Recognition: 2 theorem links
· Lean TheoremDiffSegLung: Diffusion Radiomic Distillation for Unsupervised Lung Pathology Segmentation
Pith reviewed 2026-05-13 05:15 UTC · model grok-4.3
The pith
Handcrafted radiomic descriptors can guide a diffusion model to segment lung pathologies in CT scans without annotations.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
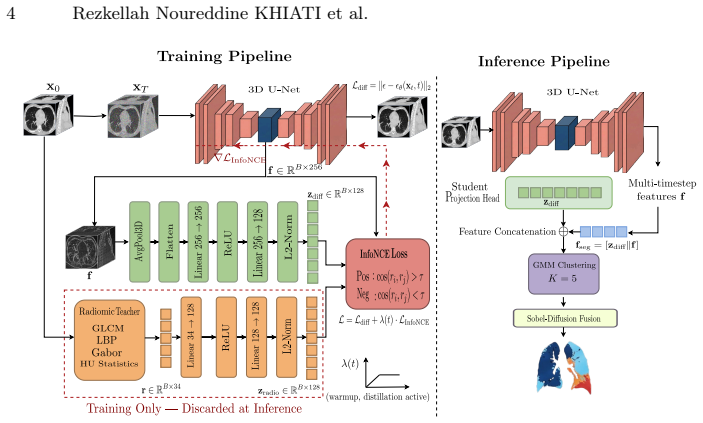
Diffusion Radiomic Distillation uses handcrafted radiomic descriptors as a physics-grounded teacher to shape the bottleneck of a 3D diffusion U-Net via a contrastive objective, transferring pathology-discriminative structure into the learned representation without annotations. At inference the teacher is discarded and multitimestep bottleneck features are clustered by a Gaussian Mixture Model with HU-guided label assignment, followed by boundary refinement. On 190 expert-annotated axial slices from four heterogeneous CT cohorts this yields improved segmentation across four pathology classes over unsupervised baselines and higher generation fidelity than prior CT diffusion models.
What carries the argument
Diffusion Radiomic Distillation: radiomic descriptors serve as a teacher to shape the diffusion U-Net bottleneck through a contrastive objective, enabling later clustering of the resulting features.
If this is right
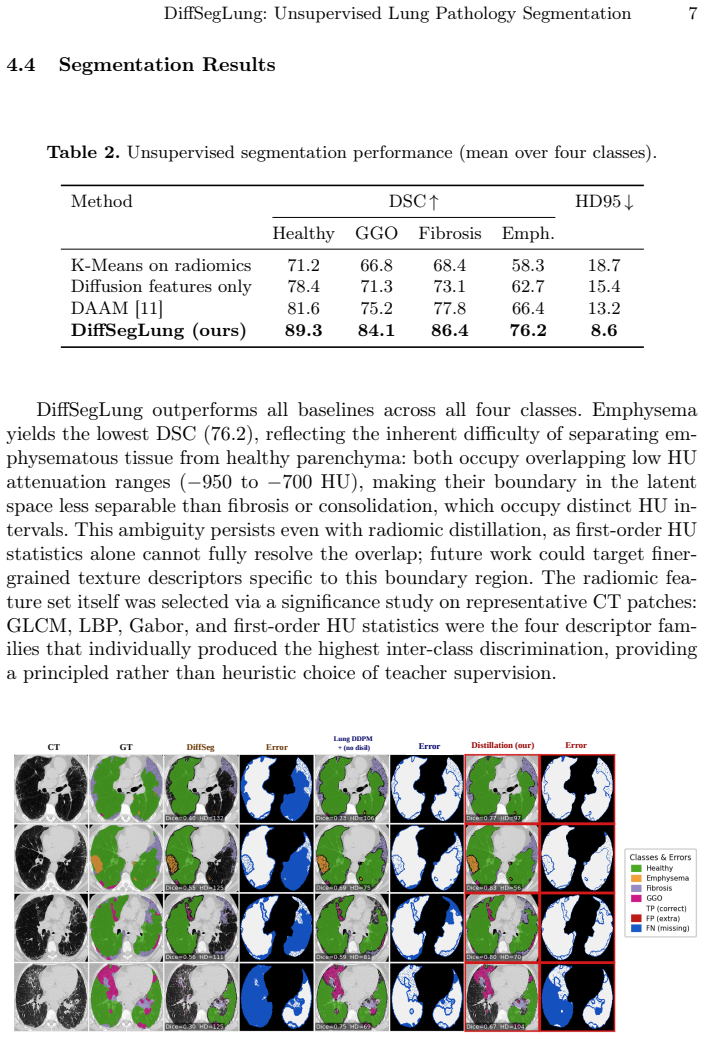
- Segmentation performance improves for all four tested pathology classes on data drawn from multiple CT sources.
