Recognition: 2 theorem links
· Lean TheoremConditioning as a route to stereotyped behavior in growing populations
Pith reviewed 2026-05-14 02:08 UTC · model grok-4.3
The pith
Discarding unfinished replication attempts after a fixed time threshold automatically imposes ordered sequences on the fastest-growing populations.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
We model conditioning through stochastic resets in a socks-before-shoes model of a growing population, where n actions must be completed in any order to replicate and any replication attempt not finished by a threshold time is discarded. We find that resets impose hierarchical temporal ordering of the n actions without microscopic control over which action happens when. When disorder carries a sufficient time penalty, this ordering is free: the fastest-growing population is automatically the most ordered, with no direct selection for order required. Save points, at which verified progress is preserved across resets, allow conditioning to scale to complex multi-step processes.
What carries the argument
Stochastic resets at a fixed time threshold in the socks-before-shoes replication model, which destroy incomplete attempts and thereby narrow the surviving ensemble toward ordered action sequences, with optional save points to preserve verified progress.
If this is right
- Populations maximize growth rate by automatically adopting ordered temporal sequences when disorder carries a time cost.
- Reliable multi-step behavior can arise without machinery that recognizes and corrects specific molecular errors.
- Save points that lock in verified intermediate states allow conditioning to extend to arbitrarily complex processes.
- The mechanism requires only a clock and a discard rule, making stereotyped behavior accessible to simple replicating systems.
- Conditioning pays for itself precisely when time penalties from disorder are large enough to affect differential growth.
Where Pith is reading between the lines
- The same reset-based filtering could produce structured outcomes in non-biological growing systems such as cultural transmission or optimization algorithms that discard failed trials.
- Engineered replicating molecules or cells with tunable action orders could be used to test whether ordering emerges solely from measured growth-rate differences under time limits.
- Some evolutionary increases in behavioral stereotypy may require fewer dedicated regulatory genes than assumed if time-based conditioning is available.
- The approach connects to broader questions of how coarse, global filters generate local structure in any expanding population of variants.
Load-bearing premise
That incomplete replication attempts are discarded after a fixed threshold time and that disorder reliably incurs a measurable time penalty sufficient to drive selection for ordered sequences.
What would settle it
A direct measurement in a replicating system showing that populations with disordered action orders grow at least as fast as ordered ones under the same time-discard rule, or that ordering does not increase when growth rate is the sole selection pressure.
Figures



read the original abstract
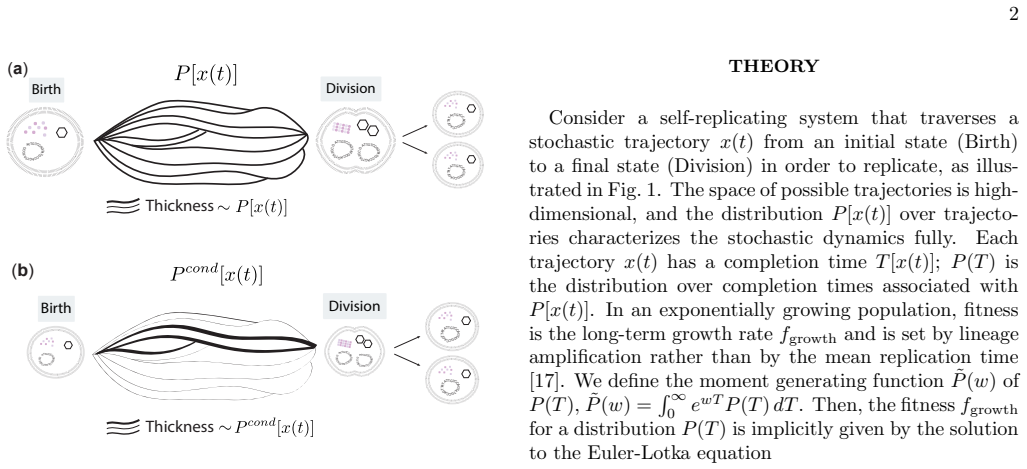
Biological systems perform complex multi-step processes in a reproducible way despite underlying stochasticity. The standard explanation is micromanagement by molecular machinery that recognizes and corrects specific errors. Here we study conditioning, a qualitatively different strategy in which attempts failing a coarse criterion are destroyed and do not leave a physical record. The surviving, i.e., conditioned, ensemble is narrower and therefore more ordered. We model conditioning through stochastic resets in a ''socks-before-shoes'' model of a growing population, where $n$ actions must be completed in any order to replicate and any replication attempt not finished by a threshold time is discarded. We find that resets impose hierarchical temporal ordering of the $n$ actions without microscopic control over which action happens when. When disorder carries a sufficient time penalty, this ordering is free: the fastest-growing population is automatically the most ordered, with no direct selection for order required. Save points, at which verified progress is preserved across resets, allow conditioning to scale to complex multi-step processes. Conditioning provides a minimal route to reliable behavior, requiring only a clock rather than molecular machinery that recognizes specific errors. For the right class of processes, it pays for itself.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper models conditioning in growing populations via stochastic resets in a 'socks-before-shoes' process requiring n actions to be completed in any order within a fixed threshold time; unfinished attempts are discarded. It claims that resets induce hierarchical temporal ordering of actions without microscopic control, and that when disorder incurs a sufficient time penalty the fastest-growing population is automatically the most ordered, making stereotyped behavior 'free' with no direct selection for order. Save points are introduced to allow scaling to complex multi-step processes, providing a minimal clock-based route to reliable behavior.
