Recognition: 2 theorem links
· Lean TheoremSEMIR: Semantic Minor-Induced Representation Learning on Graphs for Visual Segmentation
Pith reviewed 2026-05-13 07:05 UTC · model grok-4.3
The pith
SEMIR learns a compact boundary-aligned graph minor from the pixel grid to enable efficient GNN inference with exact lifting back to original labels.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
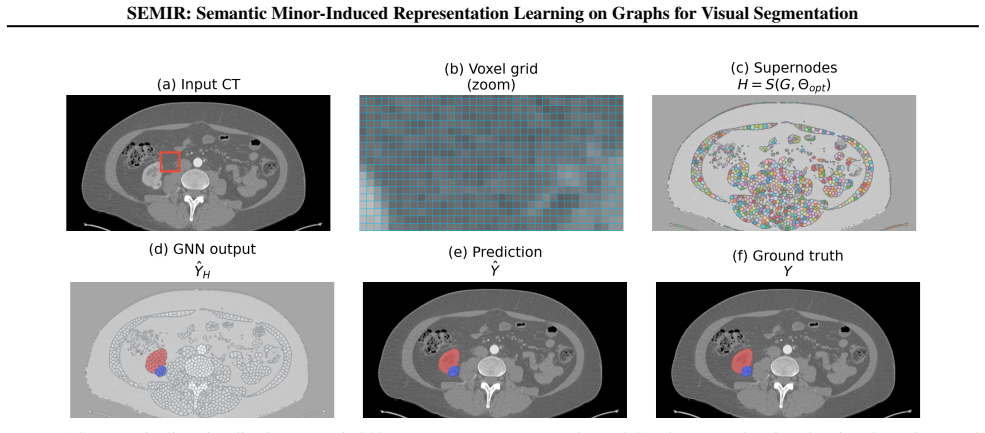
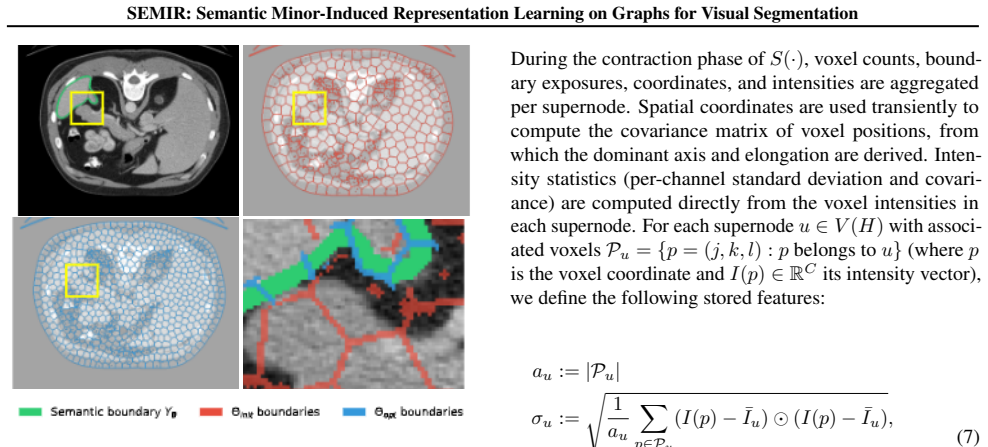
SEMIR transforms the underlying grid graph into a compact, boundary-aligned graph minor through parameterized edge contraction, node deletion, and edge deletion, while preserving an exact lifting map from minor predictions to lattice labels. Minor construction is treated as a few-shot structure learning problem solved by maximizing agreement between predicted boundary elements and target semantic edges under a boundary Dice criterion; the induced minor is then annotated with scale- and rotation-robust descriptors and processed by a GNN with relational edge features to produce labels that decode exactly to the input image.
What carries the argument
The parameterized graph minor formed by edge contraction, node deletion, and edge deletion that maintains an exact, invertible lifting map from minor-node predictions to original-grid labels.
If this is right
- Inference cost becomes independent of native image resolution once the minor is built.
- Boundary evidence for small structures is retained rather than attenuated by downsampling.
- Region-level GNN inference replaces voxel-level computation while still recovering pixel-accurate labels.
- The same learned-minor pipeline can be applied to any high-resolution grid data that contains sparse semantic targets.
- Minor parameters adapt per task and dataset without hand-tuned regionization steps.
Where Pith is reading between the lines
- The exact lifting property could allow the framework to be inserted as a drop-in replacement for pooling layers in other vision networks that require invertible downsampling.
