Recognition: unknown
MP2D: Constrained Monte Carlo Tree-Guided Diffusion for Multi-Objective Protein Sequence Design
Pith reviewed 2026-05-08 03:07 UTC · model grok-4.3
The pith
MP2D guides diffusion denoising steps with constrained Monte Carlo tree search to optimize protein sequences for multiple conflicting properties at once.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
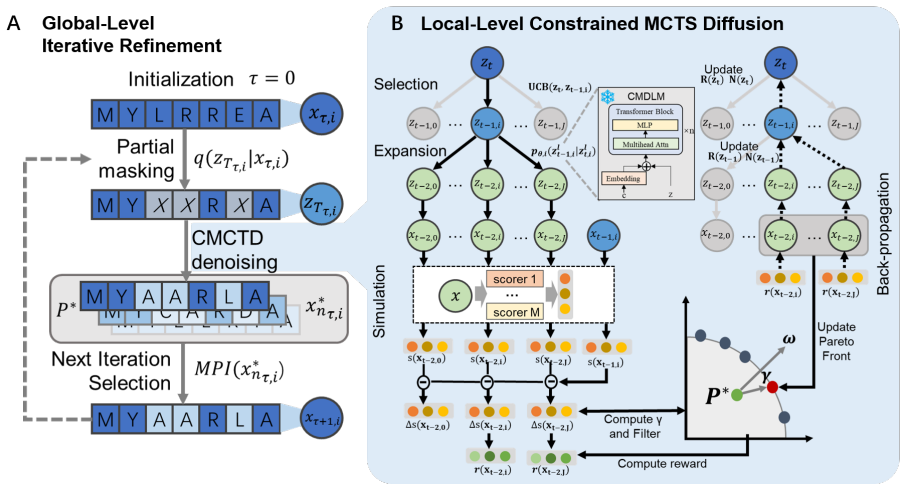
MP2D formulates diffusion denoising as a constrained sequential decision-making process, employs MCTS to explore diverse denoising trajectories guided by Pareto-based rewards, applies a dynamic Pareto constraint to prevent candidate bloat and maintain balanced trade-offs, and uses global iterative refinement to enable repeated remasking and re-optimization of candidate sequences, resulting in robust and balanced improvements across all objectives on multi-objective protein design tasks without retraining generative models.
What carries the argument
Constrained Monte Carlo tree search that explores diffusion denoising trajectories, guided by Pareto-based rewards together with dynamic Pareto constraints and global iterative refinement.
If this is right
- The same diffusion model can be reused across different sets of conflicting objectives without retraining.
- Balanced solutions emerge for four or five simultaneous properties in both antimicrobial peptide and protein binder tasks.
- Dynamic Pareto constraints keep the number of candidates manageable while preserving trade-off diversity.
- Global iterative refinement improves sequence quality beyond a single denoising run.
- The approach scales as a practical method for multi-objective functional protein design.
Where Pith is reading between the lines
- The same search-guided diffusion pattern could be tested on other sequence types such as RNA or designed small molecules.
- If the efficiency holds, it reduces the need to train separate generative models for each new combination of design goals.
- The method suggests that explicit trajectory search can handle objective spaces where pure sampling from a diffusion model tends to collapse to one corner of the Pareto front.
- A direct test would compare run-time and solution quality on proteins larger than those used in the current experiments.
Load-bearing premise
That MCTS exploration of denoising trajectories combined with dynamic Pareto constraints can reliably locate balanced multi-objective solutions without excessive cost or systematic bias toward particular trade-offs.
What would settle it
Applying MP2D to a fresh multi-objective protein design benchmark where it fails to improve all objectives simultaneously or produces more imbalanced results than existing baselines would falsify the central claim.