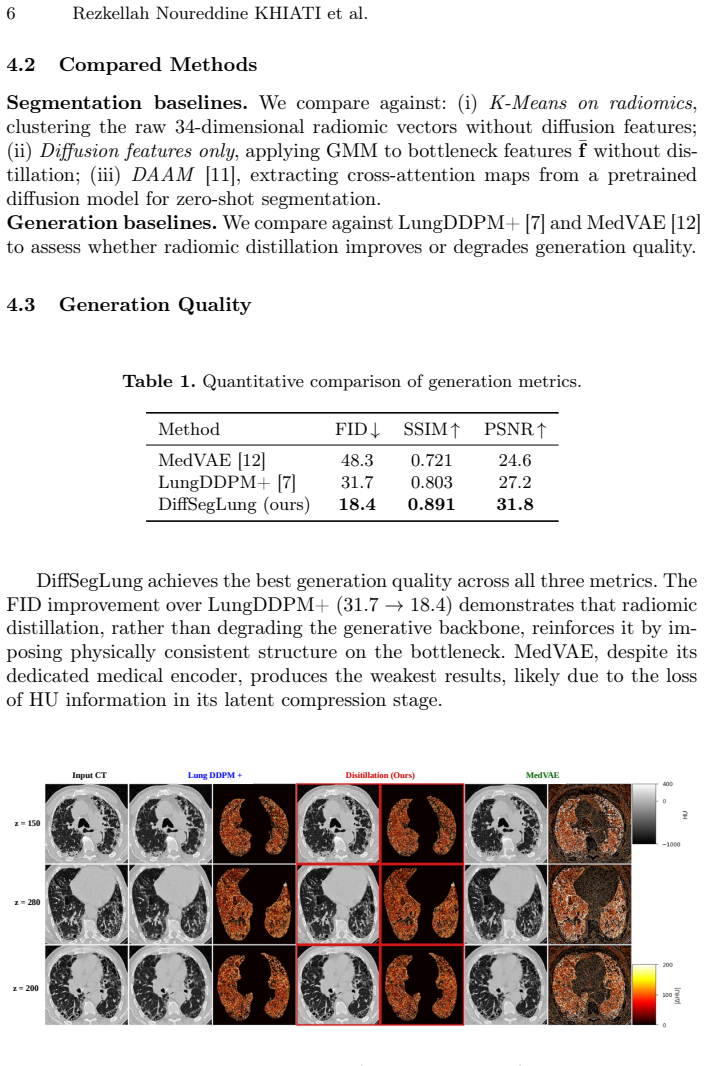
- The diffusion model itself generates higher-fidelity CT images than previous unsupervised approaches.
- The radiomic teacher can be removed at test time, leaving only feature clustering and refinement.
- The same trained features support consistent labeling across heterogeneous acquisition protocols without retraining.
Where Pith is reading between the lines
- The distillation step could be tried on other density-sensitive imaging tasks such as liver or bone lesion segmentation where similar physical descriptors exist.
- If the clustered features prove stable, they might later support downstream clinical predictions like progression risk without new labels.
- Extending the contrastive objective to include additional physical signals beyond density could further reduce reliance on handcrafted descriptors.
