Recognition: no theorem link
Beyond Manual Curation: Augmenting Targeted Protein Degradation Databases via Agentic Literature Extraction Workflows
Pith reviewed 2026-05-13 00:56 UTC · model grok-4.3
The pith
An LLM workflow refines prompts from seven annotated papers to extract structured TPD assay records and expand existing databases by 81 to 92 percent with high expert accuracy.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
With seven expert-annotated molecular glue publications the workflow reaches record-level F1 of 0.98 on extraction; the same instructions transfer to PROTAC papers by terminology substitution alone and maintain F1 above 0.93. When run at scale the method adds 81 percent more molecular glue records and 92 percent more PROTAC records, of which expert review confirms 92 percent and 82.5 percent as correct. The extracted records also capture kinetic parameters and assay context that were previously missing from the databases.
What carries the argument
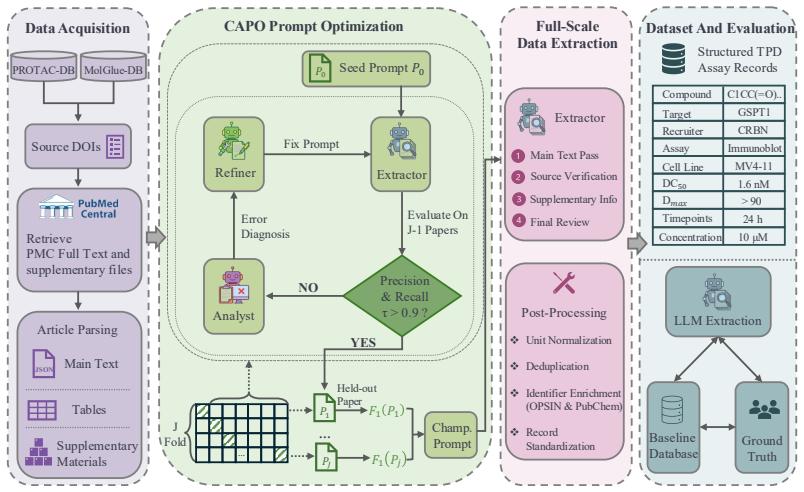
Expert-in-the-loop LLM workflow that uses a lightweight cross-validated prompt-refinement module to adapt extraction instructions from scarce annotations and applies terminology substitution to transfer performance between related compound classes.
Load-bearing premise
A handful of expert-annotated publications is enough to produce prompts that remain accurate across the much larger and more varied body of TPD literature.
What would settle it
Expert review of a new random sample of 200 extracted records drawn from papers outside the original seven would show precision or recall falling below 0.80.
Figures





read the original abstract
Predictive models in biomedicine depend on structured assay data locked in the text, tables, and supplements of primary publications. This bottleneck is especially acute in targeted protein degradation (TPD), where each assay record must combine compound identity, degradation target, recruiter, assay context, and endpoint values reported across sections, tables, and supplementary files. Inconsistent compound identifiers and incomplete or implicit assay context further demand domain-specific logic that generic LLM pipelines do not provide. Existing molecular glue and PROTAC databases are manually curated and often lack the experimental context required for downstream modeling. We formulate TPD database extraction as a domain-specific curation task and present an expert-in-the-loop LLM workflow, evaluated through a triangular comparison among LLM predictions, standardized baseline records, and expert-annotated ground truth. A lightweight cross-validated prompt-refinement module adapts extraction instructions from scarce expert annotations. With only seven annotated molecular glue publications, the workflow achieved record-level $F_1 = 0.98$ and transferred to PROTACs by terminology substitution alone, maintaining record-level $F_1 > 0.93$. Applied at scale, it expanded molecular glue and PROTAC databases by 81% and 92% records, respectively, with 92% and 82.5% of newly recovered records validated as correct upon expert review. The workflow also recovered kinetic and assay-context information essential for cross-study potency comparison and condition-aware degradation modeling. We release the workflow, prompts, evaluation code, and extracted datasets as resources for TPD data curation and AI-assisted scientific curation more broadly.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper claims to develop an expert-in-the-loop LLM workflow for extracting structured assay data (compound identity, target, recruiter, assay context, endpoints) from TPD literature to augment molecular glue and PROTAC databases. A cross-validated prompt-refinement module is applied to only seven annotated molecular glue publications, yielding record-level F1=0.98; the workflow transfers to PROTACs by terminology substitution alone (F1>0.93). Applied at scale, it expands the databases by 81% and 92% records, with 92% and 82.5% expert validation of new records. It also recovers kinetic/assay-context data for modeling, and releases the workflow, prompts, evaluation code, and extracted datasets.
