Recognition: no theorem link
Yeti: A compact protein structure tokenizer for reconstruction and multi-modal generation
Pith reviewed 2026-05-12 03:57 UTC · model grok-4.3
The pith
Yeti is a compact tokenizer that converts protein structures into discrete tokens using lookup-free quantization and flow matching, enabling strong reconstruction and multimodal sequence-structure generation from scratch.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
Yeti demonstrates that a lookup-free quantization tokenizer trained with flow matching produces highly usable discrete structure tokens, achieving the best codebook utilization and token diversity plus second-best reconstruction accuracy on multiple datasets while using roughly one-tenth the parameters of ESM3; a compact multimodal model trained from scratch on Yeti tokens and amino acid sequences then generates plausible joint sequence-structure outputs that compare favorably to models ten times larger.
What carries the argument
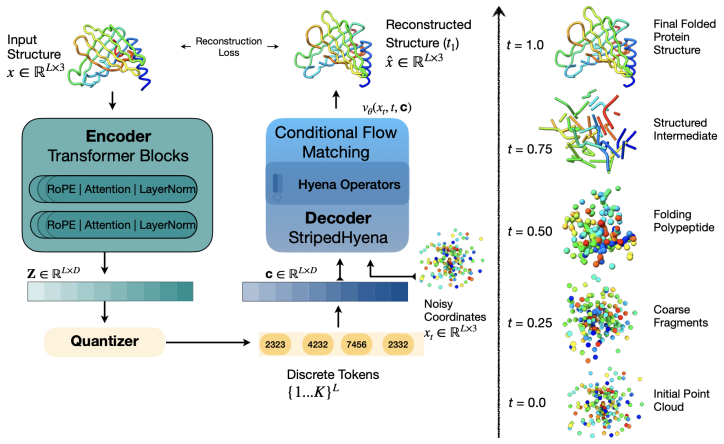
lookup-free quantization combined with an end-to-end flow matching objective that discretizes continuous protein coordinates into tokens while optimizing for both reconstruction fidelity and downstream generative use.
If this is right
- Multimodal protein models can be trained from scratch at smaller scale while still producing usable sequence-structure pairs.
- Tokenizers that maximize codebook utilization and diversity reduce the parameter count needed for competitive protein generation.
- Flow matching provides a stable training signal for learning structure tokens that preserve geometric information suitable for transformers.
- Joint unconditional generation of sequence and structure becomes feasible without separate pretraining stages for the tokenizer.
Where Pith is reading between the lines
- If Yeti tokens preserve sufficient geometric detail, they could be swapped into existing larger multimodal architectures to reduce overall model size without retraining the tokenizer.
- The emphasis on token diversity suggests Yeti may support more varied sampling in conditional generation tasks such as function-guided protein design.
- Because the multimodal model was trained without pretraining, Yeti may enable faster iteration on new protein modalities like binding sites or dynamics.
Load-bearing premise
Superior codebook utilization, token diversity, and reconstruction metrics on held-out data will translate into better generative performance when the tokenizer is used inside a multimodal model trained entirely from scratch.
What would settle it
Training the multimodal model on Yeti tokens and measuring joint sequence-structure generation quality on a new held-out protein family where the outputs show clear structural or sequence implausibility compared with 10x larger baselines.