- Because the minor is learned from boundary agreement rather than fixed heuristics, it may generalize to non-medical domains such as satellite or microscopy imagery where minority objects are similarly sparse.
- The separation of structure learning from label inference opens the possibility of reusing the same minor across multiple related tasks on the same image domain.
- If the minor remains small and stable across image batches, it could support streaming or memory-constrained deployment on large volumes.
Load-bearing premise
The boundary Dice objective produces a topology-preserving minor whose GNN predictions remain faithful to the original minority structures once lifted back to the lattice.
What would settle it
An experiment in which the lifted labels from minor predictions differ from ground-truth boundaries by more than a small tolerance on held-out images, or in which the method shows no Dice gain on minority structures despite successful minor construction.
Figures

read the original abstract
Segmenting small and sparse structures in large-scale images is fundamentally constrained by voxel-level, lattice-bound computation and extreme class imbalance -- dense, full-resolution inference scales poorly and forces most pipelines to rely on fixed regionization or downsampling, coupling computational cost to image resolution and attenuating boundary evidence precisely where minority structures are most informative. We introduce SEMIR (Semantic Minor-Induced Representation Learning), a representation framework that decouples inference from the native grid by learning a task-adapted, topology-preserving latent graph representation with exact decoding. SEMIR transforms the underlying grid graph into a compact, boundary-aligned graph minor through parameterized edge contraction, node deletion, and edge deletion, while preserving an exact lifting map from minor predictions to lattice labels. Minor construction is formalized as a few-shot structure learning problem that replaces hand-tuned preprocessing with a boundary-alignment objective: minor parameters are learned by maximizing agreement between predicted boundary elements and target-specific semantic edges under a boundary Dice criterion, and the induced minor is annotated with scale- and rotation-robust geometric and intensity descriptors and supports efficient region-level inference via message passing on a graph neural network (GNN) with relational edge features. We benchmark SEMIR on three tumor segmentation datasets -- BraTS 2021, KiTS23, and LiTS -- where targets exhibit high structural variability and distributional uncertainty. SEMIR yields consistent improvements in minority-structure Dice at practical runtime. More broadly, SEMIR establishes a framework for learning task-adapted, topology-preserving latent representations with exact decoding for high-resolution structured visual data.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper introduces SEMIR, a representation framework that decouples high-resolution image segmentation from the native voxel lattice by learning a compact, boundary-aligned graph minor of the underlying grid graph. Minor construction uses parameterized edge contraction, node deletion, and edge deletion; parameters are optimized via a boundary Dice objective that aligns predicted boundaries to target semantic edges. The resulting minor is annotated with scale- and rotation-robust descriptors and supports GNN-based inference with relational edges; an exact lifting map is claimed to recover lattice labels from minor predictions. Experiments on BraTS 2021, KiTS23, and LiTS report consistent gains in minority-structure Dice at practical runtime.
Significance. If the topology-preserving property and exact, invertible lifting map are rigorously guaranteed by the boundary-Dice procedure, the work would offer a principled way to reduce computational cost for sparse-structure segmentation while retaining boundary fidelity. The combination of learned graph minors with GNN message passing and exact decoding is a distinctive contribution that could influence other high-resolution structured-data tasks.
major comments (2)
- [§3] §3 (Minor Construction): The boundary Dice objective is the sole supervision for the contraction/deletion parameters, yet the central claim requires that the resulting minor remain topology-preserving with an exact lifting map for sparse minority nodes. No connectivity, genus, or component-count regularizer is stated, so it is possible for the learned minor to delete or over-merge internal minority nodes while still matching boundary Dice, breaking the exact lifting for sub-voxel or disconnected structures.
- [§4] §4 (GNN Inference and Lifting): The manuscript asserts that GNN predictions on the annotated minor can be lifted exactly back to lattice labels, but the lifting map is described only at the level of boundary alignment. It is unclear whether the map remains bijective after GNN message passing when the minor contains merged or deleted minority components; an explicit proof or counter-example check is needed.
minor comments (2)
- [§3] Notation for the contraction parameters and the lifting operator should be introduced with explicit symbols and a small worked example early in §3 to avoid ambiguity when reading the boundary-Dice loss.
- [§5] The abstract and §5 claim 'consistent improvements' but do not report the number of runs, standard deviations, or statistical tests; adding these would strengthen the empirical claims.
