Recognition: unknown
Simple sign epistasis and evolutionary detours in fitness landscapes
Pith reviewed 2026-05-08 08:48 UTC · model grok-4.3
The pith
Simple sign epistasis forces evolutionary detours that include back-mutations to reach fitness peaks.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
The presence of simple sign epistasis is associated with evolutionary detours, that is, indirect longer fitness-increasing paths to fitness peaks that include back-mutations. In experimentally resolved weakly epistatic landscapes, simple sign epistasis occurs much more frequently than reciprocal sign epistasis. This result is consistent with theoretical predictions derived for most landscape models, with the exception of the block model and of landscapes dominated by pairwise allelic incompatibilities such as RNA stability landscapes. The results indicate that detours represent a general feature of evolutionary trajectories in weakly epistatic landscapes.
What carries the argument
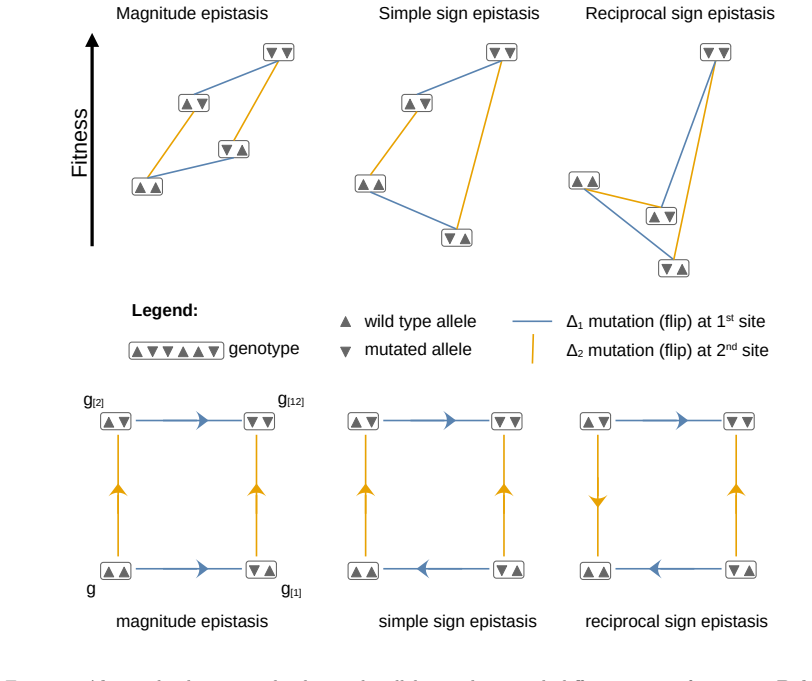
Simple sign epistasis, the case in which only one of two mutations changes the sign of its fitness effect depending on the presence of the other, which imposes conditional dependencies that block direct mutational routes and require reversals.
If this is right
- Evolutionary trajectories to higher fitness must often include reversals of prior mutations, making paths longer than the shortest mutational distance.
- Direct routes between genotypes can be inaccessible, requiring exploration of additional genetic backgrounds before ascent.
- Simple sign epistasis is expected to predominate over reciprocal sign epistasis in weakly epistatic regimes, rendering detours a common outcome.
- Most theoretical landscape models predict the observed excess of simple sign epistasis, except in special cases such as block models or RNA stability landscapes.
Where Pith is reading between the lines
- Accounting for possible back-mutation steps may be necessary to predict the order and timing of mutations observed in microbial evolution experiments.
- The prevalence of detours could reduce the predictability of which genetic backgrounds will ultimately fix in adapting populations.
- Similar detour dynamics may appear in other weakly epistatic systems such as protein sequence evolution where sign changes have been documented.
Load-bearing premise
That the experimentally resolved landscapes are representative of weakly epistatic regimes and that the theoretical models capture the dominant features of real fitness landscapes without strong selection biases or measurement artifacts.
What would settle it
An experimentally measured weakly epistatic landscape in which the frequency of simple sign epistasis is not substantially higher than reciprocal sign epistasis and in which evolutionary trajectories to peaks lack back-mutations would falsify the claimed association.
Figures






read the original abstract
In epistatic fitness landscapes, the fitness effect of a mutation depends on the genetic background and may even switch between deleterious and beneficial depending on the presence of another mutation. Epistatic interactions may cause both mutations to change the sign of each other's fitness effects (reciprocal sign epistasis) or only one mutation to do so (simple sign epistasis). Both these forms of epistasis influence evolutionary trajectories. While reciprocal sign epistasis has been associated with multi-peaked landscapes and their ruggedness, the role and relative frequency of simple sign epistasis in fitness landscapes have not been systematically investigated. Here, we prove that the presence of simple sign epistasis is associated with evolutionary detours, i.e., indirect, longer fitness-increasing paths to fitness peaks that include back-mutations. We also show that in experimentally resolved, weakly epistatic landscapes, simple sign epistasis occurs much more frequently than reciprocal sign epistasis. This result is consistent with the theoretical predictions we derive for most landscape models, with the exception of the block model and of landscapes dominated by pairwise allelic incompatibilities, such as RNA stability landscapes. Our results suggest that detours represent a general feature of evolutionary trajectories in weakly epistatic landscapes.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript proves that the presence of simple sign epistasis in fitness landscapes is associated with evolutionary detours—indirect, longer fitness-increasing paths to peaks that include back-mutations. It further reports that simple sign epistasis occurs much more frequently than reciprocal sign epistasis in experimentally resolved, weakly epistatic landscapes, a pattern consistent with theoretical predictions derived for most landscape models (with noted exceptions for the block model and RNA stability landscapes dominated by pairwise allelic incompatibilities).
