Recognition: no theorem link
ProteinJEPA: Latent prediction complements protein language models
Pith reviewed 2026-05-11 02:38 UTC · model grok-4.3
The pith
Masked-position latent prediction added to MLM outperforms pure masked language modeling on protein tasks.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
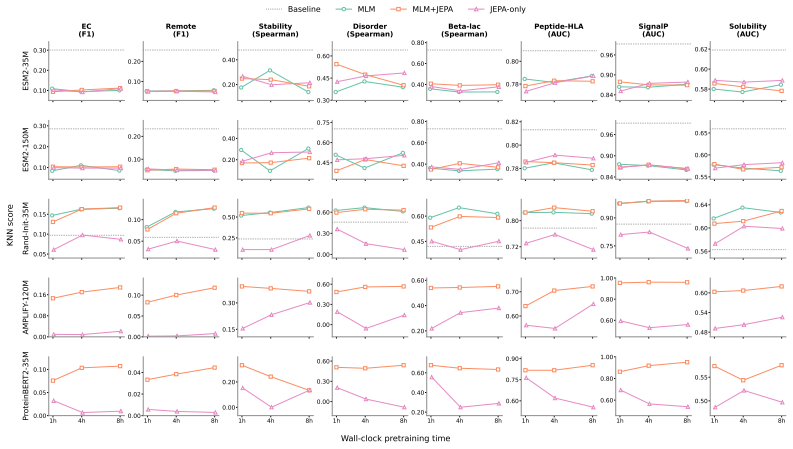
The central claim is that masked-position MLM+JEPA, which predicts latent targets exclusively at masked positions while retaining MLM cross-entropy, improves over MLM-only continuation, recording 10 wins / 3 losses / 3 ties on ESM2-35M and 11 wins / 2 losses / 3 ties on ESM2-150M across 15 linear probes and SCOPe-40 zero-shot retrieval, whereas all-position MLM+JEPA matches MLM overall and JEPA alone collapses.
What carries the argument
The masked-position MLM+JEPA objective that restricts latent target prediction to masked locations and keeps the token-level cross-entropy loss.
If this is right
- Gains occur on stability prediction, beta-lactamase fitness, variant effect, intrinsic disorder, remote homology, enzyme classification, and SCOPe-40 fold retrieval.
- Fluorescence (TAPE) and Peptide-HLA binding show more losses than wins.
- The pattern holds for continued training of pretrained ESM2 models at two sizes under matched wall-clock budgets.
- Training from random initialization yields mixed results.
- Removing the MLM component entirely causes collapse on nearly all tasks.
Where Pith is reading between the lines
- The same restriction of latent prediction to masked positions might improve JEPA hybrids in other sequence domains.
- Task-level patterns hint that latent prediction helps most on properties that depend on global structure rather than local sequence identity.
- Because the method requires no architecture changes it can be tested as a low-cost add-on to existing protein pretraining pipelines.
- The collapse of pure JEPA suggests that token-level supervision remains essential for learning useful protein representations.
Load-bearing premise
That the reported performance edges are produced by the addition of masked-position latent prediction rather than by small uncontrolled differences in optimization dynamics, data order, or random fluctuation.
What would settle it
Re-running every compared run with identical random seeds, identical data ordering, and exactly the same number of steps to check whether the win/loss ratio stays above one.