Significance. If the generalization claims hold, the work is significant for addressing the structured-data bottleneck in biomedicine, especially TPD where assay records are essential for predictive modeling. Strengths include high reported accuracy with minimal annotations, successful transfer without retraining, substantial database expansion, and expert validation of new records. The release of workflow, prompts, code, and datasets supports reproducibility and broader use in AI-assisted curation.
major comments (3)
- [Prompt Refinement section] Prompt Refinement section: The cross-validated refinement uses only seven molecular glue publications. The manuscript provides no details on selection criteria, diversity (e.g., journals, years, table formats, or supplementary usage), or how the triangular evaluation ensures independence from this narrow set. This is load-bearing for the central claim that F1=0.98 and 81%/92% expansions reflect domain-general performance rather than overfitting to the annotation sample.
- [Transfer to PROTACs subsection] Transfer to PROTACs subsection: Terminology substitution alone is used for transfer, with no ablation or control comparing performance when prompts are refined directly on PROTAC papers. Given potential differences in degradation kinetics reporting, this weakens the claim that F1>0.93 and 92% expansion are robust without domain-specific adjustments.
- [Database Expansion and Validation results] Database Expansion and Validation results: While 92%/82.5% expert validation is reported for new records, the details of how the triangular comparison (LLM predictions vs. baseline vs. expert ground truth) avoids circularity with the prompt-refinement set are insufficient to fully support the scale-up claims.
minor comments (2)
- [Abstract] Abstract: The term 'triangular comparison' is used without a one-sentence definition or pointer to the methods; adding this would improve accessibility.
- [Methods] Notation: Record-level F1 is reported consistently in results but could be explicitly defined once in methods to avoid any reader ambiguity when comparing glue and PROTAC performance.
Simulated Author's Rebuttal
We thank the referee for their constructive review and positive assessment of the work's significance for addressing the structured-data bottleneck in TPD. We address each major comment point by point below, providing clarifications and committing to revisions that strengthen the manuscript without misrepresenting our methods or results.
read point-by-point responses
-
Referee: [Prompt Refinement section] Prompt Refinement section: The cross-validated refinement uses only seven molecular glue publications. The manuscript provides no details on selection criteria, diversity (e.g., journals, years, table formats, or supplementary usage), or how the triangular evaluation ensures independence from this narrow set. This is load-bearing for the central claim that F1=0.98 and 81%/92% expansions reflect domain-general performance rather than overfitting to the annotation sample.
Authors: We agree that explicit details on the seven publications are needed to support claims of generalizability. In the revised manuscript, we will add a dedicated paragraph in the Prompt Refinement section describing the selection criteria: the publications were chosen to span 2018-2023, multiple journals (including Nature Chemical Biology, Cell Chemical Biology, and ACS Chemical Biology), and varied reporting formats (main-text tables, supplementary tables, and inline text). The triangular evaluation used independent expert annotations performed by two TPD specialists who did not participate in prompt engineering, with cross-validation folds ensuring each publication was held out during testing. We will clarify this independence to demonstrate that the F1=0.98 reflects performance beyond the annotation sample. revision: yes
-
Referee: [Transfer to PROTACs subsection] Transfer to PROTACs subsection: Terminology substitution alone is used for transfer, with no ablation or control comparing performance when prompts are refined directly on PROTAC papers. Given potential differences in degradation kinetics reporting, this weakens the claim that F1>0.93 and 92% expansion are robust without domain-specific adjustments.
