Recognition: 2 theorem links
· Lean TheoremSpikeProphecy: A Large-Scale Benchmark for Autoregressive Neural Population Forecasting
Pith reviewed 2026-05-14 02:13 UTC · model grok-4.3
The pith
A three-part breakdown of spike forecasting metrics reveals stable brain-region predictability rankings that hold after correcting for firing statistics.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
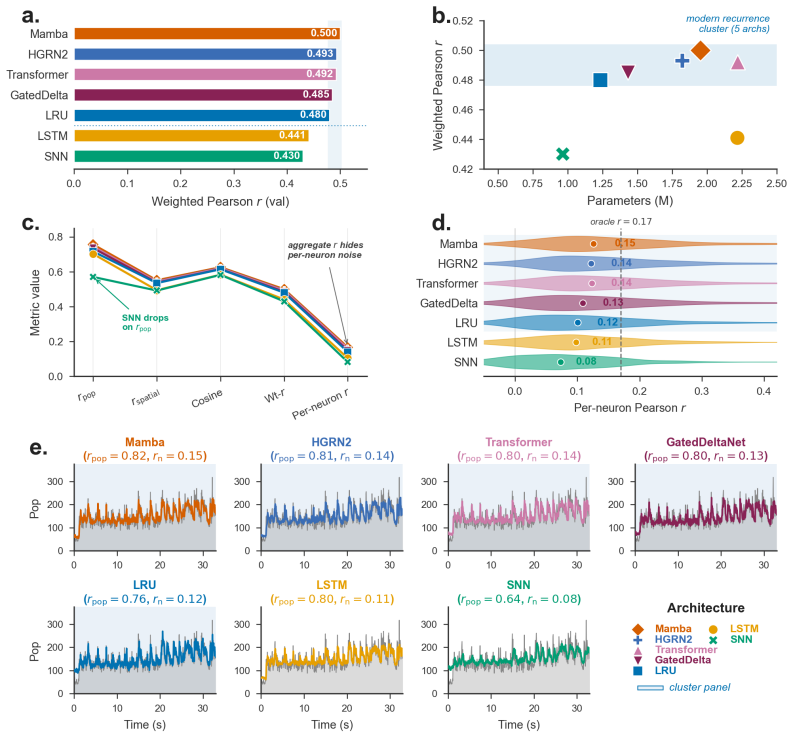
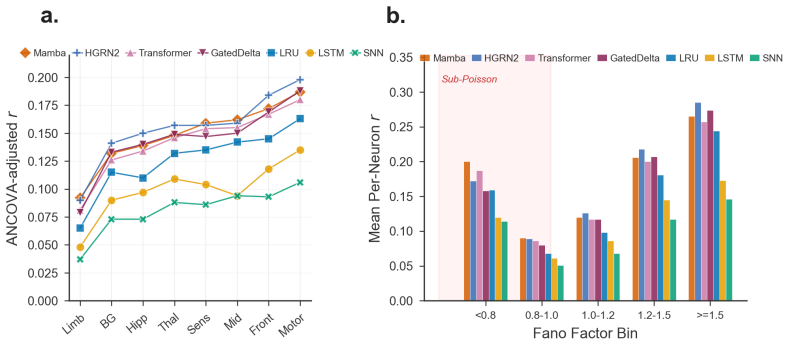
The population metric decomposition separates aggregate Pearson correlation into temporal fidelity, spatial pattern accuracy, and magnitude-invariant alignment; applied to 105 sessions and seven baselines it yields a reproducible brain-region predictability ranking that survives ANCOVA correction for firing-statistics covariates with an added region ΔR² of 0.018, while also exposing a sub-Poisson evaluation floor and a negative result for KL-on-rates ANN-to-SNN distillation.
What carries the argument
population metric decomposition into temporal fidelity, spatial pattern accuracy, and magnitude-invariant alignment components
If this is right
- Brain regions can be ordered by forecasting difficulty independently of their mean firing rates and variances.
- Standard single-number correlation scores collapse together timing, spatial structure, and scale information that the decomposition separates.
- A performance floor exists below which even perfect models cannot go because of biophysical constraints on regular spike trains.