Figures




read the original abstract
Protein language models are trained primarily with masked language modeling (MLM), which predicts amino-acid identities at masked positions. We ask whether latent-space prediction can complement these token-level objectives under matched wall-clock budget. Across pretrained and random-init protein sequence encoders at 35--150M parameters, we find that the best protein-JEPA design is not all-position latent prediction but a variant: predicting latent targets only at masked positions, and retaining the MLM cross-entropy. We call this recipe masked-position MLM+JEPA. On a 16-task downstream suite (15 frozen linear probes plus SCOPe-40 zero-shot fold retrieval), under matched wall-clock budgets, this recipe wins more tasks than it loses against MLM-only continuation: 10 wins / 3 losses / 3 ties (hereafter W/L/T) on pretrained ESM2-35M, 11/2/3 on ESM2-150M while results in pretraining from scratch are mixed (6/8/2). Gains are seen for multiple models on 11 of 16 tasks, including stability, \b{eta}\beta \b{eta}-lactamase fitness, variant effect, intrinsic disorder, remote homology, enzyme classification, and SCOPe-40 fold retrieval. Tasks with more losses than wins are Fluorescence (TAPE) and Peptide-HLA Binding. All-position MLM+JEPA matches MLM-only overall but does not reproduce the masked-position gains. JEPA-only (no MLM) collapses in nearly every experiment. We conclude that JEPA, when combined with MLM, is competitive and can outperform pure MLM in pretraining and continued training, even under matched wall-clock budgets.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper claims that a hybrid training objective called masked-position MLM+JEPA—retaining masked language modeling cross-entropy while adding latent prediction only at masked positions—outperforms pure MLM continuation for protein sequence encoders (ESM2-35M and 150M) on a 16-task downstream suite under matched wall-clock budgets. It reports 10 wins/3 losses/3 ties for the 35M pretrained model and 11/2/3 for the 150M model, with gains on tasks including stability, variant effect, remote homology, and SCOPe-40 fold retrieval; all-position JEPA variants match MLM-only while JEPA-only collapses. Results from scratch pretraining are mixed (6/8/2). The central empirical finding is that latent prediction complements token-level objectives when applied selectively at masked sites.
Significance. If the performance differences hold under rigorous controls, the result would indicate that hybrid token-latent objectives can improve protein representations for multiple biological tasks without extra wall-clock cost, offering a practical alternative to pure MLM scaling. The work provides a concrete recipe (masked-position targets plus retained MLM) that is directly testable and falsifiable on the reported suite.
major comments (2)
- [§4] §4 (Experimental results and Table 1/2): The 10/3/3 and 11/2/3 win/loss/tie counts are reported from single runs without error bars, seed-averaged statistics, or per-task significance tests. Because the central claim rests on these counts being caused by the masked-position JEPA term rather than optimization stochasticity, the absence of variance estimates makes it impossible to assess whether the observed margins exceed noise.
- [§3.3] §3.3 and §4.1 (Wall-clock budget enforcement): No per-run timing logs, gradient-step counts, or hardware utilization data are supplied to verify that MLM+JEPA and MLM-only runs executed under truly identical wall-clock envelopes. If the additional JEPA forward passes increase per-step cost, the MLM+JEPA runs may have performed fewer updates, directly threatening the matched-budget premise that underpins the reported superiority.
minor comments (2)
- [Abstract] Abstract: the token “β-lactamase” is rendered with a broken LaTeX macro (”β-lactamase”); this should be corrected for readability.
- [§2] §2 (Related work): the discussion of prior latent-prediction methods in vision and language could usefully cite the original JEPA paper and recent protein-specific contrastive or predictive objectives to clarify the precise novelty of the masked-position variant.
Simulated Author's Rebuttal
We thank the referee for the constructive comments, which help clarify the strength of our empirical claims. We address each major point below with clarifications and indicate planned revisions.
read point-by-point responses
-
Referee: [§4] §4 (Experimental results and Table 1/2): The 10/3/3 and 11/2/3 win/loss/tie counts are reported from single runs without error bars, seed-averaged statistics, or per-task significance tests. Because the central claim rests on these counts being caused by the masked-position JEPA term rather than optimization stochasticity, the absence of variance estimates makes it impossible to assess whether the observed margins exceed noise.
Authors: We acknowledge that the win/loss/tie counts derive from single training runs without error bars, seed averages, or per-task significance tests, which leaves open the possibility that some margins reflect optimization stochasticity rather than the masked-position JEPA term. At the same time, the pattern of gains is consistent across two independent model scales (35 M and 150 M) and spans multiple biologically distinct tasks, which lowers the probability that every observed improvement is noise. We agree that variance estimates would strengthen the central claim. In the revised manuscript we will add an explicit discussion of this limitation in §4 and, where additional compute permits, report standard deviations from a small number of repeated seeds on the most critical tasks. revision: partial
-
Referee: [§3.3] §3.3 and §4.1 (Wall-clock budget enforcement): No per-run timing logs, gradient-step counts, or hardware utilization data are supplied to verify that MLM+JEPA and MLM-only runs executed under truly identical wall-clock envelopes. If the additional JEPA forward passes increase per-step cost, the MLM+JEPA runs may have performed fewer updates, directly threatening the matched-budget premise that underpins the reported superiority.