Load-bearing premise
Handcrafted radiomic descriptors based on density values reliably encode the differences between pathology classes so that contrastive training plus clustering can separate them correctly without any labels or manual tuning.
What would settle it
Run the trained model on a fresh collection of expert-annotated CT slices from a new scanner or patient group and measure whether the segmentation overlap scores for each pathology class fail to exceed those obtained by clustering raw intensity values or other simple unsupervised baselines.
Figures

read the original abstract
Unsupervised segmentation of pulmonary pathologies in CT remains an open challenge due to the absence of annotated multi pathology cohorts and the failure of existing diffusion-based methods to exploit the quantitative Hounsfield Unit (HU) signal that physically distinguishes tissue classes. To address this, we propose DiffSegLung,a framework that introduces Diffusion Radiomic Distillation, in which handcrafted radiomic descriptors serve as a physics grounded teacher to shape the bottleneck of a 3D diffusion U-Net via a contrastive objective, transferring pathology discriminative structure into the learned representation without any annotations. At inference, the teacher is discarded and multitimestep bottleneck features are clustered by a Gaussian Mixture Model with HU-guided label assignment, followed by Sobel Diffusion Fusion for boundary refinement. Evaluated on 190 expert annotated axial slices drawn from four heterogeneous CT cohorts, Diff-SegLung improves segmentation across all four pathology classes over unsupervised baselines and improves generation fidelity over prior CT diffusion models.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper introduces DiffSegLung, a framework for unsupervised segmentation of lung pathologies in CT images. It employs Diffusion Radiomic Distillation where handcrafted radiomic descriptors serve as a physics-grounded teacher to guide the bottleneck features of a 3D diffusion U-Net using a contrastive objective. At inference, multi-timestep features are clustered using a Gaussian Mixture Model with HU-guided label assignment and refined with Sobel Diffusion Fusion. The method is claimed to outperform unsupervised baselines on 190 expert-annotated axial slices from four heterogeneous CT cohorts across four pathology classes and to improve generation fidelity over prior CT diffusion models.
Significance. If the results hold, this work could be significant for the field of medical image segmentation by providing an unsupervised method that leverages quantitative HU information and radiomics to handle multiple pathologies without annotations. The distillation approach from radiomics to diffusion models is innovative and addresses a key challenge in pulmonary CT analysis where labeled data is limited. The evaluation on heterogeneous cohorts adds to its potential applicability.
major comments (3)
- [Abstract] Abstract: The abstract claims performance improvements over unsupervised baselines but provides no quantitative metrics, error bars, ablation studies, or details on data splits and hyperparameter selection. This undermines the ability to assess the central claim of superiority, as the evaluation is load-bearing for the paper's contribution.
- [Method] Method (Diffusion Radiomic Distillation): The core assumption that handcrafted radiomic descriptors reliably encode pathology-discriminative structure based on HU values is not sufficiently validated. No evidence or analysis is presented showing that these features separate the four pathology classes across the heterogeneous cohorts, which is critical for the contrastive objective to transfer useful structure and for the subsequent GMM clustering to succeed without supervision.
- [Experiments] Experiments: The evaluation on 190 slices from four cohorts lacks details on how the cohorts were split for training and testing, the specific pathology classes, and statistical comparisons. Without these, the claim of improvement across all classes cannot be properly evaluated.
minor comments (2)
- [Abstract] There is an inconsistency in the naming: 'DiffSegLung' and 'Diff-SegLung' are used interchangeably.
- [Abstract] The term 'multitimestep' should be hyphenated as 'multi-timestep' for clarity.
Simulated Author's Rebuttal
We thank the referee for the constructive feedback on our manuscript. We have carefully reviewed each major comment and provide point-by-point responses below. We agree that several clarifications and additions will strengthen the paper and will incorporate them in the revised version.
read point-by-point responses
-
Referee: [Abstract] Abstract: The abstract claims performance improvements over unsupervised baselines but provides no quantitative metrics, error bars, ablation studies, or details on data splits and hyperparameter selection. This undermines the ability to assess the central claim of superiority, as the evaluation is load-bearing for the paper's contribution.