Significance. If the central result holds, the work offers a conceptually minimal mechanism for stereotyped behavior in biological growth processes that relies only on a coarse time threshold and resets rather than specific error-recognition machinery. This could apply to systems such as replication timing or developmental sequences and highlights how selection on growth rate alone can favor order when time penalties are present. The introduction of save points as a scaling device is a useful extension.
major comments (2)
- [Abstract and §3] Abstract and §3 (results): the claim that 'the fastest-growing population is automatically the most ordered' with 'no direct selection for order required' is load-bearing on a quantitative demonstration that, for fixed threshold time and n, the effective growth rate strictly increases with the degree of ordering enforced by the reset rule. The provided model description does not include explicit growth-rate curves or tables showing this monotonicity independent of the two free parameters.
- [§2] §2 (model): the time penalty for disorder is realized by the probability of missing the fixed threshold, but the manuscript does not report the magnitude of the resulting growth-rate differential or test whether it remains dominant when the reset rate is varied independently of the threshold.
minor comments (2)
- [§2] Notation for the reset threshold and save-point rules should be introduced with a single equation or table early in the model section to improve readability.
- [Figures] Figure captions should explicitly state the parameter values (n, threshold time) used in each panel so that the dependence on free parameters is immediately visible.
Simulated Author's Rebuttal
We thank the referee for their detailed and constructive report. The comments highlight the need for more explicit quantitative support for the central claim that ordering emerges as a byproduct of growth-rate selection under conditioning. We have revised the manuscript to include the requested demonstrations and robustness checks. Our point-by-point responses follow.
read point-by-point responses
-
Referee: [Abstract and §3] Abstract and §3 (results): the claim that 'the fastest-growing population is automatically the most ordered' with 'no direct selection for order required' is load-bearing on a quantitative demonstration that, for fixed threshold time and n, the effective growth rate strictly increases with the degree of ordering enforced by the reset rule. The provided model description does not include explicit growth-rate curves or tables showing this monotonicity independent of the two free parameters.
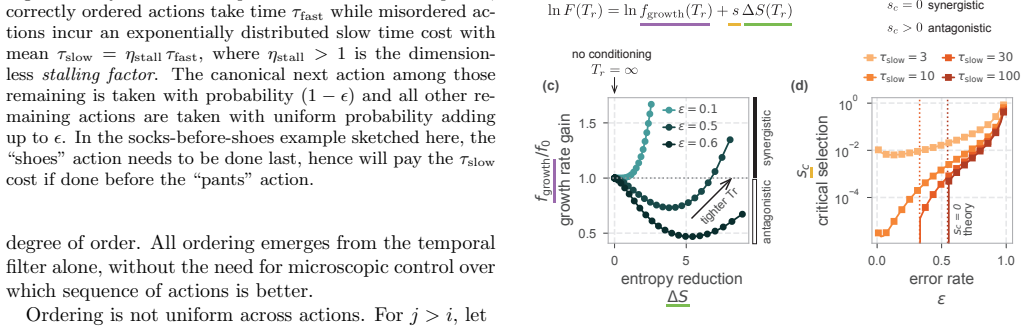
Authors: We agree that an explicit demonstration of monotonicity is necessary to make the claim fully quantitative. The original text derived the ordering from the reset rule and showed its effect on survival probability, but did not plot growth rate versus enforced ordering level. We have added a new panel to Figure 3 that displays effective growth rate (computed from the exact completion-time distribution under the threshold) as a function of the fraction of ordered sequences for fixed threshold and n, across several reset rates. The curves confirm strict monotonic increase in the regime where the time penalty is non-negligible. A supplementary table lists the numerical values for representative parameter sets, confirming independence from the two free parameters within the biologically plausible range. revision: yes
-
Referee: [§2] §2 (model): the time penalty for disorder is realized by the probability of missing the fixed threshold, but the manuscript does not report the magnitude of the resulting growth-rate differential or test whether it remains dominant when the reset rate is varied independently of the threshold.