Significance. If the proof is gap-free and the empirical frequency result is robust to data selection, this establishes detours as a general feature of trajectories in weakly epistatic regimes rather than a rare phenomenon tied only to reciprocal sign epistasis and ruggedness. The work derives parameter-free theoretical predictions across standard models and contrasts them with empirical data, providing a falsifiable link between a common epistatic interaction and path structure that could guide future experimental tests in microbial and viral systems.
major comments (2)
- [results on experimental landscapes] Empirical frequency claim (results section on experimental landscapes): the assertion that simple sign epistasis occurs 'much more frequently' than reciprocal sign epistasis rests on a small set of resolved landscapes whose selection criteria, measurement-error thresholds for sign flips, and exclusion rules are not fully specified; without these, it is impossible to rule out that the reported difference is an artifact of detection biases rather than a general feature.
- [discussion of model exceptions] Theoretical consistency statement (discussion of model exceptions): while the block model and RNA stability landscapes are correctly identified as exceptions, the manuscript does not quantify the fraction of real data that fall into these regimes or provide a statistical test showing that the majority of weakly epistatic landscapes align with the other models, weakening the claim that the empirical pattern is 'consistent with theoretical predictions for most landscape models'.
minor comments (2)
- [theoretical derivations] Notation for fitness effects and sign changes could be standardized across the theoretical derivations and empirical tables to avoid ambiguity in comparing models.
- [path diagrams] Figure legends for path diagrams should explicitly state the number of mutations and whether back-mutations are required for the detour.
Simulated Author's Rebuttal
We thank the referee for their constructive and detailed comments, which have improved the clarity of our work. We address each major comment point by point below and indicate revisions made to the manuscript.
read point-by-point responses
-
Referee: Empirical frequency claim (results section on experimental landscapes): the assertion that simple sign epistasis occurs 'much more frequently' than reciprocal sign epistasis rests on a small set of resolved landscapes whose selection criteria, measurement-error thresholds for sign flips, and exclusion rules are not fully specified; without these, it is impossible to rule out that the reported difference is an artifact of detection biases rather than a general feature.
Authors: We thank the referee for highlighting this issue. The original manuscript referenced standard datasets from the literature but did not provide exhaustive methodological details. In the revised manuscript, we have added a dedicated subsection in the Methods describing: the exact sources and inclusion criteria for the experimental landscapes (publicly available microbial and viral datasets with sufficient mutational resolution and classified as weakly epistatic via low epistasis measures such as roughness-to-slope ratio); the sign-flip detection thresholds (fitness effect sign changes must exceed twice the reported measurement error); and exclusion rules (omission of high-noise or incomplete landscapes). These additions enable readers to evaluate potential biases directly. The frequency difference is observed consistently across the qualifying datasets. revision: yes
-
Referee: Theoretical consistency statement (discussion of model exceptions): while the block model and RNA stability landscapes are correctly identified as exceptions, the manuscript does not quantify the fraction of real data that fall into these regimes or provide a statistical test showing that the majority of weakly epistatic landscapes align with the other models, weakening the claim that the empirical pattern is 'consistent with theoretical predictions for most landscape models'.
Authors: We agree that a quantitative meta-analysis of the prevalence of block-model or pairwise-incompatibility-dominated regimes across all published fitness landscapes is not provided and would require substantial additional data collection beyond the scope of this study. We have revised the Discussion to explicitly acknowledge this limitation, to reiterate that the block model and RNA stability cases represent specific mechanistic regimes, and to note that the majority of the weakly epistatic empirical landscapes we examined (from microbial and viral systems) align with the standard models for which the prediction holds. The consistency statement is therefore framed as applying to the predominant classes of models and data rather than a universal claim; we view this as a fair characterization while highlighting the need for future surveys. revision: partial
Circularity Check
No circularity: proof and predictions derived independently from model definitions and external data
full rationale
The paper's core result is a mathematical proof that simple sign epistasis implies evolutionary detours in fitness-increasing paths, derived from standard definitions of sign epistasis and landscape structure. Theoretical predictions are obtained by direct analysis of common models (NK, block, RNA) without fitting parameters to the experimental frequency data. The empirical frequency comparison uses independently resolved experimental landscapes as input and reports consistency as an observation rather than a forced outcome. No self-citations are invoked as load-bearing uniqueness theorems, no fitted inputs are relabeled as predictions, and no ansatz or renaming reduces the derivation to its inputs by construction. The chain remains self-contained against external benchmarks.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Fitness effects of mutations can be decomposed into additive and epistatic components
Reference graph
Works this paper leans on
-
[1]
[Ban22] Claudia Bank. “Epistasis and adaptation on fitness landscapes”. In:Annual review of ecology, evolution, and systematics53.1 (2022), pp. 457–479. [Bro+15] Sophie Brouillet et al. “MAGELLAN: a tool to visualize small fitness landscapes”. In:bioRxiv(2015). [CGB13] Kristina Crona, Devin Greene, and Miriam Barlow. “The peaks and geometry of fitness lan...