Figures
read the original abstract
Designing functional protein sequences that satisfy multiple desired properties is a core research focus of protein engineering. Prior methods struggle with inability or inefficiency when dealing with numerous, often conflicting, properties. We propose Multi-Property Protein Diffusion (MP2D), a unified framework for multi-objective protein sequence optimization that integrates conditional discrete diffusion with constrained MCTS and global iterative refinement. MP2D formulates diffusion denoising as a constrained sequential decision-making process and employs MCTS to explore diverse denoising trajectories guided by Pareto-based rewards. A global iterative refinement strategy further enables repeated remasking and re-optimization of candidate sequences, while a dynamic Pareto constraint prevents candidate bloat and maintains balanced trade-offs across objectives. We evaluate MP2D on two challenging multi-objective protein design tasks: antimicrobial peptide and protein binder optimization, involving four to five conflicting properties. Experimental results demonstrate that MP2D consistently outperforms existing multi-objective baselines, achieving robust and balanced improvements across all objectives without retraining generative models. These results highlight MP2D as a practical and scalable solution for multi-objective functional protein design.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper proposes MP2D, a framework integrating conditional discrete diffusion with constrained Monte Carlo Tree Search (MCTS) guided by Pareto-based rewards, dynamic Pareto constraints, and global iterative refinement via remasking and re-optimization. It formulates denoising as a sequential decision process to optimize protein sequences for multiple conflicting objectives and evaluates the method on antimicrobial peptide design and protein binder optimization tasks involving 4-5 properties, claiming consistent outperformance over multi-objective baselines without retraining the base generative model.
Significance. If the empirical claims hold with proper validation, MP2D provides a practical extension of diffusion-based generative models for multi-objective protein engineering by adding search-based guidance and refinement, potentially enabling balanced optimization of properties such as activity, stability, and specificity at lower computational cost than retraining approaches. This could be useful for tasks where pre-trained models exist but direct multi-objective sampling is challenging.
major comments (3)
- [§4] §4 (Experimental Results): The central claim of consistent outperformance and 'robust and balanced improvements across all objectives' is load-bearing, yet the reported results lack quantitative metrics (e.g., specific scores per objective, improvement deltas), baseline implementation details, ablation studies on MCTS components or the dynamic constraint, error bars, or statistical significance tests. This prevents verification that gains are reliable rather than due to stochastic variation in MCTS or task-specific tuning.
- [§3.3] §3.3 (Dynamic Pareto Constraint): The dynamic Pareto constraint is presented as preventing candidate bloat and maintaining balanced trade-offs, but no analysis quantifies its effect on the Pareto front or shows that it avoids systematic bias toward particular objectives. Without this, the claim that the method reliably discovers balanced multi-objective solutions rests on untested assumptions about the reward and constraint interaction.
- [§3.2] §3.2 (Constrained MCTS Formulation): The mapping of diffusion denoising to a constrained sequential decision process with Pareto rewards is central to the method, but the paper does not specify the exact reward function computation, tree expansion limits, or how constraints are enforced during trajectory sampling. This makes it unclear whether the approach introduces bias or excessive compute relative to simpler guidance methods.
minor comments (2)
- [Abstract / §1] The abstract and introduction would benefit from a brief table or bullet list summarizing the 4-5 objectives per task and the exact baselines compared, to improve readability before the detailed experiments.
- [§2 / §3] Notation for the diffusion process and MCTS state (e.g., how remasking is formalized) should be made consistent between §2 and §3 to avoid ambiguity in the algorithmic description.
Simulated Author's Rebuttal
We thank the referee for the constructive and detailed feedback. We address each major comment point by point below, providing clarifications from the manuscript and committing to specific revisions that will strengthen the empirical support and methodological transparency.
read point-by-point responses
-
Referee: [§4] §4 (Experimental Results): The central claim of consistent outperformance and 'robust and balanced improvements across all objectives' is load-bearing, yet the reported results lack quantitative metrics (e.g., specific scores per objective, improvement deltas), baseline implementation details, ablation studies on MCTS components or the dynamic constraint, error bars, or statistical significance tests. This prevents verification that gains are reliable rather than due to stochastic variation in MCTS or task-specific tuning.
Authors: We agree that additional quantitative detail and statistical rigor will improve verifiability. In the revised manuscript we will expand the experimental section with: (i) explicit per-objective scores and improvement deltas relative to baselines in the main tables; (ii) fuller baseline implementation details including hyperparameter settings and any public code references; (iii) new ablation studies isolating MCTS components (e.g., Pareto reward vs. standard UCT) and the dynamic constraint; (iv) error bars computed from at least five independent runs per method; and (v) statistical significance tests (paired t-tests or Wilcoxon rank-sum with Bonferroni correction) comparing MP2D against each baseline. These additions directly address the concern that observed gains could arise from stochastic variation. revision: yes
-
Referee: [§3.3] §3.3 (Dynamic Pareto Constraint): The dynamic Pareto constraint is presented as preventing candidate bloat and maintaining balanced trade-offs, but no analysis quantifies its effect on the Pareto front or shows that it avoids systematic bias toward particular objectives. Without this, the claim that the method reliably discovers balanced multi-objective solutions rests on untested assumptions about the reward and constraint interaction.