Figures




read the original abstract
Multimodal models that jointly reason over protein sequences, structures, and function annotations within a unified representation hold immense potential for integrating multimodal data and generating new proteins with designed functional properties. To utilize transformer architectures, such models require a tokenizer that converts protein structure from continuous atomic coordinates into discrete representations suitable for scalable multimodal training. The quality of such models are fundamentally upper bounded by the fidelity and expressiveness of the underlying tokenized structure. However, existing tokenizers prioritize reconstruction over generative abilities. To address these gaps, we introduce Yeti, a simple and compact protein structure tokenizer based on lookup free quantization and trained end to end with a flow matching objective for multimodal learning. Compared to existing models, Yeti generally achieves the best codebook utilization and token diversity, and second best reconstruction accuracy (with 10x fewer parameters than ESM3) on diverse datasets. To validate Yeti's generative capability, we trained a compact multimodal model jointly over its structure tokens and amino acid sequence entirely from scratch, with no pretrained initialization. The resulting multimodal model generates plausible structures under unconditional cogeneration of protein sequence and structures, achieving comparable results to 10x larger models. Together, these results demonstrate that Yeti is a compact and expressive protein structure tokenizer suitable for training multimodal models that cogenerates highly plausible sequences and structures.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper introduces Yeti, a compact protein structure tokenizer based on lookup-free quantization trained end-to-end with a flow matching objective. It reports that Yeti achieves the best codebook utilization and token diversity, along with second-best reconstruction accuracy using 10x fewer parameters than ESM3, across diverse datasets. A compact multimodal model is then trained entirely from scratch on amino acid sequences and Yeti structure tokens, demonstrating unconditional cogeneration of plausible protein sequences and structures with results comparable to 10x larger models.
Significance. If the results hold, Yeti provides an efficient, low-parameter tokenizer that could lower barriers to training multimodal protein models for sequence-structure-function reasoning and generative design. The from-scratch training of the multimodal model offers a clean baseline, and the emphasis on codebook utilization and diversity highlights an under-appreciated aspect of tokenizer quality for downstream generation.
major comments (2)
- [Multimodal generation experiments] Multimodal generation experiments (validation section): The claim that Yeti's codebook utilization and token diversity causally enable plausible unconditional cogeneration rests on training one multimodal model with Yeti tokens and comparing outputs to larger models. No ablation is presented that holds the multimodal architecture, data, and optimization fixed while substituting an alternative tokenizer (e.g., a standard VQ-VAE or ESM3-derived tokens) that matches Yeti on reconstruction but differs on utilization/diversity. Without this isolation, the tokenizer properties could be incidental to the generative results.
- [Results section, comparative tables] Comparative tables (results section): The abstract states Yeti 'generally achieves the best codebook utilization and token diversity' and 'second best reconstruction accuracy,' yet the manuscript provides no error bars, multiple random seeds, or statistical tests for these metrics across datasets. This weakens the strength of the efficiency claim relative to ESM3 and other baselines.
minor comments (2)
- [Introduction] The introduction could briefly define 'lookup free quantization' and cite the originating method rather than assuming familiarity.
- [Figures] Figure captions for generation examples should specify the exact conditioning (unconditional vs. partial) and any filtering applied to the sampled structures.
Simulated Author's Rebuttal
We thank the referee for their constructive feedback on our manuscript. We address each major comment below and describe the revisions we will make to strengthen the presentation of our results.
read point-by-point responses
-
Referee: [Multimodal generation experiments] Multimodal generation experiments (validation section): The claim that Yeti's codebook utilization and token diversity causally enable plausible unconditional cogeneration rests on training one multimodal model with Yeti tokens and comparing outputs to larger models. No ablation is presented that holds the multimodal architecture, data, and optimization fixed while substituting an alternative tokenizer (e.g., a standard VQ-VAE or ESM3-derived tokens) that matches Yeti on reconstruction but differs on utilization/diversity. Without this isolation, the tokenizer properties could be incidental to the generative results.
Authors: We appreciate the referee's point that our experiments do not isolate the causal contribution of codebook utilization and token diversity through a controlled ablation. The multimodal model was trained entirely from scratch on Yeti tokens to demonstrate that a compact architecture can achieve plausible unconditional sequence-structure cogeneration comparable to 10x larger models. We agree this leaves open the possibility that the tokenizer metrics are not the sole driver. In the revised manuscript we will add a paragraph in the discussion explicitly acknowledging that the current results are correlative rather than causal and that future work could include ablations holding the multimodal backbone fixed while swapping tokenizers with matched reconstruction fidelity. revision: partial
-
Referee: [Results section, comparative tables] Comparative tables (results section): The abstract states Yeti 'generally achieves the best codebook utilization and token diversity' and 'second best reconstruction accuracy,' yet the manuscript provides no error bars, multiple random seeds, or statistical tests for these metrics across datasets. This weakens the strength of the efficiency claim relative to ESM3 and other baselines.