Authors: The terminology-substitution approach was chosen to emphasize minimal adaptation, and the resulting F1>0.93 plus expert validation of the expanded set provide supporting evidence. However, we acknowledge that an ablation would strengthen the robustness claim. In the revision, we will add a control analysis refining prompts directly on a small set of PROTAC papers (using the same cross-validated procedure) and report the comparative F1 scores. We will also add discussion of differences in kinetics reporting between molecular glues and PROTACs to address this concern directly. revision: yes
-
Referee: [Database Expansion and Validation results] Database Expansion and Validation results: While 92%/82.5% expert validation is reported for new records, the details of how the triangular comparison (LLM predictions vs. baseline vs. expert ground truth) avoids circularity with the prompt-refinement set are insufficient to fully support the scale-up claims.
Authors: We will expand the Database Expansion and Validation section to detail the separation of sets. The expert ground truth for validating newly extracted records was generated from a random sample of publications explicitly excluded from the original seven used for prompt refinement. Baseline records are drawn from pre-existing manually curated databases, and LLM predictions apply the refined prompts to the full literature corpus. We will include a supplementary table listing the publication IDs or DOIs used in refinement versus validation to make the lack of overlap explicit and eliminate any possibility of circularity. revision: yes
Circularity Check
No circularity: performance claims rest on independent expert ground truth and cross-validation
full rationale
The paper's central results derive from a triangular evaluation comparing LLM extractions against expert-annotated ground truth and standardized baseline records, with cross-validated prompt refinement on the seven molecular glue publications and separate expert validation of newly extracted records at scale. No step reduces by construction to the workflow's own outputs, fitted parameters, or self-citations; the reported F1 scores and database expansions are measured against external human annotations rather than being tautological or self-referential. The terminology-substitution transfer to PROTACs is presented as an empirical observation validated by the same independent review process, with no uniqueness theorems, ansatzes, or renamings of prior results invoked to support the claims.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Large language models possess sufficient reasoning and extraction capabilities that can be elicited and refined through a small set of expert-annotated examples using prompt engineering.
Reference graph
Works this paper leans on
-
[1]
The landscape of biomedical research.Patterns, 5(6), 2024
Rita González-Márquez, Luca Schmidt, Benjamin M Schmidt, Philipp Berens, and Dmitry Kobak. The landscape of biomedical research.Patterns, 5(6), 2024
work page 2024
-
[2]
Andreas Bender and Isidro Cortes-Ciriano. Artificial intelligence in drug discovery: what is realistic, what are illusions? part 2: a discussion of chemical and biological data.Drug Discovery Today, 26(4):1040–1052, 2021
work page 2021
-
[3]
Guangcai Zhong, Xiaoyu Chang, Weilin Xie, and Xiangxiang Zhou. Targeted protein degradation: advances in drug discovery and clinical practice.Signal transduction and targeted therapy, 9(1):308, 2024
work page 2024
-
[4]
Yossra Gharbi and Rocío Mercado. A comprehensive review of emerging approaches in machine learning for de novo PROTAC design.Digital Discovery, 3(11):2158–2176, 2024