Simulated Author's Rebuttal
We thank the referee for the constructive comments on the topology preservation guarantees and the exactness of the lifting map in SEMIR. We address each major comment point by point below and will revise the manuscript to strengthen the relevant sections.
read point-by-point responses
-
Referee: [§3] §3 (Minor Construction): The boundary Dice objective is the sole supervision for the contraction/deletion parameters, yet the central claim requires that the resulting minor remain topology-preserving with an exact lifting map for sparse minority nodes. No connectivity, genus, or component-count regularizer is stated, so it is possible for the learned minor to delete or over-merge internal minority nodes while still matching boundary Dice, breaking the exact lifting for sub-voxel or disconnected structures.
Authors: We agree that no explicit connectivity, genus, or component-count regularizer is stated in the current manuscript. The boundary Dice objective is intended to align with semantic edges in a manner that implicitly constrains deletions and contractions to preserve minority component integrity, as the parameterization only permits operations on non-boundary regions while maintaining the induced minor structure. However, to provide a stronger formal guarantee against edge cases with disconnected or sub-voxel structures, we will introduce a lightweight component-preservation regularizer into the objective in the revised Section 3 and add corresponding ablation experiments. revision: yes
-
Referee: [§4] §4 (GNN Inference and Lifting): The manuscript asserts that GNN predictions on the annotated minor can be lifted exactly back to lattice labels, but the lifting map is described only at the level of boundary alignment. It is unclear whether the map remains bijective after GNN message passing when the minor contains merged or deleted minority components; an explicit proof or counter-example check is needed.
Authors: The lifting map is defined solely during minor construction via the history of contractions and deletions and is independent of the subsequent GNN message passing, which only assigns labels to existing minor nodes without altering the mapping. By construction, the map is bijective for all nodes retained in the minor. We will add an explicit proof of bijectivity (showing invariance to GNN operations) to the revised Section 4, together with a counter-example verification on synthetic cases involving merged minority components. revision: yes
Circularity Check
No significant circularity: derivation relies on supervised optimization of minor parameters followed by independent GNN inference
full rationale
The paper presents SEMIR as a supervised framework where minor parameters are optimized on training data via boundary Dice to align with semantic edges, after which a GNN performs region-level inference on the resulting minor with an exact lifting map back to lattice labels. This is a standard end-to-end learning pipeline with no equation or step that reduces the final predictions to the boundary fit by construction. No self-citations are invoked to justify uniqueness theorems, ansatzes, or load-bearing premises, and the central claims are supported by empirical benchmarks on external datasets rather than tautological equivalence to inputs. The topology-preservation property is asserted as following from the choice of contraction/deletion operations, which is definitional but not circular in the sense of re-deriving the target result from itself.
Axiom & Free-Parameter Ledger
free parameters (1)
- minor contraction parameters
axioms (1)
- domain assumption Parameterized edge contraction, node deletion, and edge deletion produce a topology-preserving minor with exact lifting map
invented entities (1)
-
Semantic Minor
no independent evidence
Lean theorems connected to this paper
-
IndisputableMonolith/Foundation/RealityFromDistinction.leanreality_from_one_distinction unclearSEMIR transforms the underlying grid graph into a compact, boundary-aligned graph minor through parameterized edge contraction, node deletion, and edge deletion, while preserving an exact lifting map from minor predictions to lattice labels.
-
IndisputableMonolith/Foundation/AlexanderDuality.leanalexander_duality_circle_linking unclearThe minor construction is governed by a small parameter set Θ, optimized via black-box few-shot optimization on boundary alignment objective.