Authors: The referee correctly notes the lack of error bars, multi-seed results, and statistical tests. We will rerun the tokenizer training and evaluation on the reported datasets with at least three independent random seeds, add error bars to all comparative tables, and include appropriate statistical tests (e.g., paired t-tests or Wilcoxon tests) for the key metrics of codebook utilization, token diversity, and reconstruction accuracy. These updates will be incorporated into the results section and tables in the revised manuscript. revision: yes
Circularity Check
No circularity in derivation chain; claims are empirical.
full rationale
The paper introduces Yeti via lookup-free quantization and flow-matching training, then reports empirical results on codebook utilization, token diversity, reconstruction accuracy, and multimodal generation performance on held-out data. No equations, self-definitions, fitted-parameter predictions, or self-citation chains reduce any claimed result to an input quantity by construction. All performance statements rest on direct training outcomes and external model comparisons rather than tautological reductions.
Axiom & Free-Parameter Ledger
Reference graph
Works this paper leans on
-
[1]
Nabin Giri, Liguo Wang, and Jianlin Cheng. Cryo2structdata: A large labeled cryo-em density map dataset for ai-based modeling of protein structures.Scientific Data, 11(1):458, 2024
work page 2024
-
[2]
Nabin Giri, Xiao Chen, Liguo Wang, and Jianlin Cheng. A labeled dataset for ai-based cryo-em map enhancement.Computational and Structural Biotechnology Journal, 27:2843–2850, 2025
work page 2025
-
[3]
Simulating 500 million years of evolution with a language model.Science, 387(6736):850–858, 2025
Thomas Hayes, Roshan Rao, Halil Akin, Nicholas J Sofroniew, Deniz Oktay, Zeming Lin, Robert Verkuil, Vincent Q Tran, Jonathan Deaton, Marius Wiggert, et al. Simulating 500 million years of evolution with a language model.Science, 387(6736):850–858, 2025
work page 2025
-
[4]
Dplm-2: A multimodal diffusion protein language model.arXiv preprint arXiv:2410.13782, 2024
Xinyou Wang, Zaixiang Zheng, Fei Ye, Dongyu Xue, Shujian Huang, and Quanquan Gu. Dplm-2: A multimodal diffusion protein language model.arXiv preprint arXiv:2410.13782, 2024
-
[5]
Saprot: Protein language modeling with structure-aware vocabulary.BioRxiv, pages 2023–10, 2023
Jin Su, Chenchen Han, Yuyang Zhou, Junjie Shan, Xibin Zhou, and Fajie Yuan. Saprot: Protein language modeling with structure-aware vocabulary.BioRxiv, pages 2023–10, 2023
work page 2023
-
[6]
Nabin Giri, Raj S Roy, and Jianlin Cheng. Deep learning for reconstructing protein structures from cryo-em density maps: Recent advances and future directions.Current opinion in structural biology, 79:102536, 2023
work page 2023
-
[7]
Nabin Giri and Jianlin Cheng. De novo atomic protein structure modeling for cryoem density maps using 3d transformer and hmm.Nature Communications, 15(1):5511, 2024
work page 2024
-
[8]
Protein structure tokenization: Benchmarking and new recipe.arXiv preprint arXiv:2503.00089, 2025
Xinyu Yuan, Zichen Wang, Marcus Collins, and Huzefa Rangwala. Protein structure tokenization: Benchmarking and new recipe.arXiv preprint arXiv:2503.00089, 2025
-
[9]
Andrew Liu, Axel Elaldi, Nathan Russell, and Olivia Viessmann. Bio2token: All-atom tokenization of any biomolecular structure with mamba.arXiv preprint arXiv:2410.19110, 2024
-
[10]