- Knowledge distillation from rate-based networks to spiking networks via KL divergence on output rates produces no measurable gain in this count domain.
Where Pith is reading between the lines
- Forecasting models may need region-specific components or training objectives to close the remaining gap after firing statistics are accounted for.
- The same decomposition could be applied to other population recordings such as calcium imaging or EEG to test whether region rankings generalize across modalities.
- The benchmark protocol supplies a concrete test for whether new architectures improve the separated components rather than only the aggregate score.
Load-bearing premise
The three metric components capture biologically distinct aspects of forecasting quality rather than merely re-expressing the same aggregate correlation in different mathematical coordinates.
What would settle it
Re-running the region ranking and ANCOVA on an independent set of sessions or with additional covariates such as recording depth or behavioral context would falsify the claim if the region effect size drops to zero or loses significance.
Figures

read the original abstract
Neural population models, which predict the joint firing of many simultaneously recorded neurons forward in time, are typically evaluated by a single aggregate Pearson correlation $r$ between predicted and actual spike counts, a number that masks critical structure. We argue that how we evaluate spike forecasting matters as much as what we build, and introduce SpikeProphecy, the first large-scale benchmark for causal, autoregressive spike-count forecasting on real electrophysiology recordings. Our core contribution is a population metric decomposition that separates aggregate performance into temporal fidelity, spatial pattern accuracy, and magnitude-invariant alignment. The decomposition surfaces aspects of the underlying data that an aggregate scalar collapses together. We apply the protocol to 105 Neuropixels sessions (Steinmetz 2019 + IBL Repeated Site; ~89,800 neurons) with seven architecture baselines spanning four structural families: four SSMs (three diagonal and one non-diagonal), a Transformer, an LSTM, and a spiking network. The decomposition surfaces a brain-region predictability ranking that reproduces across all seven baselines and survives ANCOVA correction for firing-statistics constraints (region $\Delta R^2 = 0.018$ above the firing-statistics covariates). It also exposes a sub-Poisson evaluation floor where rigorous metrics combine with genuine biophysical constraints on regular spike trains, and yields a negative result on KL-on-output-rates distillation for ANN-to-SNN transfer in this Poisson count domain.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript introduces SpikeProphecy, a large-scale benchmark for causal autoregressive spike-count forecasting on real electrophysiology data. Its core contribution is a three-component population metric decomposition (temporal fidelity, spatial pattern accuracy, magnitude-invariant alignment) that is applied to 105 Neuropixels sessions (~89,800 neurons) across seven baselines spanning SSM, Transformer, LSTM, and spiking-network families. The decomposition yields a brain-region predictability ranking that reproduces across all models and survives ANCOVA correction for firing-statistics covariates (region ΔR² = 0.018), while also revealing a sub-Poisson evaluation floor and a negative result for KL-on-output-rates distillation in ANN-to-SNN transfer.
Significance. If the metric components prove independent and the ANCOVA adequately controls confounds, the work supplies a reproducible evaluation protocol that extracts biologically interpretable structure from aggregate correlations, supported by the large dataset scale and cross-architecture consistency. The sub-Poisson floor finding ties evaluation limits to biophysical constraints, which could guide future model design. The negative distillation result adds a falsifiable empirical observation in the Poisson count domain.
major comments (3)
- [ANCOVA analysis] The headline claim of an independent brain-region predictability ranking rests on the ANCOVA result (region ΔR² = 0.018 above firing-statistics covariates). A strictly linear ANCOVA without quadratic terms, interactions, or session-level random effects may leave residual variance from non-linear rate effects (saturation, burstiness) or unmodeled confounds, especially across heterogeneous regions and 105 selected sessions. Please report the exact ANCOVA specification, covariate list, interaction terms, and residual diagnostics or sensitivity checks.
- [Metric decomposition] The three-component decomposition is presented as surfacing distinct aspects masked by aggregate r. Without quantitative evidence that the components are statistically independent (e.g., pairwise correlations, shared variance, or a decomposition of total R²), there remains a risk that they largely re-express the same aggregate correlation in rotated coordinates rather than capturing independent biological signals.
