Recognition: no theorem link
A Comparative Analysis of CT Degradation for LDCT Nodule Classification using Radiomics
Pith reviewed 2026-05-13 04:29 UTC · model grok-4.3
The pith
CycleGAN degradation of standard-dose CT scans creates synthetic low-dose images that train lung nodule classifiers with better balance than undegraded data.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
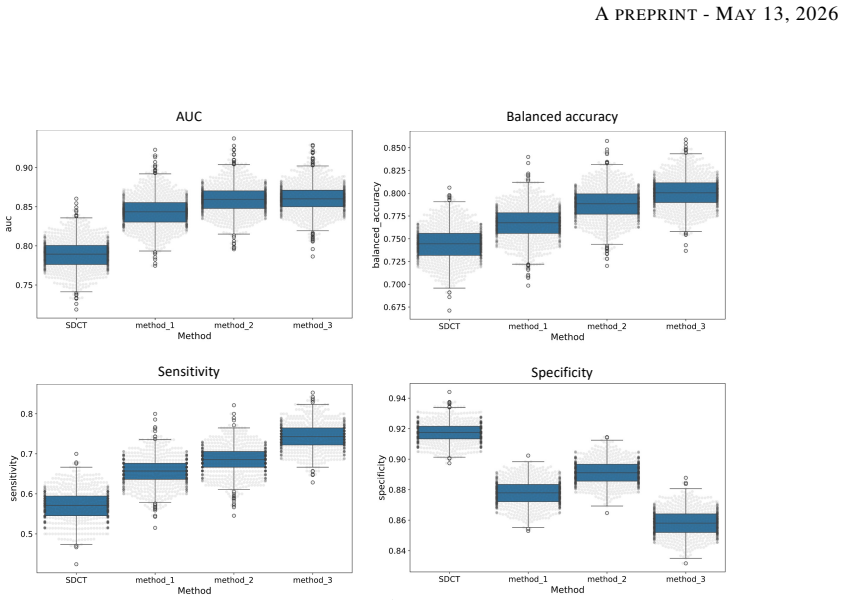
Degraded images generated using CycleGAN approach led to the most balanced performance on the classification task using Adam Booster classifier, achieving an AUC of 0.861, sensitivity of 0.743 and specificity of 0.858 in the independent test. The baseline model trained on non-degraded SDCT data failed to generalize to the real LDCT set with AUC 0.789 and sensitivity 0.571. CycleGAN also achieved the best Fréchet inception distance and kernel inception distance scores among the three degradation methods.
What carries the argument
CycleGAN unpaired image-to-image translation that simulates low-dose CT noise and contrast from standard-dose scans, followed by radiomic feature extraction for machine learning nodule classification.
If this is right
- A model trained only on standard-dose data loses sensitivity when tested on real low-dose screening scans.
- CycleGAN degradation produces images with the closest statistical match to real low-dose CT among the three methods tested.
- Synthetic low-dose data generated this way can substitute for real low-dose cases when training classifiers on the LIDC-IDRI dataset.
- The three degradation approaches differ in how well their outputs support downstream radiomics-based classification.
Where Pith is reading between the lines
- Researchers could generate large training sets for other low-dose imaging tasks without new patient scans.
- The same degradation step might be applied to improve model robustness in CT tasks beyond lung nodules.
- If the feature distributions match closely enough, this method could lower the radiation exposure needed during model development.
Load-bearing premise
Radiomic features extracted from the synthetic low-dose images behave the same way as features from real low-dose scans when used for nodule classification.
What would settle it
Train the same classifier on a held-out set of real low-dose CT cases using either the synthetic degradation pipeline or actual low-dose scans and compare the resulting AUC and sensitivity.