work page 2024
-
[5]
Jingxuan Ge, Shimeng Li, Gaoqi Weng, Huating Wang, Meijing Fang, Huiyong Sun, Yafeng Deng, Chang- Yu Hsieh, Dan Li, and Tingjun Hou. PROTAC-DB 3.0: an updated database of PROTACs with extended pharmacokinetic parameters.Nucleic Acids Research, 53(D1):D1510–D1515, 2025
work page 2025
-
[6]
MolGlueDB: an online database of molecular glues.Nucleic Acids Research, 54(D1):D1510–D1518, 2026
Xiao Wang, Zhiyao Zhuang, Chengwei Zhang, Bowen Zhang, Wei Zhan, Yifan Wang, Zhaojuan Liu, Shanwen Yuan, Wenjia Niu, Qi He, et al. MolGlueDB: an online database of molecular glues.Nucleic Acids Research, 54(D1):D1510–D1518, 2026. 10
work page 2026
-
[7]
Xinran Qin, Yinpeng Zhang, Yajunzi Wang, Yintao Zhang, Jiachen Jing, Yuyuan Zhang, Gaoxiang Xu, Haoping Teng, Tianjun Wang, Lei Fu, et al. TPDdb: the comprehensive database of targeted protein degrader.Nucleic Acids Research, 54(D1):D1683–D1691, 2026
work page 2026
-
[8]
PROTAC-PatentDB: A PROTAC patent compound dataset.Scientific Data, 12(1):1840, 2025
Hong Cai, Gengyuan Yao, Yulong Shi, Tianyi Zhang, and Yuanjia Hu. PROTAC-PatentDB: A PROTAC patent compound dataset.Scientific Data, 12(1):1840, 2025
work page 2025
-
[9]
Stefano Ribes, Eva Nittinger, Christian Tyrchan, and Rocío Mercado. Modeling PROTAC degradation activity with machine learning.Artificial Intelligence in the Life Sciences, 6:100114, 2024
work page 2024
-
[10]
Juraj Mavracic, Callum J Court, Taketomo Isazawa, Stephen R Elliott, and Jacqueline M Cole. Chem- DataExtractor 2.0: Autopopulated ontologies for materials science.Journal of Chemical Information and Modeling, 61(9):4280–4289, 2021
work page 2021
-
[11]
Taketomo Isazawa and Jacqueline M Cole. Single model for organic and inorganic chemical named entity recognition in ChemDataExtractor.Journal of Chemical Information and Modeling, 62(5):1207–1213, 2022
work page 2022
-
[12]
Kohulan Rajan, Henning Otto Brinkhaus, M Isabel Agea, Achim Zielesny, and Christoph Steinbeck. DECIMER.ai: An open platform for automated optical chemical structure identification, segmentation and recognition in scientific publications.Nature Communications, 14(1):5045, 2023
work page 2023
-
[13]
John Dagdelen, Alexander Dunn, Sanghoon Lee, Nicholas Walker, Andrew S Rosen, Gerbrand Ceder, Kristin A Persson, and Anubhav Jain. Structured information extraction from scientific text with large language models.Nature Communications, 15(1):1418, 2024
work page 2024
-
[14]
Maciej P Polak and Dane Morgan. Extracting accurate materials data from research papers with conversa- tional language models and prompt engineering.Nature Communications, 15(1):1569, 2024
work page 2024
-
[15]
Tanishq Gupta, Mohd Zaki, NM Anoop Krishnan, and Mausam. MatSciBERT: A materials domain language model for text mining and information extraction.npj Computational Materials, 8(1):102, 2022
work page 2022
-
[16]
Defne Circi, Ghazal Khalighinejad, Anlan Chen, Bhuwan Dhingra, and L Catherine Brinson. How well do large language models understand tables in materials science?Integrating Materials and Manufacturing Innovation, 13(3):669–687, 2024
work page 2024
-
[17]
Sonakshi Gupta, Akhlak Mahmood, Pranav Shetty, Aishat Adeboye, and Rampi Ramprasad. Data extraction from polymer literature using large language models.Communications Materials, 5(1):269, 2024
work page 2024
-
[18]
Magdalena Lederbauer, Siddharth Betala, Xiyao Li, Ayush Jain, Amine Sehaba, Georgia Channing, Grégoire Germain, Anamaria Leonescu, Faris Flaifil, Alfonso Amayuelas, et al. LeMat-Synth: A multi- modal toolbox to curate broad synthesis procedure databases from scientific literature.arXiv preprint arXiv:2510.26824, 2025
-
[19]