Authors: We agree that the abstract would benefit from including key quantitative results to better support the central claims. In the revised manuscript, we will update the abstract to report mean Dice scores (with standard deviations) for each pathology class, along with a brief reference to the cross-cohort evaluation protocol. Ablation studies and hyperparameter details remain in the main Experiments section and supplementary material, but we will add a concise mention of the evaluation setup in the abstract for improved readability. revision: yes
-
Referee: [Method] Method (Diffusion Radiomic Distillation): The core assumption that handcrafted radiomic descriptors reliably encode pathology-discriminative structure based on HU values is not sufficiently validated. No evidence or analysis is presented showing that these features separate the four pathology classes across the heterogeneous cohorts, which is critical for the contrastive objective to transfer useful structure and for the subsequent GMM clustering to succeed without supervision.
Authors: We acknowledge that an explicit validation of the radiomic features' ability to discriminate pathology classes would strengthen the methodological justification. We will add a new subsection or figure in the revised manuscript (e.g., t-SNE embeddings or class-separation metrics such as silhouette scores computed on the radiomic descriptors) demonstrating separation across the four classes and heterogeneous cohorts. This analysis will directly support the rationale for the contrastive distillation and the unsupervised GMM step. revision: yes
-
Referee: [Experiments] Experiments: The evaluation on 190 slices from four cohorts lacks details on how the cohorts were split for training and testing, the specific pathology classes, and statistical comparisons. Without these, the claim of improvement across all classes cannot be properly evaluated.
Authors: We agree that additional experimental details are required for full reproducibility and assessment. In the revised Experiments section, we will explicitly describe the training/testing splits (e.g., leave-one-cohort-out or stratified per-cohort ratios), list the four pathology classes, and report statistical comparisons (paired t-tests or Wilcoxon signed-rank tests with p-values) between DiffSegLung and baselines. These additions will directly address the evaluation of improvements across classes. revision: yes
Circularity Check
No significant circularity; derivation relies on external handcrafted radiomics and standard clustering
full rationale
The paper introduces handcrafted radiomic descriptors (based on HU values) as an external physics-grounded teacher for contrastive distillation into the 3D diffusion U-Net bottleneck. At inference the teacher is dropped and multitimestep features are clustered via GMM with HU-guided assignment. No equations or steps reduce the central claim to a quantity defined by parameters fitted inside the same loop, nor do they rely on self-citation chains or ansatzes smuggled from prior author work. The radiomics and GMM are presented as independent inputs, making the chain self-contained against external benchmarks.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Handcrafted radiomic descriptors provide physics-grounded, pathology-discriminative information based on Hounsfield Unit values.
Lean theorems connected to this paper
-
IndisputableMonolith/Cost/FunctionalEquation.leanwashburn_uniqueness_aczel unclearhandcrafted radiomic descriptors serve as a physics grounded teacher... 34-dimensional radiomic vector r(i) comprising GLCM statistics (14 features), LBP histograms (8 features), Gabor filter responses (8 features), and first-order HU statistics (4 features)
-
IndisputableMonolith/Foundation/AbsoluteFloorClosure.leanabsolute_floor_iff_bare_distinguishability unclearGMM with K=5 components... HU-guided label assignment... comparing each cluster’s mean HU h̄k against clinically established thresholds