Foldtoken: Learning protein language via vector quantization and beyond
Zhangyang Gao, Cheng Tan, Jue Wang, Yufei Huang, Lirong Wu, and Stan Z Li. Foldtoken: Learning protein language via vector quantization and beyond. InProceedings of the AAAI Conference on Artificial Intelligence, volume 39, pages 219–227, 2025
work page 2025
-
[11]
Highly accurate protein structure prediction with alphafold.nature, 596(7873):583–589, 2021
John Jumper, Richard Evans, Alexander Pritzel, Tim Green, Michael Figurnov, Olaf Ronneberger, Kathryn Tunyasuvunakool, Russ Bates, Augustin Žídek, Anna Potapenko, et al. Highly accurate protein structure prediction with alphafold.nature, 596(7873):583–589, 2021
work page 2021
-
[12]
Neural discrete representation learning.Advances in neural information processing systems, 30, 2017
Aaron Van Den Oord, Oriol Vinyals, et al. Neural discrete representation learning.Advances in neural information processing systems, 30, 2017
work page 2017
-
[13]
Stephen K Burley, Helen M Berman, Gerard J Kleywegt, John L Markley, Haruki Nakamura, and Sameer Velankar. Protein data bank (pdb): the single global macromolecular structure archive.Protein crystallography: methods and protocols, pages 627–641, 2017
work page 2017
-
[14]
Learning inverse folding from millions of predicted structures
Chloe Hsu, Robert Verkuil, Jason Liu, Zeming Lin, Brian Hie, Tom Sercu, Adam Lerer, and Alexander Rives. Learning inverse folding from millions of predicted structures. InInternational conference on machine learning, pages 8946–8970. PMLR, 2022
work page 2022
-
[15]
Language Model Beats Diffusion -- Tokenizer is Key to Visual Generation
Lijun Yu, José Lezama, Nitesh B Gundavarapu, Luca Versari, Kihyuk Sohn, David Minnen, Yong Cheng, Vighnesh Birodkar, Agrim Gupta, Xiuye Gu, et al. Language model beats diffusion–tokenizer is key to visual generation.arXiv preprint arXiv:2310.05737, 2023
work page internal anchor Pith review arXiv 2023
-
[16]
Flow autoencoders are effective protein tokenizers.arXiv preprint arXiv:2510.00351, 2025
Rohit Dilip, Evan Zhang, Ayush Varshney, and David Van Valen. Flow autoencoders are effective protein tokenizers.arXiv preprint arXiv:2510.00351, 2025
-
[17]
Finite scalar quantization: Vq-vae made simple.arXiv preprint arXiv:2309.15505, 2023
Fabian Mentzer, David Minnen, Eirikur Agustsson, and Michael Tschannen. Finite scalar quantization: Vq-vae made simple.arXiv preprint arXiv:2309.15505, 2023
-
[18]
Adaptive protein tokenization.arXiv preprint arXiv:2602.06418, 2026
Rohit Dilip, Ayush Varshney, and David Van Valen. Adaptive protein tokenization.arXiv preprint arXiv:2602.06418, 2026
-
[19]
Lei Zhu, Fangyun Wei, Yanye Lu, and Dong Chen. Scaling the codebook size of vq-gan to 100,000 with a utilization rate of 99%.Advances in Neural Information Processing Systems, 37:12612–12635, 2024
work page 2024
-
[20]
Philippe Hansen-Estruch, David Yan, Ching-Yao Chung, Orr Zohar, Jialiang Wang, Tingbo Hou, Tao Xu, Sriram Vishwanath, Peter Vajda, and Xinlei Chen. Learnings from scaling visual tokenizers for reconstruction and generation.arXiv preprint arXiv:2501.09755, 2025. 10
-
[21]
Jingfeng Yao, Bin Yang, and Xinggang Wang. Reconstruction vs. generation: Taming optimization dilemma in latent diffusion models. InProceedings of the Computer Vision and Pattern Recognition Conference, pages 15703–15712, 2025
work page 2025
-
[22]
StripedHyena: Moving Beyond Transformers with Hybrid Signal Processing Models, 12 2023