Authors: We acknowledge the value of explicit quantification. We will add a dedicated analysis (new subsection in §3.3 or supplementary material) that reports Pareto-front metrics (hypervolume, spread, and coverage) with and without the dynamic constraint across both tasks. We will also include objective-wise bias diagnostics (e.g., normalized contribution of each objective to selected sequences) and visualizations of front evolution to demonstrate that the constraint maintains balance without favoring any single objective. These experiments will be run on the same evaluation protocols already described. revision: yes
-
Referee: [§3.2] §3.2 (Constrained MCTS Formulation): The mapping of diffusion denoising to a constrained sequential decision process with Pareto rewards is central to the method, but the paper does not specify the exact reward function computation, tree expansion limits, or how constraints are enforced during trajectory sampling. This makes it unclear whether the approach introduces bias or excessive compute relative to simpler guidance methods.
Authors: We will revise §3.2 to supply the missing implementation details: the exact Pareto reward function (including the scalarization and normalization steps), concrete tree-expansion limits (maximum simulations per node, depth, and branching factor), and the precise constraint-enforcement procedure (pruning of invalid partial trajectories and back-propagation rules). These additions will allow readers to reproduce the search procedure and to compare its computational profile with simpler guidance baselines. revision: yes
Circularity Check
No significant circularity detected
full rationale
The paper proposes MP2D as an algorithmic integration of conditional discrete diffusion, constrained MCTS, Pareto rewards, dynamic constraints, and iterative refinement for multi-objective protein sequence design. No equations, derivations, or first-principles predictions are presented in the abstract or description that reduce by construction to fitted inputs, self-definitions, or self-citation chains. Outperformance claims rest on empirical evaluation against baselines on antimicrobial peptide and protein binder tasks rather than any internal mathematical equivalence or renamed known result. The method is framed as a practical extension of existing diffusion and search techniques without load-bearing self-referential steps.
Axiom & Free-Parameter Ledger
Reference graph
Works this paper leans on
-
[1]
Protein generation with evolutionary dif- fusion: sequence is all you need.BioRxiv, pages 2023–09,
[Alamdariet al., 2023 ] Sarah Alamdari, Nitya Thakkar, Ri- anne Van Den Berg, Neil Tenenholtz, Robert Strome, Alan M Moses, Alex X Lu, Nicol`o Fusi, Ava P Amini, and Kevin K Yang. Protein generation with evolutionary dif- fusion: sequence is all you need.BioRxiv, pages 2023–09,
2023
-
[2]
Sms-emoa: Multiobjective selection based on dominated hypervolume.European journal of operational research, 181(3):1653–1669,
[Beumeet al., 2007 ] Nicola Beume, Boris Naujoks, and Michael Emmerich. Sms-emoa: Multiobjective selection based on dominated hypervolume.European journal of operational research, 181(3):1653–1669,
2007
-
[3]
Monte-carlo tree search: A new framework for game ai
[Chaslotet al., 2008 ] Guillaume Chaslot, Sander Bakkes, Istvan Szita, and Pieter Spronck. Monte-carlo tree search: A new framework for game ai. InProceedings of the AAAI Conference on Artificial Intelligence and Interactive Digi- tal Entertainment, volume 4, pages 216–217,
2008
-
[4]
A web server and mobile app for computing hemolytic potency of peptides.Scientific re- ports, 6(1):22843,