Authors: We matched wall-clock budgets by first measuring per-step wall-clock time for each objective on the same hardware and then reducing the step count for the MLM+JEPA runs to compensate for the modest extra cost of the latent-prediction head. This calibration was performed internally before the final runs. While detailed timing logs were omitted from the original submission, the budgets were deliberately equalized. In the revision we will add an appendix containing per-run timing measurements, total gradient-step counts, and hardware specifications so that readers can verify the matched-budget condition directly. revision: yes
Circularity Check
No circularity: purely empirical objective comparison
full rationale
The paper reports experimental results from training protein encoders with MLM, JEPA variants, and their combinations, then evaluates downstream performance on a fixed 16-task suite under claimed matched wall-clock budgets. No derivation, uniqueness theorem, ansatz, or fitted parameter is invoked whose output is then relabeled as a prediction; the W/L/T counts are direct measurements, not quantities defined by construction from the inputs. Self-citations (if any) are not load-bearing for any central claim. The analysis is self-contained as standard empirical benchmarking.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Masked language modeling and JEPA-style latent prediction are valid self-supervised signals for sequence data
Reference graph
Works this paper leans on
-
[1]
Bioinformatics , author =. 2022 , pages =. doi:10.1093/bioinformatics/btac020 , abstract =
-
[2]
Hayes, Thomas and Rao, Roshan and Akin, Halil and Sofroniew, Nicholas J. and Oktay, Deniz and Lin, Zeming and Verkuil, Robert and Tran, Vincent Q. and Deaton, Jonathan and Wiggert, Marius and Badkundri, Rohil and Shafkat, Irhum and Gong, Jun and Derry, Alexander and Molina, Raul S. and Thomas, Neil and Khan, Yousuf A. and Mishra, Chetan and Kim, Carolyn a...
-
[3]
Lawrence and Ma, Jerry and Fergus, Rob , year =
Rives, Alexander and Goyal, Siddharth and Meier, Joshua and Guo, Demi and Ott, Myle and Zitnick, C. Lawrence and Ma, Jerry and Fergus, Rob , year =. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences , doi =
-
[4]
Medium-sized protein language models perform well at transfer learning on realistic datasets , volume =. Scientific Reports , author =. 2025 , keywords =. doi:10.1038/s41598-025-05674-x , abstract =
-
[5]
Balestriero, Randall and LeCun, Yann , month = nov, year =. doi:10.48550/arXiv.2511.08544 , abstract =
-
[6]
Huang, Hai and LeCun, Yann and Balestriero, Randall , month = oct, year =. doi:10.48550/arXiv.2509.14252 , abstract =
-
[7]
Discovering. Neural Computation , author =. 1993 , pages =. doi:10.1162/neco.1993.5.4.625 , abstract =
-
[8]
and Ye, Chengzhong and Song, Yun S
Benegas, Gonzalo and Albors, Carlos and Aw, Alan J. and Ye, Chengzhong and Song, Yun S. , month = oct, year =. doi:10.1101/2023.10.10.561776 , abstract =
-
[9]
doi: 10.1038/s41586-021-03819-2
Highly accurate protein structure prediction with. Nature , author =. doi:10.1038/s41586-021-03819-2 , language =
-
[10]
Maes, Lucas and Lidec, Quentin Le and Scieur, Damien and LeCun, Yann and Balestriero, Randall , month = mar, year =. doi:10.48550/arXiv.2603.19312 , abstract =
-
[11]
Computational and Structural Biotechnology Journal , author =
The language of proteins:. Computational and Structural Biotechnology Journal , author =. 2021 , pages =. doi:10.1016/j.csbj.2021.03.022 , language =