Reference graph
Works this paper leans on
-
[1]
SLIC Superpixels Compared to State-of-the-Art Superpixel Methods , year=
Achanta, Radhakrishna and Shaji, Appu and Smith, Kevin and Lucchi, Aurelien and Fua, Pascal and Süsstrunk, Sabine , journal=. SLIC Superpixels Compared to State-of-the-Art Superpixel Methods , year=
-
[2]
Image segmentation methods: Overview, challenges, and future directions , author=. 2024 Seventh International Women in Data Science Conference at Prince Sultan University (WiDS PSU) , pages=. 2024 , organization=
work page 2024
-
[5]
and Villarini, Barbara , TITLE =
El Badaoui, Rim and Bonmati Coll, Ester and Psarrou, Alexandra and Asaturyan, Hykoush A. and Villarini, Barbara , TITLE =. Journal of Imaging , VOLUME =. 2025 , NUMBER =
work page 2025
-
[6]
The RSNA-ASNR-MICCAI BraTS 2021 Benchmark on Brain Tumor Segmentation and Radiogenomic Classification , author=. 2021 , eprint=
work page 2021
-
[7]
Algorithms for Hyper-Parameter Optimization , url =
Bergstra, James and Bardenet, R\'. Algorithms for Hyper-Parameter Optimization , url =. Advances in Neural Information Processing Systems , editor =
-
[8]
International Conference on Neural Information Processing , pages=
Deep extremely randomized trees , author=. International Conference on Neural Information Processing , pages=. 2019 , organization=
work page 2019
-
[10]
Bolya, Daniel and Fu, Cheng-Yang and Dai, Xiaoliang and Zhang, Peizhao and Feichtenhofer, Christoph and Hoffman, Judy , booktitle=. Token Merging: Your
-
[11]
Bonato, Beatrice and Nanni, Loris and Bertoldo, Alessandra , TITLE =. Sensors , VOLUME =. 2025 , NUMBER =
work page 2025
-
[14]
46th Annual IEEE Symposium on Foundations of Computer Science (FOCS'05) , pages=
Algorithmic graph minor theory: Decomposition, approximation, and coloring , author=. 46th Annual IEEE Symposium on Foundations of Computer Science (FOCS'05) , pages=. 2005 , organization=
work page 2005
-
[15]
The Lancet Digital Health , volume=
Spatially aware graph neural networks and cross-level molecular profile prediction in colon cancer histopathology: a retrospective multi-cohort study , author=. The Lancet Digital Health , volume=. 2022 , publisher=
work page 2022
-
[16]
International journal of computer vision , volume=
Efficient graph-based image segmentation , author=. International journal of computer vision , volume=. 2004 , publisher=
work page 2004
-
[17]
Gao, Yuxiao and Jiang, Yang and Peng, Yanhong and Yuan, Fujiang and Zhang, Xinyue and Wang, Jianfeng , TITLE =. Tomography , VOLUME =. 2025 , NUMBER =
work page 2025
-
[18]
Frontiers in Radiology , volume=
Evaluating the effect of voxel size on the accuracy of 3D volumetric analysis measurements of brain tumors , author=. Frontiers in Radiology , volume=. 2025 , publisher=
work page 2025
-
[19]
Instance-level semantic segmentation of nuclei based on multimodal structure encoding , author=. BMC bioinformatics , volume=. 2025 , publisher=
work page 2025
-
[20]
The KiTS21 Challenge: Automatic segmentation of kidneys, renal tumors, and renal cysts in corticomedullary-phase CT , author=. 2023 , eprint=
work page 2023
-
[21]
Pixel-wise Modulated Dice Loss for Medical Image Segmentation , author=. 2025 , eprint=
work page 2025
-
[22]
International Conference on Learning Representations , year=
Strategies for Pre-training Graph Neural Networks , author=. International Conference on Learning Representations , year=
-
[24]
3D Segmentation of Kidneys, Kidney Tumors and Cysts on CT Images - KiTS23 Challenge
Kaczmarska, Marta and Majek, Karol. 3D Segmentation of Kidneys, Kidney Tumors and Cysts on CT Images - KiTS23 Challenge. Kidney and Kidney Tumor Segmentation. 2024
work page 2024
-
[25]
Proceedings of The 2nd International Conference on Medical Imaging with Deep Learning , pages =
Boundary loss for highly unbalanced segmentation , author =. Proceedings of The 2nd International Conference on Medical Imaging with Deep Learning , pages =. 2019 , editor =
work page 2019
- [26]
-
[27]
IEEE Transactions on Pattern Analysis and Machine Intelligence , volume=
Superadditivity and convex optimization for globally optimal cell segmentation using deformable shape models , author=. IEEE Transactions on Pattern Analysis and Machine Intelligence , volume=. 2022 , publisher=
work page 2022
-
[28]
Lei, Tao and Wang, Risheng and Zhang, Yuxiao and Wan, Yong and Liu, Chang and Nandi, Asoke K. , journal=. DefED-Net: Deformable Encoder-Decoder Network for Liver and Liver Tumor Segmentation , year=
-
[29]
H-DenseUNet: Hybrid Densely Connected UNet for Liver and Tumor Segmentation From CT Volumes , year=
Li, Xiaomeng and Chen, Hao and Qi, Xiaojuan and Dou, Qi and Fu, Chi-Wing and Heng, Pheng-Ann , journal=. H-DenseUNet: Hybrid Densely Connected UNet for Liver and Tumor Segmentation From CT Volumes , year=
-
[31]