Figures






read the original abstract
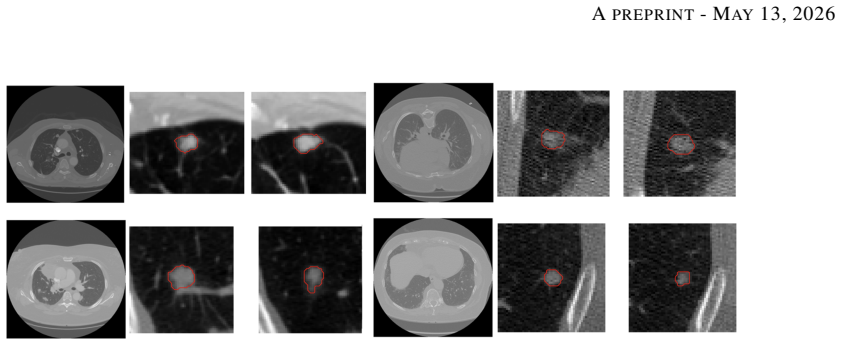
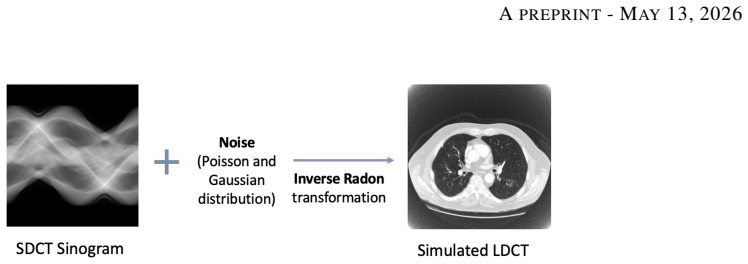
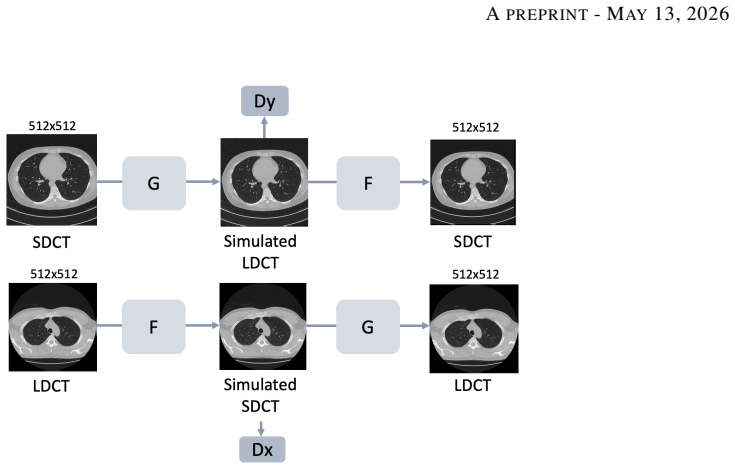
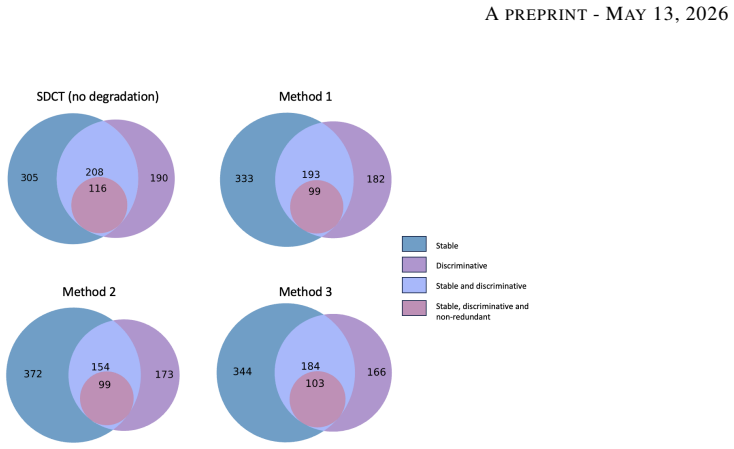
Low-dose computed tomography (LDCT) is the standard modality for lung cancer screening, known for its low radiation dose but high noise levels. While existing literature focuses on denoising LDCT images, comparative research on simulating LDCT characteristics to directly use these images for model development is lacking. This study shifts the focus from denoising images to degrading available standard-dose CT (SDCT) data, generating synthetic images for data augmentation to train classifiers for screening-detected nodules. We compare three degradation methods: (1) a sinogram domain statistical noise insertion; (2) replicate a validated physics-based simulation using Pix2Pix; and (3) unpaired CycleGAN. The generated images were utilized to simulate LDCT screening scenario replacing 695 SDCT cases from the LIDC-IDRI dataset, from which radiomic features were extracted to train machine learning models for lung nodule classification. Regarding image quality, CycleGAN achieved the best Fr\'echet inception distance (0.1734) and kernel inception distance (0.0813; 0.1002) scores, indicating distributional alignment with the target low-dose domain. In the nodule classification task, results confirmed the necessity of domain adaptation since a baseline model trained on non-degraded SDCT data failed to generalize to the real LDCT set (AUC 0.789) with a low sensitivity (0.571). Degraded images generated using CycleGAN approach led to the most balanced performance on the classification task using Adam Booster classifier, achieving an AUC of 0.861, sensitivity of 0.743 and specificity of 0.858 in the independent test. Our findings confirm that generating synthetic LDCT data from standard-dose scans is a viable strategy for training robust nodule classifiers for screening detected nodules.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper claims that among three degradation methods applied to SDCT images to simulate LDCT (sinogram statistical noise insertion, Pix2Pix, and unpaired CycleGAN), the CycleGAN approach produces synthetic images with the closest distributional alignment to real LDCT (FID 0.1734, KID 0.0813/0.1002). When used to augment the LIDC-IDRI dataset for radiomics-based nodule classification, it yields superior performance with the Adam Booster classifier on an independent real LDCT test set (AUC 0.861, sensitivity 0.743, specificity 0.858), compared to baseline SDCT training (AUC 0.789, sensitivity 0.571). The authors conclude this is a viable strategy for training robust classifiers for screening-detected nodules.