[Chaudharyet al., 2016 ] Kumardeep Chaudhary, Ritesh Ku- mar, Sandeep Singh, Abhishek Tuknait, Ankur Gautam, Deepika Mathur, Priya Anand, Grish C Varshney, and Gajendra PS Raghava. A web server and mobile app for computing hemolytic potency of peptides.Scientific re- ports, 6(1):22843,
2016
-
[5]
[Chen and Li, 2023] Tao Chen and Miqing Li. The weights can be harmful: Pareto search versus weighted search in multi-objective search-based software engineering.ACM Transactions on Software Engineering and Methodology, 32(1):1–40,
2023
-
[6]
Amp-diffusion: Integrat- ing latent diffusion with protein language models for an- timicrobial peptide generation.bioRxiv, pages 2024–03,
[Chenet al., 2024 ] Tianlai Chen, Pranay Vure, Rishab Pu- lugurta, and Pranam Chatterjee. Amp-diffusion: Integrat- ing latent diffusion with protein language models for an- timicrobial peptide generation.bioRxiv, pages 2024–03,
2024
-
[7]
Multi-objective-guided discrete flow matching for controllable biological sequence design
[Chenet al., 2025 ] Tong Chen, Yinuo Zhang, Sophia Tang, and Pranam Chatterjee. Multi-objective-guided discrete flow matching for controllable biological sequence design. arXiv preprint arXiv:2505.07086,
-
[8]
Mopso: A proposal for multiple objective particle swarm optimization
[Coello and Lechuga, 2002] CA Coello Coello and Max- imino Salazar Lechuga. Mopso: A proposal for multiple objective particle swarm optimization. InProceedings of the 2002 Congress on Evolutionary Computation. CEC’02 (Cat. No. 02TH8600), volume 2, pages 1051–1056. IEEE,
2002
-
[9]
Uniprot: a world- wide hub of protein knowledge.Nucleic acids research, 47(D1):D506–D515,
[Consortium, 2019] UniProt Consortium. Uniprot: a world- wide hub of protein knowledge.Nucleic acids research, 47(D1):D506–D515,
2019
-
[10]
Protgpt2 is a deep unsupervised lan- guage model for protein design.Nature communications, 13(1):4348,
[Ferruzet al., 2022 ] Noelia Ferruz, Steffen Schmidt, and Birte H ¨ocker. Protgpt2 is a deep unsupervised lan- guage model for protein design.Nature communications, 13(1):4348,
2022
-
[11]
Computational design of proteins targeting the con- served stem region of influenza hemagglutinin.Science, 332(6031):816–821,
[Fleishmanet al., 2011 ] Sarel J Fleishman, Timothy A Whitehead, Damian C Ekiert, Cyrille Dreyfus, Jacob E Corn, Eva-Maria Strauch, Ian A Wilson, and David Baker. Computational design of proteins targeting the con- served stem region of influenza hemagglutinin.Science, 332(6031):816–821,
2011
-
[12]
Interactive hyperparameter optimization in multi-objective problems via preference learning
[Giovanelliet al., 2024 ] Joseph Giovanelli, Alexander Tornede, Tanja Tornede, and Marius Lindauer. Interactive hyperparameter optimization in multi-objective problems via preference learning. InProceedings of the AAAI Conference on Artificial Intelligence, volume 38, pages 12172–12180,
2024
-
[13]
Classifier-Free Diffusion Guidance
[Ho and Salimans, 2022] Jonathan Ho and Tim Salimans. Classifier-free diffusion guidance.arXiv preprint arXiv:2207.12598,
work page internal anchor Pith review arXiv 2022
-
[14]
Denoising diffusion probabilistic models.Advances in neural information processing systems, 33:6840–6851,
[Hoet al., 2020 ] Jonathan Ho, Ajay Jain, and Pieter Abbeel. Denoising diffusion probabilistic models.Advances in neural information processing systems, 33:6840–6851,
2020
-
[15]
Performance comparison of nsga-ii and nsga-iii on various many-objective test prob- lems
[Ishibuchiet al., 2016 ] Hisao Ishibuchi, Ryo Imada, Yu Se- toguchi, and Yusuke Nojima. Performance comparison of nsga-ii and nsga-iii on various many-objective test prob- lems. In2016 IEEE Congress on Evolutionary Computa- tion (CEC), pages 3045–3052. IEEE,
2016
-
[16]