Michael Poli, Jue Wang, Stefano Massaroli, Jeffrey Quesnelle, Ryan Carlow, Eric Nguyen, and Armin Thomas. StripedHyena: Moving Beyond Transformers with Hybrid Signal Processing Models, 12 2023
work page 2023
-
[23]
Attention is all you need.Advances in neural information processing systems, 30, 2017
Ashish Vaswani, Noam Shazeer, Niki Parmar, Jakob Uszkoreit, Llion Jones, Aidan N Gomez, Łukasz Kaiser, and Illia Polosukhin. Attention is all you need.Advances in neural information processing systems, 30, 2017
work page 2017
-
[24]
Roformer: Enhanced transformer with rotary position embedding, 2021
Jianlin Su, Yu Lu, Shengfeng Pan, Bo Wen, and Yunfeng Liu. Roformer: Enhanced transformer with rotary position embedding, 2021
work page 2021
-
[25]
Flow Matching for Generative Modeling
Yaron Lipman, Ricky TQ Chen, Heli Ben-Hamu, Maximilian Nickel, and Matt Le. Flow matching for generative modeling.arXiv preprint arXiv:2210.02747, 2022
work page internal anchor Pith review Pith/arXiv arXiv 2022
-
[26]
Classifier-Free Diffusion Guidance
Jonathan Ho and Tim Salimans. Classifier-free diffusion guidance.arXiv preprint arXiv:2207.12598, 2022
work page internal anchor Pith review Pith/arXiv arXiv 2022
-
[27]
Fast and accurate protein structure search with foldseek
Michel Van Kempen, Stephanie S Kim, Charlotte Tumescheit, Milot Mirdita, Jeongjae Lee, Cameron LM Gilchrist, Johannes Söding, and Martin Steinegger. Fast and accurate protein structure search with foldseek. Nature biotechnology, 42(2):243–246, 2024
work page 2024
-
[28]
Jacob Austin, Daniel D Johnson, Jonathan Ho, Daniel Tarlow, and Rianne Van Den Berg. Structured denoising diffusion models in discrete state-spaces.Advances in neural information processing systems, 34:17981–17993, 2021
work page 2021
-
[29]
Runpeng Yu, Qi Li, and Xinchao Wang. Discrete diffusion in large language and multimodal models: A survey, 2025.URL https://arxiv. org/abs/2506.13759
-
[30]
Robust deep learning–based protein sequence design using proteinmpnn.Science, 378(6615):49–56, 2022
Justas Dauparas, Ivan Anishchenko, Nathaniel Bennett, Hua Bai, Robert J Ragotte, Lukas F Milles, Basile IM Wicky, Alexis Courbet, Rob J de Haas, Neville Bethel, et al. Robust deep learning–based protein sequence design using proteinmpnn.Science, 378(6615):49–56, 2022
work page 2022
-
[31]
Zeming Lin, Halil Akin, Roshan Rao, Brian Hie, Zhongkai Zhu, Wenting Lu, Nikita Smetanin, Robert Verkuil, Ori Kabeli, Yaniv Shmueli, et al. Evolutionary-scale prediction of atomic-level protein structure with a language model.Science, 379(6637):1123–1130, 2023
work page 2023
-
[32]
Chentong Wang, Sarah Alamdari, Carles Domingo-Enrich, Ava P Amini, and Kevin K Yang. Toward deep learning sequence–structure co-generation for protein design.Current Opinion in Structural Biology, 91:103018, 2025
work page 2025
-
[33]
Diffusion language models are versatile protein learners.arXiv preprint arXiv:2402.18567, 2024
Xinyou Wang, Zaixiang Zheng, Fei Ye, Dongyu Xue, Shujian Huang, and Quanquan Gu. Diffusion language models are versatile protein learners.arXiv preprint arXiv:2402.18567, 2024
-
[34]
Tomas Geffner, Kieran Didi, Zuobai Zhang, Danny Reidenbach, Zhonglin Cao, Jason Yim, Mario Geiger, Christian Dallago, Emine Kucukbenli, Arash Vahdat, et al. Proteina: Scaling flow-based protein structure generative models.arXiv preprint arXiv:2503.00710, 2025
-
[35]
Scaling Laws for Neural Language Models