AG-MS3D-CNN multiscale attention guided 3D convolutional neural network for robust brain tumor segmentation across MRI protocols , author=. Scientific Reports , volume=. 2025 , publisher=
work page 2025
-
[34]
Bulletin of the American Mathematical Society , volume=
Graph minor theory , author=. Bulletin of the American Mathematical Society , volume=
-
[36]
Token Pooling in Vision Transformers , author=. NeurIPS Workshop , year=
-
[37]
Mienye, Ibomoiye Domor and Viriri, Serestina , TITLE =. Information , VOLUME =. 2025 , NUMBER =
work page 2025
-
[38]
Automated 3D Segmentation of Kidneys and Tumors in MICCAI KiTS 2023 Challenge
Myronenko, Andriy and Yang, Dong and He, Yufan and Xu, Daguang. Automated 3D Segmentation of Kidneys and Tumors in MICCAI KiTS 2023 Challenge. Kidney and Kidney Tumor Segmentation. 2024
work page 2023
-
[39]
Neubert, Peer and Protzel, Peter , booktitle=. Compact Watershed and Preemptive SLIC: On Improving Trade-offs of Superpixel Segmentation Algorithms , year=
-
[40]
Convolutional Occupancy Networks for Medical Imaging with Applications to the KiTS23 Challenge , author=. 2025 , booktitle=
work page 2025
-
[41]
Pandey, Sumit and Toshali and Perslev, Mathias and Dam, Erik B. Advancing Kidney, Kidney Tumor, Cyst Segmentation: A Multi-Planner U-Net Approach for the KiTS23 Challenge. Kidney and Kidney Tumor Segmentation. 2024
work page 2024
-
[42]
Peng, Yaopeng and Chen, Danny Z. and Sonka, Milan , booktitle=. U-Net V2: Rethinking the Skip Connections of U-Net for Medical Image Segmentation , year=
-
[43]
Perera, Shehan and Navard, Pouyan and Yilmaz, Alper , title =. Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) Workshops , month =. 2024 , pages =
work page 2024
-
[44]
Signal, Image and Video Processing , volume=
Lgma-net: liver and tumor segmentation methods based on local--global feature mergence and attention mechanisms , author=. Signal, Image and Video Processing , volume=. 2025 , publisher=
work page 2025
-
[45]
Graph minors. XX. Wagner's conjecture , author=. Journal of Combinatorial Theory, Series B , volume=. 2004 , publisher=
work page 2004
-
[48]
Su, Hao and Wei, Shunjun and Yan, Min and Wang, Chen and Shi, Jun and Zhang, Xiaoling , booktitle=. Object Detection and Instance Segmentation in Remote Sensing Imagery Based on Precise Mask R-CNN , year=
-
[49]
Segmenting the non-enhancing compartment of brain tumor MRIs , year=
Tobias, Schaffer and Alexander, Brawanski and Simon, Wein and Maria, Tomé Ana and Elmar W., Lang , booktitle=. Segmenting the non-enhancing compartment of brain tumor MRIs , year=
-
[52]
International Conference on Learning Representations , year=
How Powerful are Graph Neural Networks? , author=. International Conference on Learning Representations , year=
-
[53]
Quantitative Imaging in Medicine and Surgery , volume=
Automatic computed tomography image segmentation method for liver tumor based on a modified tokenized multilayer perceptron and attention mechanism , author=. Quantitative Imaging in Medicine and Surgery , volume=
-
[55]
Hierarchical Graph Representation Learning with Differentiable Pooling , author=. NeurIPS , year=
-
[57]
2025 IEEE International Conference on Real-time Computing and Robotics (RCAR) , pages=
Exploring Semi-Supervised Domain Adaptation for Precise Kidney Tumor Segmentation , author=. 2025 IEEE International Conference on Real-time Computing and Robotics (RCAR) , pages=. 2025 , organization=
work page 2025
- [59]
-
[60]
Slic superpixels compared to state-of-the-art superpixel methods
Achanta, R., Shaji, A., Smith, K., Lucchi, A., Fua, P., and Süsstrunk, S. Slic superpixels compared to state-of-the-art superpixel methods. IEEE Transactions on Pattern Analysis and Machine Intelligence, 34 0 (11): 0 2274--2282, 2012. doi:10.1109/TPAMI.2012.120
-
[61]
Alonso-Monsalve, S., Whitehead, L. H., Aurisano, A., and Sanchez, L. E. Submanifold sparse convolutional networks for automated 3d segmentation of kidneys and kidney tumours in computed tomography. arXiv preprint arXiv:2511.04334, 2025
work page internal anchor Pith review Pith/arXiv arXiv 2025
-
[62]
J., Sun, X., Qin, Y., and Langbein, F
Alwadee, E. J., Sun, X., Qin, Y., and Langbein, F. C. Latup-net: A lightweight 3d attention u-net with parallel convolutions for brain tumor segmentation. Computers in Biology and Medicine, 184: 0 109353, 2025. ISSN 0010-4825. doi:https://doi.org/10.1016/j.compbiomed.2024.109353. URL https://www.sciencedirect.com/science/article/pii/S0010482524014380
-
[63]
arXiv preprint arXiv:2107.02314 (2021)
Baid, U., Ghodasara, S., Mohan, S., Bilello, M., Calabrese, E., Colak, E., Farahani, K., Kalpathy-Cramer, J., Kitamura, F. C., Pati, S., Prevedello, L. M., Rudie, J. D., Sako, C., Shinohara, R. T., Bergquist, T., Chai, R., Eddy, J., Elliott, J., Reade, W., Schaffter, T., Yu, T., Zheng, J., Moawad, A. W., Coelho, L. O., McDonnell, O., Miller, E., Moron, F....