-
[12]
doi:10.1101/2020.07.12.199554 , abstract =
Elnaggar, Ahmed and Heinzinger, Michael and Dallago, Christian and Rehawi, Ghalia and Wang, Yu and Jones, Llion and Gibbs, Tom and Feher, Tamas and Angerer, Christoph and Steinegger, Martin and Bhowmik, Debsindhu and Rost, Burkhard , month = jul, year =. doi:10.1101/2020.07.12.199554 , abstract =
-
[13]
NAR Genomics and Bioinformatics , author =
Detecting anomalous proteins using deep representations , volume =. NAR Genomics and Bioinformatics , author =. 2024 , pages =. doi:10.1093/nargab/lqae021 , abstract =
-
[14]
Fournier, Quentin and Vernon, Robert M. and Sloot, Almer van der and Schulz, Benjamin and Chandar, Sarath and Langmead, Christopher James , month = jan, year =. Protein. doi:10.1101/2024.09.23.614603 , abstract =
-
[15]
Progen2: exploring the boundaries of protein language models.arXiv preprint arXiv:2206.13517, 2022
Nijkamp, Erik and Ruffolo, Jeffrey and Weinstein, Eli N. and Naik, Nikhil and Madani, Ali , month = jun, year =. doi:10.48550/arXiv.2206.13517 , abstract =
-
[16]
doi:10.1101/2023.10.01.560349 , abstract =
Su, Jin and Han, Chenchen and Zhou, Yuyang and Shan, Junjie and Zhou, Xibin and Yuan, Fajie , month = mar, year =. doi:10.1101/2023.10.01.560349 , abstract =
-
[17]
arXiv:1906.08230 [cs, q-bio, stat] , author =
Evaluating. arXiv:1906.08230 [cs, q-bio, stat] , author =. 2019 , note =
-
[18]
Patrick Wilhelm, Thorsten Wittkopp, and Odej Kao
Warner, Benjamin and Chaffin, Antoine and Clavié, Benjamin and Weller, Orion and Hallström, Oskar and Taghadouini, Said and Gallagher, Alexis and Biswas, Raja and Ladhak, Faisal and Aarsen, Tom and Cooper, Nathan and Adams, Griffin and Howard, Jeremy and Poli, Iacopo , month = dec, year =. Smarter,. doi:10.48550/arXiv.2412.13663 , abstract =
-
[19]
Bioinformatics (Oxford, England) , author =. 2015 , keywords =. doi:10.1093/bioinformatics/btu739 , abstract =
-
[20]
Bioinformatics (Oxford, England) , author =. 2014 , pages =. doi:10.1093/bioinformatics/btt725 , abstract =
-
[21]
Nucleic Acids Research , author =. 2014 , keywords =. doi:10.1093/nar/gku363 , abstract =
-
[22]
Dass, Rupashree and Mulder, Frans A. A. and Nielsen, Jakob Toudahl , month = sep, year =. Scientific Reports , publisher =. doi:10.1038/s41598-020-71716-1 , abstract =
-
[23]
Nature Biotechnology , author =. 2022 , keywords =. doi:10.1038/s41587-021-01156-3 , abstract =
-
[24]
Nucleic Acids Research , author =. 1999 , pages =. doi:10.1093/nar/27.1.254 , language =
-
[25]
Assran, Mahmoud and Duval, Quentin and Misra, Ishan and Bojanowski, Piotr and Vincent, Pascal and Rabbat, Michael and LeCun, Yann and Ballas, Nicolas , month = apr, year =. Self-. doi:10.48550/arXiv.2301.08243 , abstract =
-
[26]
Revisiting Feature Prediction for Learning Visual Representations from Video
Bardes, Adrien and Garrido, Quentin and Ponce, Jean and Chen, Xinlei and Rabbat, Michael and LeCun, Yann and Assran, Mahmoud and Ballas, Nicolas , month = feb, year =. Revisiting. doi:10.48550/arXiv.2404.08471 , abstract =
work page internal anchor Pith review doi:10.48550/arxiv.2404.08471
-
[27]
Larey, Ariel and Dahan, Elay and Bleiweiss, Amit and Kellerman, Raizy and Leib, Guy and Nayshool, Omri and Ofer, Dan and Zinger, Tal and Dominissini, Dan and Rechavi, Gideon and Bussola, Nicole and Lee, Simon and O'Connell, Shane and Hoang, Dung and Wirth, Marissa and Charney, Alexander W. and Daniel, Nati and Shavit, Yoli , month = feb, year =. doi:10.48...