Jared Kaplan, Sam McCandlish, Tom Henighan, Tom B Brown, Benjamin Chess, Rewon Child, Scott Gray, Alec Radford, Jeffrey Wu, and Dario Amodei. Scaling laws for neural language models.arXiv preprint arXiv:2001.08361, 2020
work page internal anchor Pith review Pith/arXiv arXiv 2001
-
[36]
Levinthal’s paradox.Proceedings of the National Academy of Sciences, 89(1):20–22, 1992
Robert Zwanzig, Attila Szabo, and Biman Bagchi. Levinthal’s paradox.Proceedings of the National Academy of Sciences, 89(1):20–22, 1992
work page 1992
-
[37]
Are there pathways for protein folding?Journal de chimie physique, 65:44–45, 1968
Cyrus Levinthal. Are there pathways for protein folding?Journal de chimie physique, 65:44–45, 1968
work page 1968
-
[38]
José Nelson Onuchic, Nicholas D Socci, Zaida Luthey-Schulten, and Peter G Wolynes. Protein folding funnels: the nature of the transition state ensemble.Folding and Design, 1(6):441–450, 1996
work page 1996
-
[39]
José Nelson Onuchic, Zaida Luthey-Schulten, and Peter G Wolynes. Theory of protein folding: the energy landscape perspective.Annual review of physical chemistry, 48(1):545–600, 1997
work page 1997
-
[40]
Protein-folding dynamics.Nature, 260(5550):404–406, 1976
Martin Karplus and David L Weaver. Protein-folding dynamics.Nature, 260(5550):404–406, 1976
work page 1976
-
[41]
Principles that govern the folding of protein chains.Science, 181(4096):223–230, 1973
Christian B Anfinsen. Principles that govern the folding of protein chains.Science, 181(4096):223–230, 1973. 11
work page 1973
-
[42]
Theory for the folding and stability of globular proteins.Biochemistry, 24(6):1501–1509, 1985
Ken A Dill. Theory for the folding and stability of globular proteins.Biochemistry, 24(6):1501–1509, 1985
work page 1985
-
[43]
Is burst hydrophobic collapse necessary for protein folding? Biochemistry, 34(9):3066–3076, 1995
AM Gutin, VI Abkevich, and EI Shakhnovich. Is burst hydrophobic collapse necessary for protein folding? Biochemistry, 34(9):3066–3076, 1995
work page 1995
-
[44]
Laura S Itzhaki, Daniel E Otzen, and Alan R Fersht. The structure of the transition state for folding of chymotrypsin inhibitor 2 analysed by protein engineering methods: evidence for a nucleation-condensation mechanism for protein folding.Journal of molecular biology, 254(2):260–288, 1995
work page 1995
-
[45]
Donald B Wetlaufer. Nucleation, rapid folding, and globular intrachain regions in proteins.Proceedings of the National Academy of Sciences, 70(3):697–701, 1973
work page 1973
-
[46]
Shi-Jie Chen, Mubashir Hassan, Robert L Jernigan, Kejue Jia, Daisuke Kihara, Andrzej Kloczkowski, Sergei Kotelnikov, Dima Kozakov, Jie Liang, Adam Liwo, et al. Protein folds vs. protein folding: Differing questions, different challenges.Proceedings of the National Academy of Sciences, 120(1):e2214423119, 2023
work page 2023
-
[47]
Yang Zhang and Jeffrey Skolnick. Scoring function for automated assessment of protein structure template quality.Proteins: Structure, Function, and Bioinformatics, 57(4):702–710, 2004
work page 2004
-
[48]
Gilles Labesse, N Colloc’h, Joël Pothier, and J-P Mornon. P-sea: a new efficient assignment of secondary structure from cαtrace of proteins.Bioinformatics, 13(3):291–295, 1997
work page 1997
-
[49]
Tomas Geffner, Kieran Didi, Zhonglin Cao, Danny Reidenbach, Zuobai Zhang, Christian Dallago, Emine Kucukbenli, Karsten Kreis, and Arash Vahdat. La-proteina: Atomistic protein generation via partially latent flow matching.arXiv preprint arXiv:2507.09466, 2025
-
[50]