-
[64]
Algorithms for hyper-parameter optimization
Bergstra, J., Bardenet, R., Bengio, Y., and K\' e gl, B. Algorithms for hyper-parameter optimization. In Shawe-Taylor, J., Zemel, R., Bartlett, P., Pereira, F., and Weinberger, K. (eds.), Advances in Neural Information Processing Systems, volume 24. Curran Associates, Inc., 2011. URL https://proceedings.neurips.cc/paper_files/paper/2011/file/86e8f7ab32cfd...
work page 2011
-
[65]
Deep extremely randomized trees
Berrouachedi, A., Jaziri, R., and Bernard, G. Deep extremely randomized trees. In International Conference on Neural Information Processing, pp.\ 717--729. Springer, 2019
work page 2019
-
[66]
B., Vorontsov, E., Ben-Cohen, A., Kaissis, G., Szeskin, A., Jacobs, C., Mamani, G
Bilic, P., Christ, P., Li, H. B., Vorontsov, E., Ben-Cohen, A., Kaissis, G., Szeskin, A., Jacobs, C., Mamani, G. E. H., Chartrand, G., Lohöfer, F., Holch, J. W., Sommer, W., Hofmann, F., Hostettler, A., Lev-Cohain, N., Drozdzal, M., Amitai, M. M., Vivanti, R., Sosna, J., Ezhov, I., Sekuboyina, A., Navarro, F., Kofler, F., Paetzold, J. C., Shit, S., Hu, X....
-
[67]
Bonato, B., Nanni, L., and Bertoldo, A. Advancing precision: A comprehensive review of mri segmentation datasets from brats challenges (2012–2025). Sensors, 25 0 (6), 2025. ISSN 1424-8220. doi:10.3390/s25061838. URL https://www.mdpi.com/1424-8220/25/6/1838
-
[68]
Brussee, S., Buzzanca, G., Schrader, A. M., and Kers, J. Graph neural networks in histopathology: Emerging trends and future directions. Medical Image Analysis, 101: 0 103444, 2025. ISSN 1361-8415. doi:https://doi.org/10.1016/j.media.2024.103444. URL https://www.sciencedirect.com/science/article/pii/S1361841524003694
-
[69]
Cheng, G., Han, J., Zhou, P., and Guo, L. Multi-class geospatial object detection and geographic image classification based on collection of part detectors. ISPRS Journal of Photogrammetry and Remote Sensing, 98: 0 119--132, 2014. ISSN 0924-2716. doi:https://doi.org/10.1016/j.isprsjprs.2014.10.002. URL https://www.sciencedirect.com/science/article/pii/S09...