A graph-based genetic algo- rithm and generative model/monte carlo tree search for the exploration of chemical space.Chemical science, 10(12):3567–3572,
[Jensen, 2019] Jan H Jensen. A graph-based genetic algo- rithm and generative model/monte carlo tree search for the exploration of chemical space.Chemical science, 10(12):3567–3572,
2019
-
[17]
Highly accu- rate protein structure prediction with alphafold.nature, 596(7873):583–589,
[Jumperet al., 2021 ] John Jumper, Richard Evans, Alexan- der Pritzel, Tim Green, Michael Figurnov, Olaf Ron- neberger, Kathryn Tunyasuvunakool, Russ Bates, Au- gustin ˇZ´ıdek, Anna Potapenko, et al. Highly accu- rate protein structure prediction with alphafold.nature, 596(7873):583–589,
2021
-
[18]
Rethinkmcts: Refining erroneous thoughts in monte carlo tree search for code generation
[Liet al., 2025 ] Qingyao Li, Wei Xia, Xinyi Dai, Kounian- hua Du, Weiwen Liu, Yasheng Wang, Ruiming Tang, Yong Yu, and Weinan Zhang. Rethinkmcts: Refining erroneous thoughts in monte carlo tree search for code generation. In Proceedings of the 2025 Conference on Empirical Meth- ods in Natural Language Processing, pages 8103–8121,
2025
-
[19]
Language models of protein sequences at the scale of evolution enable accurate structure prediction
[Linet al., 2022 ] Zeming Lin, Halil Akin, Roshan Rao, Brian Hie, Zhongkai Zhu, Wenting Lu, Allan dos San- tos Costa, Maryam Fazel-Zarandi, Tom Sercu, Sal Can- dido, et al. Language models of protein sequences at the scale of evolution enable accurate structure prediction. BioRxiv, 2022:500902,
2022
-
[20]
[Liuet al., 2025b ] Mengdi Liu, Xiaoxue Cheng, Zhangyang Gao, Hong Chang, Cheng Tan, Shiguang Shan, and Xilin Chen. Protinvtree: Deliberate protein inverse folding with reward-guided tree search.arXiv preprint arXiv:2506.00925,
-
[21]
Monte carlo tree diffusion with multiple experts for protein design.arXiv preprint arXiv:2509.15796,
[Liuet al., 2025c ] Xuefeng Liu, Mingxuan Cao, Songhao Jiang, Xiao Luo, Xiaotian Duan, Mengdi Wang, Tobin R Sosnick, Jinbo Xu, and Rick Stevens. Monte carlo tree diffusion with multiple experts for protein design.arXiv preprint arXiv:2509.15796,
-
[22]
[Luet al., 2025 ] Yining Lu, Zilong Wang, Shiyang Li, Xin Liu, Changlong Yu, Qingyu Yin, Zhan Shi, Zixuan Zhang, and Meng Jiang. Learning to optimize multi-objective alignment through dynamic reward weighting.arXiv preprint arXiv:2509.11452,
-
[23]
[Meshchaninovet al., 2024 ] Viacheslav Meshchaninov, Pavel Strashnov, Andrey Shevtsov, Fedor Nikolaev, Nikita Ivanisenko, Olga Kardymon, and Dmitry Vetrov. Diffusion on language model embeddings for protein sequence generation.arXiv preprint arXiv:2403.03726,
-
[24]
Searching for the pareto frontier in multi- objective protein design.Biophysical reviews, 9(4):339– 344,
[Nandaet al., 2017 ] Vikas Nanda, Sandeep V Belure, and Ofer M Shir. Searching for the pareto frontier in multi- objective protein design.Biophysical reviews, 9(4):339– 344,
2017
-
[25]
Pareto multi objective optimization
[Ngatchouet al., 2005 ] Patrick Ngatchou, Anahita Zarei, and A El-Sharkawi. Pareto multi objective optimization. InProceedings of the 13th international conference on, in- telligent systems application to power systems, pages 84–
2005
-
[26]
Dbaasp v3: database of antimicrobial/cytotoxic activity and structure of peptides as a resource for development of new therapeutics.Nucleic acids research, 49(D1):D288– D297,
[Pirtskhalavaet al., 2021 ] Malak Pirtskhalava, Anthony A Amstrong, Maia Grigolava, Mindia Chubinidze, Evgenia Alimbarashvili, Boris Vishnepolsky, Andrei Gabrielian, Alex Rosenthal, Darrell E Hurt, and Michael Tartakovsky. Dbaasp v3: database of antimicrobial/cytotoxic activity and structure of peptides as a resource for development of new therapeutics.Nu...