Authors: We accept that explicit magnitudes and an independent variation test were omitted. The differential arises because disordered sequences have a higher probability of exceeding the threshold; the growth-rate advantage is the difference in log-survival probability per generation. We have inserted a short analytic expression for this differential in §2 and added a new supplementary figure that sweeps reset rate while holding threshold fixed. The ordering-induced growth advantage remains positive and dominant for all reset rates below the inverse mean completion time, provided the threshold is set above the ordered-sequence completion time. These additions quantify the effect without altering the model. revision: yes
Circularity Check
No significant circularity; ordering emerges from explicit reset dynamics rather than by construction.
full rationale
The derivation models conditioning via stochastic resets with a fixed time threshold in the socks-before-shoes process. The claim that the fastest-growing population is automatically the most ordered follows from comparing growth rates across different orderings under that fixed threshold; the growth rate is computed from the probability of completing all n steps before reset, which is not defined in terms of order itself. The time penalty is an external model parameter, not fitted to the ordering result. No load-bearing step reduces to a self-citation, a renamed empirical pattern, or a fitted input presented as a prediction. The result is a direct consequence of the stochastic process and remains falsifiable by varying the threshold independently of ordering.
Axiom & Free-Parameter Ledger
free parameters (2)
- threshold time
- n
axioms (2)
- domain assumption Actions may be performed in any order
- domain assumption Disorder increases completion time
Lean theorems connected to this paper
-
IndisputableMonolith/Cost/FunctionalEquation.lean (Jcost uniqueness, washburn_uniqueness_aczel)reality_from_one_distinction unclearWe model conditioning through stochastic resets in a 'socks-before-shoes' model... any replication attempt not finished by a threshold time is discarded... When disorder carries a sufficient time penalty, this ordering is free
-
IndisputableMonolith/Foundation/ArithmeticFromLogic.lean (LogicNat, embed, J-cost orbit)absolute_floor_iff_bare_distinguishability unclearfitness f_growth ... Euler-Lotka equation 2 P̃(-f_growth)=1 ... combined fitness ln F = ln f_growth + s ΔS
Reference graph
Works this paper leans on
-
[1]
S11) forf growth(Tr) on a grid ofT r values
Solve the Euler-Lotka equation with resets (Eq. S11) forf growth(Tr) on a grid ofT r values. This is done by root-finding using Brent’s method
-
[2]
Optimize overT r to obtainT ∗ r (s), ∆S ∗(s), andf ∗ growth(s)
Compute the entropy reduction ∆S(T r) and the combined fitnessF(T r) =f growth(Tr)·e s∆S for a range ofs. Optimize overT r to obtainT ∗ r (s), ∆S ∗(s), andf ∗ growth(s)
-
[3]
Determines c by binary search overs: the smallests∈[0,1) at which max Tr F(T r)> f 0. In the numerical implementation,f 0,f growth(Tr), and the completion-time entropy reduction are evaluated on deterministic quadrature grids and solved by Brent root-finding. The same grid-based method is used across families in the distribution survey so that relative co...