-
[70]
Demaine, E. D., Hajiaghayi, M. T., and Kawarabayashi, K.-i. Algorithmic graph minor theory: Decomposition, approximation, and coloring. In 46th Annual IEEE Symposium on Foundations of Computer Science (FOCS'05), pp.\ 637--646. IEEE, 2005
work page 2005
-
[71]
Ding, K., Zhou, M., Wang, H., Zhang, S., and Metaxas, D. N. Spatially aware graph neural networks and cross-level molecular profile prediction in colon cancer histopathology: a retrospective multi-cohort study. The Lancet Digital Health, 4 0 (11): 0 e787--e795, 2022
work page 2022
-
[72]
El Badaoui, R., Bonmati Coll, E., Psarrou, A., Asaturyan, H. A., and Villarini, B. Enhanced catbrats for brain tumour semantic segmentation. Journal of Imaging, 11 0 (1), 2025. ISSN 2313-433X. doi:10.3390/jimaging11010008. URL https://www.mdpi.com/2313-433X/11/1/8
-
[73]
Felzenszwalb, P. F. and Huttenlocher, D. P. Efficient graph-based image segmentation. International journal of computer vision, 59 0 (2): 0 167--181, 2004
work page 2004
-
[74]
Medical image segmentation: A comprehensive review of deep learning-based methods
Gao, Y., Jiang, Y., Peng, Y., Yuan, F., Zhang, X., and Wang, J. Medical image segmentation: A comprehensive review of deep learning-based methods. Tomography, 11 0 (5), 2025. ISSN 2379-139X. doi:10.3390/tomography11050052. URL https://www.mdpi.com/2379-139X/11/5/52
-
[75]
Ghankot, R. S., Singh, M., Desroches, S. T., Jester, N., Mahajan, A., Lorr, S., Buono, F. D., Wiznia, D. H., Johnson, M. H., and Tommasini, S. M. Evaluating the effect of voxel size on the accuracy of 3d volumetric analysis measurements of brain tumors. Frontiers in Radiology, 5: 0 1618261, 2025
work page 2025
-
[76]
Instance-level semantic segmentation of nuclei based on multimodal structure encoding
Guan, B., Chu, G., Wang, Z., Li, J., and Yi, B. Instance-level semantic segmentation of nuclei based on multimodal structure encoding. BMC bioinformatics, 26 0 (1): 0 42, 2025
work page 2025
-
[77]
Heller, N., Isensee, F., Trofimova, D., Tejpaul, R., Zhao, Z., Chen, H., Wang, L., Golts, A., Khapun, D., Shats, D., Shoshan, Y., Gilboa-Solomon, F., George, Y., Yang, X., Zhang, J., Zhang, J., Xia, Y., Wu, M., Liu, Z., Walczak, E., McSweeney, S., Vasdev, R., Hornung, C., Solaiman, R., Schoephoerster, J., Abernathy, B., Wu, D., Abdulkadir, S., Byun, B., S...
- [78]
-
[79]
Strategies for pre-training graph neural networks
Hu*, W., Liu*, B., Gomes, J., Zitnik, M., Liang, P., Pande, V., and Leskovec, J. Strategies for pre-training graph neural networks. In International Conference on Learning Representations, 2020. URL https://openreview.net/forum?id=HJlWWJSFDH
work page 2020
-
[80]
Jiangtao, W., Ruhaiyem, N. I. R., and Panpan, F. A comprehensive review of u-net and its variants: Advances and applications in medical image segmentation. IET Image Processing, 19 0 (1): 0 e70019, 2025. doi:https://doi.org/10.1049/ipr2.70019. URL https://ietresearch.onlinelibrary.wiley.com/doi/abs/10.1049/ipr2.70019
-
[81]
Kaczmarska, M. and Majek, K. 3d segmentation of kidneys, kidney tumors and cysts on ct images - kits23 challenge. In Heller, N., Wood, A., Isensee, F., R \"a dsch, T., Teipaul, R., Papanikolopoulos, N., and Weight, C. (eds.), Kidney and Kidney Tumor Segmentation, pp.\ 149--155, Cham, 2024. Springer Nature Switzerland. ISBN 978-3-031-54806-2
work page 2024
-
[82]
Kingma, D. P. and Ba, J. Adam: A method for stochastic optimization, 2017. URL https://arxiv.org/abs/1412.6980
work page internal anchor Pith review Pith/arXiv arXiv 2017
-
[83]
Kostrykin, L. and Rohr, K. Superadditivity and convex optimization for globally optimal cell segmentation using deformable shape models. IEEE Transactions on Pattern Analysis and Machine Intelligence, 45 0 (3): 0 3831--3847, 2022
work page 2022
-
[84]
Lei, T., Wang, R., Zhang, Y., Wan, Y., Liu, C., and Nandi, A. K. Defed-net: Deformable encoder-decoder network for liver and liver tumor segmentation. IEEE Transactions on Radiation and Plasma Medical Sciences, 6 0 (1): 0 68--78, 2022. doi:10.1109/TRPMS.2021.3059780