2021
-
[27]
Overcoming the challenges in machine learning-guided antimicrobial peptide design
[Plisson, 2022] Fabien Plisson. Overcoming the challenges in machine learning-guided antimicrobial peptide design. InProceedings of the 36th European and the 12th interna- tional peptide symposium, Sitges, Spain, pages 207–210,
2022
-
[28]
Toxinpred 3.0: An improved method for predicting the toxicity of peptides.Computers in biology and medicine, 179:108926,
[Rathoreet al., 2024 ] Anand Singh Rathore, Shubham Choudhury, Akanksha Arora, Purva Tijare, and Gajen- dra PS Raghava. Toxinpred 3.0: An improved method for predicting the toxicity of peptides.Computers in biology and medicine, 179:108926,
2024
-
[29]
Prediction of hemolytic peptides and their hemolytic concentration.Communications Biology, 8(1):176,
[Rathoreet al., 2025 ] Anand Singh Rathore, Nishant Kumar, Shubham Choudhury, Naman Kumar Mehta, and Gajen- dra PS Raghava. Prediction of hemolytic peptides and their hemolytic concentration.Communications Biology, 8(1):176,
2025
-
[30]
Ex- panding functional protein sequence spaces using gener- ative adversarial networks.Nature Machine Intelligence, 3(4):324–333,
[Repeckaet al., 2021 ] Donatas Repecka, Vykintas Jau- niskis, Laurynas Karpus, Elzbieta Rembeza, Irmantas Rokaitis, Jan Zrimec, Simona Poviloniene, Audrius Lau- rynenas, Sandra Viknander, Wissam Abuajwa, et al. Ex- panding functional protein sequence spaces using gener- ative adversarial networks.Nature Machine Intelligence, 3(4):324–333,
2021
-
[31]
Dramp 3.0: an enhanced comprehensive data repository of antimicrobial peptides
[Shiet al., 2022 ] Guobang Shi, Xinyue Kang, Fanyi Dong, Yanchao Liu, Ning Zhu, Yuxuan Hu, Hanmei Xu, Xingzhen Lao, and Heng Zheng. Dramp 3.0: an enhanced comprehensive data repository of antimicrobial peptides. Nucleic acids research, 50(D1):D488–D496,
2022
-
[32]
Peptune: De novo generation of therapeu- tic peptides with multi-objective-guided discrete diffusion
[Tanget al., 2025 ] Sophia Tang, Yinuo Zhang, and Pranam Chatterjee. Peptune: De novo generation of therapeu- tic peptides with multi-objective-guided discrete diffusion. ArXiv, pages arXiv–2412,
2025
-
[33]
Protein dynamism and evolvability.Science, 324(5924):203–207,
[Tokuriki and Tawfik, 2009] Nobuhiko Tokuriki and Dan S Tawfik. Protein dynamism and evolvability.Science, 324(5924):203–207,
2009
-
[34]
arXiv preprint arXiv:2502.14944 , year =
[Ueharaet al., 2025 ] Masatoshi Uehara, Xingyu Su, Yulai Zhao, Xiner Li, Aviv Regev, Shuiwang Ji, Sergey Levine, and Tommaso Biancalani. Reward-guided iterative refine- ment in diffusion models at test-time with applications to protein and dna design.arXiv preprint arXiv:2502.14944,
-
[35]
Deep learning improves antimicrobial peptide recognition.Bioinformatics, 34(16):2740–2747,
[Veltriet al., 2018 ] Daniel Veltri, Uday Kamath, and Amarda Shehu. Deep learning improves antimicrobial peptide recognition.Bioinformatics, 34(16):2740–2747,
2018
-
[36]
[Wanget al., 2024a ] Li Wang, Xiangzheng Fu, Jiahao Yang, Xinyi Zhang, Xiucai Ye, Yiping Liu, Tetsuya Sakurai, and Xiangxiang Zeng. Moformer: Multi-objective an- timicrobial peptide generation based on conditional trans- former joint multi-modal fusion descriptor.arXiv preprint arXiv:2406.02610,
-
[37]
Diffusion language models are versatile protein learners.arXiv preprint arXiv:2402.18567, 2024
[Wanget al., 2024b ] Xinyou Wang, Zaixiang Zheng, Fei Ye, Dongyu Xue, Shujian Huang, and Quanquan Gu. Diffu- sion language models are versatile protein learners.arXiv preprint arXiv:2402.18567,
-
[38]
Hmamp: Designing highly potent antimicrobial peptides using a hypervolume- driven multiobjective deep generative model.Journal of Medicinal Chemistry, 68(8):8346–8360,