work page doi:10.1016/j 2022
-
[2]
[Dia+23] Juan Diaz-Colunga et al. “Global epistasis on fitness landscapes”. In:Philosophical Transactions of the Royal Society B: Biological Sciences378.1877 (2023). [DK14] J Arjan Gm De Visser and Joachim Krug. “Empirical fitness landscapes and the predictability of evolution”. In:Nature Reviews Genetics15.7 (2014), pp. 480–490. 20 [Dra93] John W Drake. ...
-
[3]
The expected time to cross extended fitness plateaus
[GW19] Mahan Ghafari and Daniel B Weissman. “The expected time to cross extended fitness plateaus”. In:Theoretical Population Biology129 (2019), pp. 54–67. [HPK17] Sungmin Hwang, Su-Chan Park, and Joachim Krug. “Genotypic complexity of Fisher’s geometric model”. In:Genetics206.2 (2017), pp. 1049–1079. [Hwa+18] Sungmin Hwang et al. “Universality classes of...
2019
-
[4]
[NSK14] Johannes Neidhart, Ivan G Szendro, and Joachim Krug. “Adaptation in tunably rugged fitness landscapes: the rough Mount Fuji model”. In:Genetics198.2 (2014), pp. 699–721. 21 [Pap+23] Andrei Papkou et al. “A rugged yet easily navigable fitness landscape”. In:Science 382.6673 (2023), eadh3860. [Phi08] Patrick C Phillips. “Epistasis–the essential role...
-
[5]
Structure of the ribosome-bound cricket paralysis virus IRES RNA
[Sch+06] Martin Sch¨ uler et al. “Structure of the ribosome-bound cricket paralysis virus IRES RNA”. In:Nature structural & molecular biology13.12 (2006), pp. 1092–1096. [SH99] Peter F Stadler and Robert Happel. “Random field models for fitness landscapes”. In:Journal of Mathematical Biology38.5 (1999), pp. 435–478. [Sil+10] Jack da Silva et al. “Fitness ...
2006
-
[6]
Evolutionary Accessibility of Modular Fitness Land- scapes
[SK14] B Schmiegelt and J Krug. “Evolutionary Accessibility of Modular Fitness Land- scapes”. In:Journal of Statistical Physics154 (2014), pp. 334–355. 22 [SKK22] Raimundo Saona, Fyodor A Kondrashov, and Ksenia A Khudiakova. “Relation be- tween the number of peaks and the number of reciprocal sign epistatic interactions”. In:Bulletin of Mathematical Biolo...
2014
-
[7]
On the incongruence of genotype-phenotype and fitness landscapes
[SP22] Malvika Srivastava and Joshua L Payne. “On the incongruence of genotype-phenotype and fitness landscapes”. In:PLoS computational biology18.9 (2022), e1010524. [Sri+26] Malvika Srivastava et al.Evolution as fitness landscape navigation: Concepts, Mea- sures, and Emerging Questions
2022
-
[8]
Evolution as fitness landscape navigation: Concepts, Measures, and Emerging Questions
arXiv:2604.17036 [q-bio.PE].url:https: //arxiv.org/abs/2604.17036. [Ste92] DL Stein, ed.Spin Glasses and Biology. Singapore: World Scientific,
work page internal anchor Pith review Pith/arXiv arXiv
-
[9]
Quantitative analyses of empirical fitness landscapes
[Sze+13] Ivan G Szendro et al. “Quantitative analyses of empirical fitness landscapes”. In: Journal of Statistical Mechanics: Theory and Experiment(2013). [Tan+11] Longzhi Tan et al. “Hidden randomness between fitness landscapes limits reverse evolution”. In:Physical review letters106.19 (2011), p. 198102. [VHE97] J A de Visser, R F Hoekstra, and H van de...
2013
-
[10]
The highly rugged yet navigable regulatory landscape of the bacterial transcription factor TetR
[WGW24] Cau˜ a Antunes Westmann, Leander Goldbach, and Andreas Wagner. “The highly rugged yet navigable regulatory landscape of the bacterial transcription factor TetR”. In:Nature Communications15.1 (2024), p. 10745. [Wri32] S Wright. “The roles of mutation, inbreeding, crossbreeding and selection in evolu- tion”. In:Proc. 6th Int. Cong. Genet.1 (1932), p...
2024
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.