Reference graph
Works this paper leans on
-
[1]
Radiology307(5), e222998 (2023)
Amudala Puchakayala, P.R., Sthanam, V.L., Nakhmani, A., et al.: Radiomics for improved detection of chronic obstructive pulmonary disease in low-dose and standard-dose chest CT scans. Radiology307(5), e222998 (2023)
work page 2023
-
[2]
Bandara, W.G.C., Nair, N.G., Patel, V.M.: DDPM-CD: Remote sensing change detection using denoising diffusion probabilistic models (2022)
work page 2022
-
[3]
Baranchuk, D., Rubachev, I., Voynov, A., Khrulkov, V., Babenko, A.: Label- efficient semantic segmentation with diffusion models (2022)
work page 2022
-
[4]
Radi- ology290(3), 783–792 (2019)
Beig, N., Khorrami, M., Alilou, M., et al.: Perinodular and intranodular radiomic features on lung CT images distinguish adenocarcinomas from granulomas. Radi- ology290(3), 783–792 (2019)
work page 2019
-
[5]
Denoising Diffusion Probabilistic Models
Ho, J., Jain, A., Abbeel, P.: Denoising diffusion probabilistic models. CoRR abs/2006.11239(2020),https://arxiv.org/abs/2006.11239
work page internal anchor Pith review Pith/arXiv arXiv 2006
-
[6]
IEEE Transactions on Biomedical Engineering73(3), 1134–1145 (2026)
Jiang, Y., Lemaréchal, Y., Plante, S., Bafaro, J., Abi-Rjeile, J., Joubert, P., De- sprés, P., Manem, V.: Lung-DDPM: Semantic layout-guided diffusion models for thoracic CT image synthesis. IEEE Transactions on Biomedical Engineering73(3), 1134–1145 (2026). https://doi.org/10.1109/tbme.2025.3599011
-
[7]
Computers in Biology and Medicine199, 111290 (2025)
Jiang, Y., Shariftabrizi, A., Manem, V.S.: Lung-DDPM+: Efficient thoracic CT image synthesis using diffusion probabilistic model. Computers in Biology and Medicine199, 111290 (2025). https://doi.org/10.1016/j.compbiomed.2025.111290
-
[8]
Advances in Neural Information Processing Systems35, 5775–5787 (2022)
Lu, C., Zhou, Y., Bao, F., Chen, J., Li, C., Zhu, J.: DPM-Solver: A fast ODE solver for diffusion probabilistic model sampling in around 10 steps. Advances in Neural Information Processing Systems35, 5775–5787 (2022)
work page 2022
-
[9]
Luo, X., Hu, M., Liao, W., Zhai, S., Song, T., Wang, G., Zhang, S.: Scribble- supervised medical image segmentation via dual-branch network and dynamically mixed pseudo labels supervision (2022)
work page 2022
-
[10]
IEEE Transactions on Medical Imaging40(12), 3820–3831 (2021)
Qin, D., Bu, J.J., Liu, Z., Shen, X., Zhou, S., Gu, J.J., Wang, Z.H., Wu, L., Dai, H.F.: Efficient medical image segmentation based on knowledge dis- tillation. IEEE Transactions on Medical Imaging40(12), 3820–3831 (2021). https://doi.org/10.1109/TMI.2021.3098703
-
[11]
Tian, J., Aggarwal, L., Colaco, A., Kira, Z., Gonzalez-Franco, M.: Diffuse, attend, and segment: Unsupervised zero-shot segmentation using stable diffusion (2024)
work page 2024
-
[12]
Varma, M., Kumar, A., van der Sluijs, R., et al.: MedVAE: Efficient automated interpretation of medical images with large-scale generalizable autoencoders (2025)
work page 2025
-
[13]
Wu, J., Fu, R., Fang, H., Zhang, Y., Yang, Y., Xiong, H., Liu, H., Xu, Y.: Med- SegDiff:Medicalimagesegmentationwithdiffusionprobabilisticmodel.In:Medical Imaging with Deep Learning. pp. 1623–1639. PMLR (2024)
work page 2024
-
[14]
Investigative Radiology 47, 596–602 (2012)
Zach, J.A., Newell, J.D., Schroeder, J.D., et al.: Quantitative computed tomogra- phy of the lungs and airways in healthy nonsmoking adults. Investigative Radiology 47, 596–602 (2012)
work page 2012
-
[15]
Computers in Biology and Medicine164, 107284 (2023)
Zhao, L., Qian, X., Guo, Y., Song, J., Hou, J., Gong, J.: MSKD: Structured knowledge distillation for efficient medical image segmen- tation. Computers in Biology and Medicine164, 107284 (2023). https://doi.org/10.1016/j.compbiomed.2023.107284
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.