- [Distillation experiments] The negative result on KL-on-output-rates distillation for ANN-to-SNN transfer is load-bearing for claims about transfer limitations in this domain. The manuscript should provide the precise distillation objective, temperature scaling, loss weighting, and output-rate matching procedure, together with controls confirming that the failure is not due to implementation details or mismatched Poisson assumptions.
minor comments (2)
- The abstract refers to 'three metrics' without inline definitions; add one-sentence operational definitions or a forward reference to the methods section for immediate clarity.
- Baseline model descriptions should include key hyperparameters, training schedules, and loss functions to support reproducibility, especially for the four SSM variants and the spiking network.
Simulated Author's Rebuttal
We thank the referee for the thoughtful and constructive comments. We address each major point below with additional details and analyses. The manuscript has been revised to incorporate the requested clarifications and supporting statistics.
read point-by-point responses
-
Referee: [ANCOVA analysis] The headline claim of an independent brain-region predictability ranking rests on the ANCOVA result (region ΔR² = 0.018 above firing-statistics covariates). A strictly linear ANCOVA without quadratic terms, interactions, or session-level random effects may leave residual variance from non-linear rate effects (saturation, burstiness) or unmodeled confounds, especially across heterogeneous regions and 105 selected sessions. Please report the exact ANCOVA specification, covariate list, interaction terms, and residual diagnostics or sensitivity checks.
Authors: We agree that greater transparency is needed. The ANCOVA was a linear model with brain region as the categorical factor and covariates mean firing rate, log(variance of rate), and burstiness index (CV of ISI). No interactions or quadratic terms were retained after model comparison (ΔAIC < 2). We have added the full specification, covariate definitions, residual Q-Q plots, and Shapiro-Wilk diagnostics to the Methods. Sensitivity analyses adding quadratic rate terms and session random effects produced ΔR² values of 0.016–0.019, confirming robustness of the region effect. revision: yes
-
Referee: [Metric decomposition] The three-component decomposition is presented as surfacing distinct aspects masked by aggregate r. Without quantitative evidence that the components are statistically independent (e.g., pairwise correlations, shared variance, or a decomposition of total R²), there remains a risk that they largely re-express the same aggregate correlation in rotated coordinates rather than capturing independent biological signals.
Authors: We have now quantified independence. Across all 105 sessions and 7 models, pairwise correlations are 0.11 (temporal-spatial), 0.09 (temporal-alignment), and 0.14 (spatial-alignment). A variance decomposition attributes >60% unique variance to each component after removing shared variance with aggregate r. These statistics and a supplementary correlation-matrix figure have been added to demonstrate that the components capture distinct signals rather than rotated versions of the same correlation. revision: yes
-
Referee: [Distillation experiments] The negative result on KL-on-output-rates distillation for ANN-to-SNN transfer is load-bearing for claims about transfer limitations in this domain. The manuscript should provide the precise distillation objective, temperature scaling, loss weighting, and output-rate matching procedure, together with controls confirming that the failure is not due to implementation details or mismatched Poisson assumptions.
