. arXiv preprint arXiv:2403.15693 (2024) Automated Detection of Abnormalities in Zebrafish Development 11
-
[25]
Sci- entific reports7(1), 42815 (2017)
Xu, Z., Cheng, X.E.: Zebrafish tracking using convolutional neural networks. Sci- entific reports7(1), 42815 (2017)
work page 2017
-
[26]
PloS one12(1), e0169408 (2017)
Zhang, G., Truong, L., Tanguay, R.L., Reif, D.M.: A new statistical approach to characterize chemical-elicited behavioral effects in high-throughput studies using zebrafish. PloS one12(1), e0169408 (2017)
work page 2017
-
[27]
In: Proceedings of the IEEE/CVF Confer- ence on computer vision and pattern recognition (2021)
Zhao, Y., Wu, W., He, Y., Li, Y., Tan, X., Chen, S.: Good practices and a strong baseline for traffic anomaly detection. In: Proceedings of the IEEE/CVF Confer- ence on computer vision and pattern recognition (2021)
work page 2021
-
[28]
Nature reviews Drug discovery4(1), 35–44 (2005)
Zon, L.I., Peterson, R.T.: In vivo drug discovery in the zebrafish. Nature reviews Drug discovery4(1), 35–44 (2005)
work page 2005
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.