Significance. If substantiated, this work provides evidence that image-to-image translation models like CycleGAN can effectively simulate LDCT degradation for data augmentation in radiomics pipelines. This has potential significance for medical imaging AI, as it leverages more readily available SDCT data to improve model performance on noisy LDCT without additional radiation exposure or data collection. The reported improvements in balanced classification metrics highlight practical benefits for lung cancer screening applications.
major comments (1)
- [Results] The improved classification performance with CycleGAN-degraded images (AUC 0.861) is presented as resulting from better simulation of LDCT characteristics. However, the manuscript provides no quantitative analysis comparing the distributions of radiomic features extracted from the CycleGAN synthetic LDCT images versus real LDCT images (such as per-feature statistical tests or multivariate distances). This is a load-bearing gap because the central claim that the synthetics enable generalization to real LDCT depends on the features behaving similarly, rather than the improvement arising from generic augmentation or other artifacts.
minor comments (2)
- [Methods] The experimental setup lacks explicit description of how the 695 SDCT cases were selected for replacement, the composition of the independent test set, and whether stratified splitting or cross-validation was employed to ensure robust evaluation.
- [Abstract] The KID scores are reported as 0.0813; 0.1002 without clarifying what the two values represent (e.g., different kernel sizes or train/test splits).
Simulated Author's Rebuttal
We thank the referee for their thorough and constructive review of our manuscript. We have carefully considered the major comment and provide a point-by-point response below, including plans for revision.
read point-by-point responses
-
Referee: [Results] The improved classification performance with CycleGAN-degraded images (AUC 0.861) is presented as resulting from better simulation of LDCT characteristics. However, the manuscript provides no quantitative analysis comparing the distributions of radiomic features extracted from the CycleGAN synthetic LDCT images versus real LDCT images (such as per-feature statistical tests or multivariate distances). This is a load-bearing gap because the central claim that the synthetics enable generalization to real LDCT depends on the features behaving similarly, rather than the improvement arising from generic augmentation or other artifacts.
Authors: We agree that this represents a substantive gap in the current manuscript. While the CycleGAN approach demonstrated the strongest image-level distributional alignment via FID (0.1734) and KID (0.0813/0.1002), these metrics do not directly confirm similarity in the specific radiomic feature space used for nodule classification. The observed improvement in generalization to the independent real LDCT test set (AUC 0.861 vs. baseline 0.789) could indeed partly reflect generic augmentation benefits rather than targeted LDCT simulation. In the revised manuscript, we will add quantitative comparisons of radiomic feature distributions, including per-feature statistical tests (e.g., Kolmogorov-Smirnov or Wilcoxon rank-sum tests with multiple-comparison correction) and multivariate measures (e.g., Mahalanobis distance or Earth Mover's Distance on the feature vectors) between CycleGAN-degraded images and real LDCT. We will also report these results alongside the classification metrics to better substantiate the mechanism of improvement. revision: yes
Circularity Check
Empirical comparison study with no derivation chain or self-referential reductions
full rationale
The manuscript is a standard empirical ML pipeline: three image-degradation techniques (sinogram noise insertion, Pix2Pix, CycleGAN) are applied to SDCT scans from LIDC-IDRI to produce synthetic LDCT images; radiomic features are extracted; classifiers are trained and evaluated on a held-out real LDCT test set. No equations, uniqueness theorems, fitted parameters renamed as predictions, or self-citations are invoked to derive the central performance numbers (AUC 0.861 etc.). All reported metrics are direct experimental outcomes on external data, so the result does not reduce to its inputs by construction.