-
[85]
H-denseunet: Hybrid densely connected unet for liver and tumor segmentation from ct volumes
Li, X., Chen, H., Qi, X., Dou, Q., Fu, C.-W., and Heng, P.-A. H-denseunet: Hybrid densely connected unet for liver and tumor segmentation from ct volumes. IEEE Transactions on Medical Imaging, 37 0 (12): 0 2663--2674, 2018. doi:10.1109/TMI.2018.2845918
-
[86]
Liao, J., Wang, H., Gu, H., and Cai, Y. Liver tumor segmentation method combining multi-axis attention and conditional generative adversarial networks. PLOS ONE, 19 0 (12): 0 1--24, 12 2024. doi:10.1371/journal.pone.0312105. URL https://doi.org/10.1371/journal.pone.0312105
-
[87]
Lilhore, U. K., Sunder, R., Simaiya, S., Alsafyani, M., Monish Khan, M., Alroobaea, R., Alsufyani, H., and Baqasah, A. M. Ag-ms3d-cnn multiscale attention guided 3d convolutional neural network for robust brain tumor segmentation across mri protocols. Scientific Reports, 15 0 (1): 0 24306, 2025
work page 2025
-
[88]
Superpixel region merging based on deep network for medical image segmentation
Liu, H., Wang, H., Wu, Y., and Xing, L. Superpixel region merging based on deep network for medical image segmentation. ACM Trans. Intell. Syst. Technol., 11 0 (4), May 2020. ISSN 2157-6904. doi:10.1145/3386090. URL https://doi.org/10.1145/3386090
-
[89]
Limt: A multi-task liver image benchmark dataset
Liu, Z., Han, K., Ma, S., Zhu, Y., Chen, J., Lyu, C., Qiu, X., Qian, C., Song, Y., Liu, Y., et al. Limt: A multi-task liver image benchmark dataset. arXiv preprint arXiv:2511.19889, 2025
-
[90]
Lov \'a sz, L. Graph minor theory. Bulletin of the American Mathematical Society, 43 0 (1): 0 75--86, 2006
work page 2006
-
[91]
Rethinking u-net: Task-adaptive mixture of skip connections for enhanced medical image segmentation
Luo, Z., Zhu, X., Zhang, L., and Sun, B. Rethinking u-net: Task-adaptive mixture of skip connections for enhanced medical image segmentation. Proceedings of the AAAI Conference on Artificial Intelligence, 39 0 (6): 0 5874--5882, Apr. 2025. doi:10.1609/aaai.v39i6.32627. URL https://ojs.aaai.org/index.php/AAAI/article/view/32627
-
[92]
Mienye, I. D. and Viriri, S. Graph neural networks in medical imaging: Methods, applications and future directions. Information, 16 0 (12), 2025. ISSN 2078-2489. doi:10.3390/info16121051. URL https://www.mdpi.com/2078-2489/16/12/1051
-
[93]
Automated 3d segmentation of kidneys and tumors in miccai kits 2023 challenge
Myronenko, A., Yang, D., He, Y., and Xu, D. Automated 3d segmentation of kidneys and tumors in miccai kits 2023 challenge. In Heller, N., Wood, A., Isensee, F., R \"a dsch, T., Teipaul, R., Papanikolopoulos, N., and Weight, C. (eds.), Kidney and Kidney Tumor Segmentation, pp.\ 1--7, Cham, 2024. Springer Nature Switzerland. ISBN 978-3-031-54806-2
work page 2023
-
[94]
Neubert, P. and Protzel, P. Compact watershed and preemptive slic: On improving trade-offs of superpixel segmentation algorithms. In 2014 22nd International Conference on Pattern Recognition, pp.\ 996--1001, 2014. doi:10.1109/ICPR.2014.181
-
[95]
Nowakowski, L. and Patel, R. Convolutional occupancy networks for medical imaging with applications to the kits23 challenge. In IEEE-EMBS International Conference on Biomedical and Health Informatics 2025, 2025
work page 2025
-
[96]
Pandey, S., Toshali, Perslev, M., and Dam, E. B. Advancing kidney, kidney tumor, cyst segmentation: A multi-planner u-net approach for the kits23 challenge. In Heller, N., Wood, A., Isensee, F., R \"a dsch, T., Teipaul, R., Papanikolopoulos, N., and Weight, C. (eds.), Kidney and Kidney Tumor Segmentation, pp.\ 143--148, Cham, 2024. Springer Nature Switzer...
work page 2024
-
[97]
Peng, Y., Chen, D. Z., and Sonka, M. U-net v2: Rethinking the skip connections of u-net for medical image segmentation. In 2025 IEEE 22nd International Symposium on Biomedical Imaging (ISBI), pp.\ 1--5, 2025. doi:10.1109/ISBI60581.2025.10980742
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.