-
[28]
Bioinformatics , author =. 2007 , note =. doi:10.1093/bioinformatics/btm098 , abstract =
-
[29]
Nature Methods , author =. 2025 , keywords =. doi:10.1038/s41592-025-02656-9 , abstract =
-
[30]
Nucleic acids research , author =. 2014 , keywords =. doi:10.1093/nar/gkt1240 , abstract =
-
[31]
FlashAttention-2: Faster Attention with Better Parallelism and Work Partitioning
Dao, Tri , month = jul, year =. doi:10.48550/arXiv.2307.08691 , abstract =
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.2307.08691
-
[32]
bioRxiv , author =. 2021 , pages =. doi:10.1101/2021.02.12.430858 , abstract =
-
[33]
Learning the language of viral evolution and escape , volume =. Science , author =. 2021 , pages =. doi:10.1126/science.abd7331 , abstract =
-
[34]
Ofer, Dan and Linial, Michal , month = aug, year =. Protein. Viruses , publisher =. doi:10.3390/v17091199 , abstract =
-
[35]
Unsupervised evolution of protein and antibody complexes with a structure-informed language model , volume =. Science , author =. 2024 , pmid =. doi:10.1126/science.adk8946 , abstract =
-
[36]
and Alamdari, Sarah and Lee, Alex J
Yang, Kevin K. and Alamdari, Sarah and Lee, Alex J. and Kaymak-Loveless, Kaeli and Char, Samir and Brixi, Garyk and Domingo-Enrich, Carles and Wang, Chentong and Lyu, Suyue and Fusi, Nicolo and Tenenholtz, Neil and Amini, Ava P. , month = jul, year =. The. doi:10.1101/2025.07.21.665991 , abstract =
-
[37]
Bioinformatics Advances , author =
Folding the unfoldable: using. Bioinformatics Advances , author =. 2022 , pages =. doi:10.1093/bioadv/vbab043 , abstract =
-
[38]
Cheng, Jun and Novati, Guido and Pan, Joshua and Bycroft, Clare and Žemgulytė, Akvilė and Applebaum, Taylor and Pritzel, Alexander and Wong, Lai Hong and Zielinski, Michal and Sargeant, Tobias and Schneider, Rosalia G. and Senior, Andrew W. and Jumper, John and Hassabis, Demis and Kohli, Pushmeet and Avsec, Žiga , month = sep, year =. Accurate proteome-wi...
-
[39]
Genomic heterogeneity inflates the performance of variant pathogenicity predictions , copyright =
Lu, Baiyu and Liu, Xueshen and Lin, Po-Yu and Brandes, Nadav , month = sep, year =. Genomic heterogeneity inflates the performance of variant pathogenicity predictions , copyright =. doi:10.1101/2025.09.05.674459 , abstract =
-
[40]
bioRxiv , author =. 2023 , pages =. doi:10.1101/2023.12.07.570727 , abstract =
-
[41]
Nucleic Acids Research , author =
The. Nucleic Acids Research , author =. 2023 , keywords =. doi:10.1093/nar/gkac1000 , abstract =
-
[42]
Global analysis of protein folding using massively parallel design, synthesis, and testing , volume =. Science , author =. 2017 , pages =. doi:10.1126/science.aan0693 , language =
-
[43]
Sarkisyan, Karen S. and Bolotin, Dmitry A. and Meer, Margarita V. and Usmanova, Dinara R. and Mishin, Alexander S. and Sharonov, George V. and Ivankov, Dmitry N. and Bozhanova, Nina G. and Baranov, Mikhail S. and Soylemez, Onuralp and Bogatyreva, Natalya S. and Vlasov, Peter K. and Egorov, Evgeny S. and Logacheva, Maria D. and Kondrashov, Alexey S. and Ch...
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.