Axiom & Free-Parameter Ledger
free parameters (2)
- CycleGAN training parameters
- Noise insertion parameters
axioms (1)
- domain assumption Synthetic degraded images preserve discriminative radiomic features for nodule classification
Reference graph
Works this paper leans on
-
[1]
H. Sung et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries.CA: A Cancer Journal for Clinicians, 71(3):209–249, May 2021
work page 2020
-
[2]
H. J. de Koning et al. Reduced lung-cancer mortality with volume CT screening in a randomized trial.New England Journal of Medicine, 382(6):503–513, February 2020
work page 2020
-
[3]
The National Lung Screening Trial Research Team. Reduced lung-cancer mortality with low-dose computed tomographic screening.New England Journal of Medicine, 365(5):395–409, August 2011
work page 2011
-
[4]
W. Kim, S. Y . Jeon, G. Byun, H. Yoo, and J. H. Choi. A systematic review of deep learning-based denoising for low-dose computed tomography from a perceptual quality perspective.Biomedical Engineering Letters, 14(6):1153–1173, August 2024
work page 2024
-
[5]
K. A. S. H. Kulathilake, N. A. Abdullah, A. Q. M. Sabri, and K. W. Lai. A review on deep learning approaches for low-dose computed tomography restoration.Complex & Intelligent Systems, 9(3):2713–2745, June 2023. 13 APREPRINT- MAY13, 2026
work page 2023
-
[6]
T. Zhao, M. McNitt-Gray, and D. Ruan. A convolutional neural network for ultra-low-dose CT denoising and emphysema screening.Medical Physics, 46(9):3941–3950, September 2019
work page 2019
-
[7]
M. Gholizadeh-Ansari, J. Alirezaie, and P. Babyn. Deep learning for low-dose CT denoising using perceptual loss and edge detection layer.Journal of Digital Imaging, 33(2):504–515, September 2019
work page 2019
-
[8]
H. Chen et al. Low-dose CT denoising with convolutional neural network. InProceedings of the International Symposium on Biomedical Imaging (ISBI), pages 143–146, June 2017
work page 2017
- [9]
-
[10]
C. H. McCollough et al. Low-dose CT for the detection and classification of metastatic liver lesions: Results of the 2016 low dose CT grand challenge.Medical Physics, 44(10):e339–e352, October 2017
work page 2016
-
[11]
S. G. Armato et al. The lung image database consortium (LIDC) and image database resource initiative (IDRI): A completed reference database of lung nodules on CT scans.Medical Physics, 38(2), 2011
work page 2011
-
[12]
T. R. Moen et al. Low-dose CT image and projection dataset.Medical Physics, 48(2):902–911, February 2021
work page 2021
-
[13]
T. Snowsill et al. Low-dose computed tomography for lung cancer screening in high-risk populations: a systematic review and economic evaluation.Health Technology Assessment, 22(69):1–276, November 2018
work page 2018
-
[14]
C. Rampinelli, D. Origgi, and M. Bellomi. Low-dose CT: technique, reading methods and image interpretation. Cancer Imaging, 12(3):548, 2013
work page 2013
-
[15]
L. Yu, M. Shiung, D. Jondal, and C. H. McCollough. Development and validation of a practical lower-dose- simulation tool for optimizing computed tomography scan protocols.Journal of Computer Assisted Tomography, 36(4):477–487, 2012
work page 2012
-
[16]
D. Zeng et al. A simple low-dose X-ray CT simulation from high-dose scan.IEEE Transactions on Nuclear Science, 62(5):2226–2233, October 2015
work page 2015
- [17]
-
[18]
J. Y . Zhu, T. Park, P. Isola, and A. A. Efros. Unpaired image-to-image translation using cycle-consistent adversarial networks. InProceedings of the IEEE International Conference on Computer Vision (ICCV), pages 2242–2251, March 2017
work page 2017
-
[19]
J. Liu, A. Corti, V . D. A. Corino, and L. Mainardi. Lung nodule classification using radiomics model trained on degraded SDCT images.Computer Methods and Programs in Biomedicine, 257:108474, December 2024
work page 2024
-
[20]
J. J. M. van Griethuysen et al. Computational radiomics system to decode the radiographic phenotype.Cancer Research, 77(21):e104–e107, November 2017
work page 2017
-
[21]
F. Lo Iacono, R. Maragna, G. Pontone, and V . D. A. Corino. A novel data augmentation method for radiomics analysis using image perturbations.Journal of Imaging Informatics in Medicine, 37(5):2401–2414, May 2024
work page 2024
-
[22]
J. Christensen et al. ACR lung-RADS v2022: Assessment categories and management recommendations.Chest, 165(3):738–753, March 2024. 14
work page 2024
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.