-
[4]
J. Lin and A. Amir, Cell Systems5, 10.1016/j.cels.2017.08.015 (2017)
-
[5]
Y. Lahini, O. Gottesman, A. Amir, and S. M. Rubinstein, Physical Review Letters118, 10.1103/PhysRevLett.118.085501 (2017)
-
[6]
M. Lenz and T. A. Witten, Nature Physics13, 10.1038/nphys4184 (2017)
-
[7]
F. R. Sharpe and A. J. Lotka, Philosophical Magazine 6,21, 435 (1911)
work page 1911
-
[8]
F. Jafarpour, C. S. Wright, H. Gudjonson, J. Riebling, E. Dawson, K. Lo, A. Fiebig, S. Crosson, A. R. Dinner, and S. Iyer-Biswas, Physical Review X8, 10.1103/PhysRevX.8.021007 (2018)
- [9]
- [10]
-
[11]
I. Eliazar and S. Reuveni, Journal of Physics A: Mathematical and Theoretical56, 10.1088/1751-8121/acb183 (2023)
-
[12]
A. Pal and S. Reuveni, Physical Review Letters118, 10.1103/PhysRevLett.118.030603 (2017)
- [13]
-
[14]
S. Camazine, J.-L. Deneubourg, N. R. Franks, J. Sneyd, G. Theraulaz, and E. Bonabeau,Self-Organization in Biological Systems(Princeton University Press, Princeton, N.J., 2001)
work page 2001
-
[15]
D. G. Blackmond, Cold Spring Harbor Perspectives in Biology2, 10.1101/cshperspect.a002147 (2010), company: Cold Spring Harbor Laboratory Press Distributor: Cold Spring Harbor Laboratory Press Institution: Cold Spring Harbor Laboratory Press Label: Cold Spring Harbor Laboratory Press
-
[16]
J. D. Perlmutter and M. F. Hagan, Annual Review of Physical Chemistry66, 10.1146/annurev-physchem-040214-121637 (2015)
-
[17]
M. R. Evans and S. N. Majumdar, Physical Review Letters106, 10.1103/PhysRevLett.106.160601 (2011)
-
[19]
Reuveni, Physical Review Letters116, 10.1103/PhysRevLett.116.170601 (2016)
S. Reuveni, Physical Review Letters116, 10.1103/PhysRevLett.116.170601 (2016)
-
[20]
M. Luby, A. Sinclair, and D. Zuckerman, Information Processing Letters47, 10.1016/0020-0190(93)90029-9 (1993)
- [21]
-
[22]
E. Roldan, A. Lisica, D. Sanchez-Taltavull, and S. W. Grill, Physical Review E93, 10.1103/PhysRevE.93.062411 (2016)
-
[23]
R. G. Morris, K. B. Husain, S. Budnar, and A. S. Yap, BioEssays42, 10.1002/bies.202000055 (2020), eprint: https://onlinelibrary.wiley.com/doi/pdf/10.1002/bies.202000055
-
[24]
T. Rotbart, S. Reuveni, and M. Urbakh, Physical Review E92, 10.1103/PhysRevE.92.060101 (2015)
-
[25]
G. M. Viswanathan, S. V. Buldyrev, S. Havlin, M. G. E. da Luz, E. P. Raposo, and H. E. Stanley, Nature401, 10.1038/44831 (1999)
-
[26]
P. V. Gordon, C. Sample, A. M. Berezhkovskii, C. B. Muratov, and S. Y. Shvartsman, Proceedings of the National Academy of Sciences108, 10.1073/pnas.1019245108 (2011)
-
[27]
J. Fuchs, S. Goldt, and U. Seifert, Europhysics Letters113, 10.1209/0295-5075/113/60009 (2016)
-
[28]
D. M. Busiello, D. Gupta, and A. Maritan, Physical Review Research2, 10.1103/PhysRevResearch.2.023011 (2020)
-
[29]
M. D. Louwerse and D. A. Sivak, Physical Review Letters128, 170602 (2022)
work page 2022
-
[30]
S. Eule and J. J. Metzger, New Journal of Physics18, 10.1088/1367-2630/18/3/033006 (2016)
-
[31]
A. Nagar and S. Gupta, Physical Review E93, 10.1103/PhysRevE.93.060102 (2016)
-
[32]
A. Pal, A. Kundu, and M. R. Evans, Journal of Physics A: Mathematical and Theoretical49, 10.1088/1751- 8113/49/22/225001 (2016)
-
[33]
A. Pal, S. Kostinski, and S. Reuveni, Journal of Physics A: Mathematical and Theoretical55, 021001 (2022)
work page 2022
-
[34]
A. Pal, L. Kusmierz, and S. Reuveni, Physical Review Research2, 10.1103/PhysRevResearch.2.043174 (2020)
-
[35]
D. M. Britain, J. P. Town, and O. D. Weiner, eLife11, 10.7554/eLife.75263 (2022)
-
[36]