Mara Schilling-Wilhelmi, Martiño Ríos-García, Sherjeel Shabih, María Victoria Gil, Santiago Miret, Christoph T Koch, José A Márquez, and Kevin Maik Jablonka. From text to insight: large language models for chemical data extraction.Chemical Society Reviews, 54(3):1125–1150, 2025
work page 2025
-
[20]
Nils Dunlop, Francisco Erazo, Farzaneh Jalalypour, and Rocío Mercado. Predicting PROTAC-mediated ternary complexes with AlphaFold3 and Boltz-1.Digital Discovery, 4(12):3782–3809, 2025
work page 2025
-
[21]
Stefano Ribes, Ranxuan Zhang, Télio Cropsal, Anders Källberg, Christian Tyrchan, Eva Nittinger, and Rocío Mercado. PROTAC-Splitter: a machine learning framework for automated identification of PROTAC substructures.Journal of Cheminformatics, 18(1):30, 2026
work page 2026
-
[22]
An autonomous living database for perovskite photovoltaics.arXiv preprint arXiv:2601.17807, 2026
Sherjeel Shabih, Hampus Näsström, Sharat Patil, Asmin Askin, Keely Dodd-Clements, Jessica He- lisa Hautrive Rossato, Hugo Gajardoni de Lemos, Yuxin Liu, Florian Mathies, Natalia Maticiuc, et al. An autonomous living database for perovskite photovoltaics.arXiv preprint arXiv:2601.17807, 2026
-
[23]
Dimakis, Ion Stoica, Dan Klein, Matei Zaharia, and Omar Khattab
Lakshya A Agrawal, Shangyin Tan, Dilara Soylu, Noah Ziems, Rishi Khare, Krista Opsahl-Ong, Arnav Singhvi, Herumb Shandilya, Michael J Ryan, Meng Jiang, Christopher Potts, Koushik Sen, Alexandros G. Dimakis, Ion Stoica, Dan Klein, Matei Zaharia, and Omar Khattab. GEPA: Reflective prompt evolution can outperform reinforcement learning, 2025
work page 2025
-
[24]
Joshi, Hanna Moazam, Heather Miller, Matei Zaharia, and Christopher Potts
Omar Khattab, Arnav Singhvi, Paridhi Maheshwari, Zhiyuan Zhang, Keshav Santhanam, Sri Vardhamanan, Saiful Haq, Ashutosh Sharma, Thomas T. Joshi, Hanna Moazam, Heather Miller, Matei Zaharia, and Christopher Potts. DSPy: Compiling declarative language model calls into self-improving pipelines. 2024. 11
work page 2024
-
[25]
Martiño Ríos-García and Kevin Maik Jablonka. LLM-as-Judge meets LLM-as-Optimizer: Enhancing or- ganic data extraction evaluations through dual LLM approaches. InAI4Mat-ICLR-2025: AI for Accelerated Materials Design Workshop, ICLR 2025, 2025
work page 2025
-
[26]
Gisele Nishiguchi, Fatemeh Keramatnia, Jaeki Min, Yunchao Chang, Barbara Jonchere, Sourav Das, Marisa Actis, Jeanine Price, Divyabharathi Chepyala, Brandon Young, et al. Identification of potent, selective, and orally bioavailable small-molecule GSPT1/2 degraders from a focused library of cereblon modulators.Journal of Medicinal Chemistry, 64(11):7296–7311, 2021
work page 2021
- [27]
-
[28]
python-docx.https://python-docx.readthedocs.io/, 2026
Steve Canny. python-docx.https://python-docx.readthedocs.io/, 2026
work page 2026
-
[29]
Chemical name to structure: OPSIN, an open source solution, 2011
Daniel M Lowe, Peter T Corbett, Peter Murray-Rust, and Robert C Glen. Chemical name to structure: OPSIN, an open source solution, 2011
work page 2011
-
[30]
Pubchem 2025 update.Nucleic Acids Research, 53(D1):D1516– D1525, 2025
Sunghwan Kim, Jie Chen, Tiejun Cheng, Asta Gindulyte, Jia He, Siqian He, Qingliang Li, Benjamin A Shoemaker, Paul A Thiessen, Bo Yu, et al. Pubchem 2025 update.Nucleic Acids Research, 53(D1):D1516– D1525, 2025
work page 2025
-
[31]
Matthew Swain. PubChemPy, April 2017. 12 A Ground truth expert paper annotation Selected publications.This section lists the publications used for expert ground-truth annotation from the processed MG and PROTAC cohorts (Tabs. 3 & 4). Ground-truth annotations were created by expert PhD and postdoc chemists & machine learning researchers (details below). Pu...