[Wanget al., 2025 ] Li Wang, Yiping Liu, Xiangzheng Fu, Xiucai Ye, Junfeng Shi, Gary G Yen, Quan Zou, Xiangx- iang Zeng, and Dongsheng Cao. Hmamp: Designing highly potent antimicrobial peptides using a hypervolume- driven multiobjective deep generative model.Journal of Medicinal Chemistry, 68(8):8346–8360,
2025
-
[39]
High-resolution de novo structure prediction from primary sequence.BioRxiv, pages 2022–07,
[Wuet al., 2022 ] Ruidong Wu, Fan Ding, Rui Wang, Rui Shen, Xiwen Zhang, Shitong Luo, Chenpeng Su, Zuo- fan Wu, Qi Xie, Bonnie Berger, et al. High-resolution de novo structure prediction from primary sequence.BioRxiv, pages 2022–07,
2022
-
[40]
dbamp 3.0: updated resource of antimicrobial activity and structural annotation of peptides in the post-pandemic era
[Yaoet al., 2025 ] Lantian Yao, Jiahui Guan, Peilin Xie, Chia-Ru Chung, Zhihao Zhao, Danhong Dong, Yilin Guo, Wenyang Zhang, Junyang Deng, Yuxuan Pang, et al. dbamp 3.0: updated resource of antimicrobial activity and structural annotation of peptides in the post-pandemic era. Nucleic acids research, 53(D1):D364–D376,
2025
-
[41]
Monte carlo tree diffusion for system 2 planning, 2026
[Yoonet al., 2025 ] Jaesik Yoon, Hyeonseo Cho, Doojin Baek, Yoshua Bengio, and Sungjin Ahn. Monte carlo tree diffusion for system 2 planning.arXiv preprint arXiv:2502.07202,
-
[42]
Multi-cgan: deep generative model- based multiproperty antimicrobial peptide design.Journal of Chemical Information and Modeling, 64(1):316–326,
[Yuet al., 2023 ] Haoqing Yu, Ruheng Wang, Jianbo Qiao, and Leyi Wei. Multi-cgan: deep generative model- based multiproperty antimicrobial peptide design.Journal of Chemical Information and Modeling, 64(1):316–326,
2023
-
[43]
Efficient retrosynthetic planning with mcts exploration en- hanced a* search.Communications Chemistry, 7(1):52,
[Zhaoet al., 2024 ] Dengwei Zhao, Shikui Tu, and Lei Xu. Efficient retrosynthetic planning with mcts exploration en- hanced a* search.Communications Chemistry, 7(1):52,
2024
-
[44]
Spea2: Improving the strength pareto evo- lutionary algorithm.TIK report, 103,
[Zitzleret al., 2001 ] Eckart Zitzler, Marco Laumanns, and Lothar Thiele. Spea2: Improving the strength pareto evo- lutionary algorithm.TIK report, 103,
2001
-
[45]
All models use standard amino-acid tokenization with a sin- gle [MASK] token from ESM-2 [Linet al., 2022 ]
The hidden dropout rate is set to 0.0, layer norm eps to 0.00001, max po- sitional embedding to 1026 and GeLU activation functions. All models use standard amino-acid tokenization with a sin- gle [MASK] token from ESM-2 [Linet al., 2022 ]. The con- dition token is first projected with an added embedding layer and then summed with the embedded input after ...
2022
-
[46]
For condi- tional funetunes on functional peptides, we leveraged LoRA on layer 19-29 with rank 16 and dropout 0.1
Pretraining uses a masked diffusion objective with a linear noise schedule. For condi- tional funetunes on functional peptides, we leveraged LoRA on layer 19-29 with rank 16 and dropout 0.1. Classifier-free guidance is enabled by randomly dropping condition labels with probability 0.1 and cfg scale is set to 1.7. We used a batch size of 64 and AdamW optim...
2000
-
[47]
For the noise schedule, we used linear schedule
During generation, sequences are initialized as fully masked and iteratively denoised. For the noise schedule, we used linear schedule. A.2 Hyperparameter Settings The hyperparameter settings of CMCTD are summarized in Table S1 and the hyperparameter settings of global iterative refinement are summarized in Table S2. B MIC predictor B.1 Dataset We collect...
2022
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.