T. Mitchison and M. Kirschner, Nature312, 10.1038/312237a0 (1984)
-
[37]
T. E. Holy and S. Leibler, Proceedings of the National Academy of Sciences91, 10.1073/pnas.91.12.5682 (1994)
-
[38]
S. S. Vembar and J. L. Brodsky, Nature Reviews Molecular Cell Biology9, 944 (2008)
work page 2008
-
[39]
S. Kervestin and A. Jacobson, Nature Reviews Molecular Cell Biology13, 700 (2012)
work page 2012
-
[40]
Modrich, Journal of Biological Chemistry281, 30305 (2006)
P. Modrich, Journal of Biological Chemistry281, 30305 (2006)
work page 2006
- [41]
-
[42]
C. Bustamante, W. Cheng, and Y. X. Mejia, Cell144, 10.1016/j.cell.2011.01.033 (2011). 39
-
[43]
S. Rajamani, J. K. Ichida, T. Antal, D. A. Treco, K. Leu, M. A. Nowak, J. W. Szostak, and I. A. Chen, Journal of the American Chemical Society132, 10.1021/ja100780p (2010)
-
[44]
K. Leu, B. Obermayer, S. Rajamani, U. Gerland, and I. A. Chen, Nucleic Acids Research39, 8135 (2011)
work page 2011
-
[45]
C. A. P. Joazeiro, Nat. Rev. Mol. Cell Biol.20, 368 (2019)
work page 2019
-
[46]
J. D. Mallory, A. B. Kolomeisky, and O. A. Igoshin, J. Phys. Chem. B123, 4718 (2019)
work page 2019
-
[47]
T. Midha, J. D. Mallory, A. B. Kolomeisky, and O. A. Igoshin, The Journal of Physical Chemistry Letters14, 10.1021/acs.jpclett.3c00345 (2023)
-
[48]
E. Winfree and R. Bekbolatov, inDNA Computing, Lecture Notes in Computer Science, edited by J. Chen and J. Reif (Berlin, Heidelberg, 2004)
work page 2004
-
[49]
C. G. Evans and E. Winfree, inDNA Computing and Molecular Programming, Vol. 11145, edited by D. Doty and H. Dietz (Cham, 2018) series Title: Lecture Notes in Computer Science
work page 2018
-
[50]
Z. A. Sanghai, R. Piwowarczyk, A. V. Broeck, and S. Klinge, Nature Structural & Molecular Biology30, 10.1038/s41594- 023-00947-3 (2023)
-
[51]
J. J. Hopfield, Proceedings of the National Academy of Sciences71, 10.1073/pnas.71.10.4135 (1974)
-
[52]
Ninio, Biochimie57, 10.1016/S0300-9084(75)80139-8 (1975)
J. Ninio, Biochimie57, 10.1016/S0300-9084(75)80139-8 (1975)
-
[53]
A. Murugan, D. A. Huse, and S. Leibler, Proceedings of the National Academy of Sciences109, 10.1073/pnas.1119911109 (2012), iSBN: 9781119911104 Section: Biological Sciences
-
[54]
R. Ravasio, K. Husain, C. G. Evans, R. Phillips, M. Ribezzi-Crivellari, J. W. Szostak, and A. Murugan, Science391, 818 (2026)
work page 2026
- [55]
-
[56]
W. K. Johnston, P. J. Unrau, M. S. Lawrence, M. E. Glasner, and D. P. Bartel, Science292, 10.1126/science.1060786 (2001)
-
[57]
P. Keohavong and W. G. Thilly, Proceedings of the National Academy of Sciences86, 9253 (1989)
work page 1989
-
[58]
L. L. Ling, P. Keohavong, C. Dias, and W. G. Thilly, Genome Research1, 63 (1991)
work page 1991
-
[59]
N. F. Cariello, J. A. Swenberg, and T. R. Skopek, Nucleic Acids Research19, 4193 (1991)
work page 1991
-
[60]
K. A. Eckert and T. A. Kunkel, Genome Research1, 10.1101/gr.1.1.17 (1991)
-
[61]
K. R. Tindall and T. A. Kunkel, Biochemistry27, 6008 (1988)
work page 1988
-
[62]
P. Handal-Marquez, L. L. Torres, and V. B. Pinheiro, bioRxiv 10.1101/2022.11.08.515583 (2022), preprint posted November 8, 2022
-
[63]
J. G. Paez, M. Lin, R. Beroukhim, J. C. Lee, X. Zhao, D. J. Richter, S. Gabriel, P. Herman, H. Sasaki, D. Altshuler, C. Li, M. Meyerson, and W. R. Sellers, Nucleic Acids Research32, e71 (2004)
work page 2004
-
[64]
P. P´ erez-Arnaiz, J. M. L´ azaro, M. Salas, and M. de Vega, Nucleic Acids Research34, 3107 (2006)
work page 2006
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.