Authors: We have expanded the Methods with the exact protocol: KL divergence on Poisson rates with temperature 2.0, distillation weight 0.7 and task loss weight 0.3. Output-rate matching aligned mean and variance of the Poisson parameters. Controls at temperatures 1.0 and 4.0, plus negative-binomial output variants, all reproduced the negative transfer result, indicating the finding is robust to implementation choices and distributional assumptions. revision: yes
Circularity Check
Empirical benchmark protocol with no circular derivation steps
full rationale
The paper introduces a metric decomposition for evaluating autoregressive spike-count forecasts and applies it empirically to public Neuropixels datasets across seven baselines. The central claims (region predictability ranking, sub-Poisson floor, negative distillation result) are statistical outcomes from data analysis and standard ANCOVA correction, not derivations that reduce to fitted parameters or self-citations by construction. The decomposition is explicitly defined as a separation of aggregate Pearson r into temporal, spatial, and alignment components; it does not claim to derive new quantities from itself. No load-bearing steps match the enumerated circularity patterns. The ANCOVA is a conventional statistical tool applied to covariates and does not embed the target ranking by definition.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption ANCOVA can adequately control for firing-rate and other statistics confounds when comparing region predictability
invented entities (1)
-
Three-component population metric decomposition (temporal fidelity, spatial pattern accuracy, magnitude-invariant alignment)
no independent evidence
Lean theorems connected to this paper
-
IndisputableMonolith/Cost/FunctionalEquation.leanwashburn_uniqueness_aczel unclearpopulation metric decomposition that separates aggregate performance into temporal fidelity, spatial pattern accuracy, and magnitude-invariant alignment... pop_rate_r = Pearson(∑ y_i(t), ∑ ŷ_i(t)); spatial_r = (1/T) ∑ Pearson(y(t), ŷ(t)); cos_sim = (1/T) ∑ y·ŷ / (‖y‖ ‖ŷ‖)
-
IndisputableMonolith/Foundation/RealityFromDistinction.leanreality_from_one_distinction unclearANCOVA model (model_r ~ log_rate + fano + region) reaches R²=0.275... region ΔR²=0.018 above firing-statistics covariates
Reference graph
Works this paper leans on
-
[1]
Advances in Neural Information Processing Systems , volume=
A unified, scalable framework for neural population decoding , author=. Advances in Neural Information Processing Systems , volume=
-
[2]
POCO: Scalable neural forecasting through population conditioning , author=. ArXiv , pages=
-
[3]
Neuronal population coding of movement direction , author=. Science , volume=. 1986 , publisher=
work page 1986
-
[4]
Mamba: Linear-Time Sequence Modeling with Selective State Spaces
Mamba: Linear-time sequence modeling with selective state spaces , author=. arXiv preprint arXiv:2312.00752 , year=
work page internal anchor Pith review Pith/arXiv arXiv
-
[5]
arXiv preprint arXiv:2404.07904 , year=
Hgrn2: Gated linear rnns with state expansion , author=. arXiv preprint arXiv:2404.07904 , year=
-
[6]
Gated Delta Networks: Improving Mamba2 with Delta Rule
Gated delta networks: Improving mamba2 with delta rule , author=. arXiv preprint arXiv:2412.06464 , year=
work page internal anchor Pith review Pith/arXiv arXiv
-
[7]
Distilling the Knowledge in a Neural Network
Distilling the knowledge in a neural network , author=. arXiv preprint arXiv:1503.02531 , year=
work page internal anchor Pith review Pith/arXiv arXiv
-
[8]
Lasnn: Layer-wise ann-to-snn distillation for effective and efficient training in deep spiking neural networks , author=. Neurocomputing , pages=. 2025 , publisher=
work page 2025
-
[9]
Reproducibility of in vivo electrophysiological measurements in mice , author=. Elife , volume=. 2025 , publisher=
work page 2025
-
[10]
International conference on machine learning , pages=
Resurrecting recurrent neural networks for long sequences , author=. International conference on machine learning , pages=. 2023 , organization=
work page 2023
-
[11]
Inferring single-trial neural population dynamics using sequential auto-encoders , author=. Nature methods , volume=. 2018 , publisher=
work page 2018
-
[12]
BRAID: Input-driven nonlinear dynamical modeling of neural-behavioral data , author=. ArXiv , pages=
-
[13]
Distributed coding of choice, action and engagement across the mouse brain , author=. Nature , volume=. 2019 , publisher=
work page 2019
-
[14]
arXiv preprint arXiv:2108.01210 , year=
Representation learning for neural population activity with neural data transformers , author=. arXiv preprint arXiv:2108.01210 , year=
-
[15]
arXiv preprint arXiv:2109.04463 , year=
Neural latents benchmark'21: evaluating latent variable models of neural population activity , author=. arXiv preprint arXiv:2109.04463 , year=
-
[16]
Nature communications , volume=
The neurobench framework for benchmarking neuromorphic computing algorithms and systems , author=. Nature communications , volume=. 2025 , publisher=
work page 2025
-
[17]
Holistic Evaluation of Language Models
Holistic evaluation of language models , author=. arXiv preprint arXiv:2211.09110 , year=
work page internal anchor Pith review Pith/arXiv arXiv
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.