-
[32]
Look for IUPAC names, SMILES strings, or full chemical names for compounds you previously extracted (e.g., if you extracted "compound 5", look for its IUPAC name, SMILES string, or chemical structure name). ãÑ ãÑ
-
[33]
- This helps match compounds across different parts of the paper
For each compound where you find an IUPAC name or SMILES: - Keep all other fields (Compound_Name, Degradation_Target, DC50, Dmax, etc.) EXACTLY the same as your previous extraction.ãÑ - ONLY fill in or update the IUPAC_Name and SMILES fields. - This helps match compounds across different parts of the paper. Secondary Task: Extract Any New Compound Data
-
[34]
If you find any NEW molecular glue degradation data that was NOT in your previous extraction, extract it completely.ãÑ
-
[35]
Include any additional compounds, assays, or cell lines found in this file. Important Notes: - When updating existing compounds with IUPAC names or SMILES, preserve all original data.ãÑ - If a compound already has an IUPAC name or SMILES, only update if you find a more complete or accurate one.ãÑ - Return all data points (both updated and new) in the same...
-
[36]
Identify any molecular glue degradation data that was NOT in your previous extraction.ãÑ
-
[37]
Extract any missing fields (DC50, Dmax, Cell_Line, Assay, etc.) that can now be filled.ãÑ
-
[38]
Look for IUPAC names, SMILES strings, or full chemical names for compounds you previously extracted.ãÑ
-
[39]
Include any new compounds, assays, or cell lines found in this file. Important for IUPAC Names and SMILES: - If you find IUPAC names or SMILES strings for compounds you already extracted, fill in the IUPAC_Name and SMILES fields while keeping otherãÑ fields the same. Return all data points (both updated and new) in the same JSON format as before. Output a...
-
[40]
Identify any compounds you extracted that are still missing IUPAC names or SMILES stringsãÑ
-
[41]
Search the main text for IUPAC names, SMILES strings, or full chemical names for these compoundsãÑ
-
[42]
For each compound where you find an IUPAC name or SMILES: 23 - Keep all other fields (Compound_Name, Degradation_Target, DC50, Dmax, etc.) EXACTLY the same as your previous extractionãÑ - ONLY fill in or update the IUPAC_Name and SMILES fields - Match the compound by its name (e.g., "compound 5", "molecule A") or by its degradation dataãÑ **Important:** -...
-
[43]
**Target Sections**: You are ONLY allowed to modify`# Goal`and`# Extraction Principles`.ãÑ
-
[44]
**Frozen Sections**: Do **NOT** modify`# Required Fields`or`# Output Format` under any circumstances.ãÑ
-
[45]
Identify the root cause of the errors and fix the prompt generally
**Generalization**: Avoid overfitting. Identify the root cause of the errors and fix the prompt generally. Do not hardcode details about specific compounds, targets, recruiters, or cell lines found in the error report. ãÑ ãÑ
-
[46]
**Strict Conciseness**: You may only update or add **maximum 2 sentences** in total. Prioritize the most critical fix. The total added/modified text should be approximately \leq {char_limit} characters. ãÑ ãÑ # Output Format Return **only** a valid, raw JSON object (no markdown code blocks, no pre-text): {{ "updated_prompt": "The complete updated prompt s...
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.