Gemini: A Family of Highly Capable Multimodal Models
Gemini Team, Rohan Anil, Sebastian Borgeaud, Jean-Baptiste Alayrac, Jiahui Yu, Radu Soricut, Johan Schalkwyk, Andrew M Dai, Anja Hauth, Katie Millican, et al. Gemini: a family of highly capable multimodal models.arXiv preprint arXiv:2312.11805, 2023
work page internal anchor Pith review Pith/arXiv arXiv 2023
-
[51]
Chameleon: Mixed-Modal Early-Fusion Foundation Models
Chameleon Team. Chameleon: Mixed-modal early-fusion foundation models.arXiv preprint arXiv:2405.09818, 2024
work page internal anchor Pith review Pith/arXiv arXiv 2024
-
[52]
Eric Nguyen, Michael Poli, Matthew G Durrant, Brian Kang, Dhruva Katrekar, David B Li, Liam J Bartie, Armin W Thomas, Samuel H King, Garyk Brixi, et al. Sequence modeling and design from molecular to genome scale with evo.Science, 386(6723):eado9336, 2024
work page 2024
-
[53]
Genome modelling and design across all domains of life with evo 2.Nature, pages 1–13, 2026
Garyk Brixi, Matthew G Durrant, Jerome Ku, Mohsen Naghipourfar, Michael Poli, Gwanggyu Sun, Greg Brockman, Daniel Chang, Alison Fanton, Gabriel A Gonzalez, et al. Genome modelling and design across all domains of life with evo 2.Nature, pages 1–13, 2026
work page 2026
-
[54]
Jerome Ku, Eric Nguyen, David W Romero, Garyk Brixi, Brandon Yang, Anton V orontsov, Ali Taghibakhshi, Amy X Lu, Dave P Burke, Greg Brockman, et al. Systems and algorithms for convo- lutional multi-hybrid language models at scale.arXiv preprint arXiv:2503.01868, 2025
-
[55]
Andrew Campbell, Jason Yim, Regina Barzilay, Tom Rainforth, and Tommi Jaakkola. Generative flows on discrete state-spaces: Enabling multimodal flows with applications to protein co-design.arXiv preprint arXiv:2402.04997, 2024
-
[56]
Sidney Lyayuga Lisanza, Jacob Merle Gershon, Samuel WK Tipps, Jeremiah Nelson Sims, Lucas Arnoldt, Samuel J Hendel, Miriam K Simma, Ge Liu, Muna Yase, Hongwei Wu, et al. Multistate and functional protein design using rosettafold sequence space diffusion.Nature biotechnology, 43(8):1288–1298, 2025
work page 2025
-
[57]
Minkyung Baek, Frank DiMaio, Ivan Anishchenko, Justas Dauparas, Sergey Ovchinnikov, Gyu Rie Lee, Jue Wang, Qian Cong, Lisa N Kinch, R Dustin Schaeffer, et al. Accurate prediction of protein structures and interactions using a three-track neural network.Science, 373(6557):871–876, 2021
work page 2021
-
[58]
Generating functional and multistate proteins with a multimodal diffusion transformer.bioRxiv, 2025
Bowen Jing, Anna Sappington, Mihir Bafna, Ravi Shah, Adrina Tang, Rohith Krishna, Adam Klivans, Daniel J Diaz, and Bonnie Berger. Generating functional and multistate proteins with a multimodal diffusion transformer.bioRxiv, 2025
work page 2025
-
[59]
Scalable diffusion models with transformers
William Peebles and Saining Xie. Scalable diffusion models with transformers. InProceedings of the IEEE/CVF international conference on computer vision, pages 4195–4205, 2023
work page 2023
-
[60]
Zeming Lin, Halil Akin, Roshan Rao, Brian Hie, Zhongkai Zhu, Wenting Lu, Nikita Smetanin, Robert Verkuil, Ori Kabeli, Yaniv Shmueli, et al. Evolutionary-scale prediction of atomic-level protein structure with a language model.Science, 379(6637):1123–1130, 2023. 12
work page 2023
-
[61]
Procyon: A multimodal foundation model for protein phenotypes.BioRxiv, pages 2024–12, 2025
Owen Queen, Yepeng Huang, Robert Calef, Valentina Giunchiglia, Tianlong Chen, George Dasoulas, LeAnn Tai, Gianmarco Abbadessa, Owain Howell, Michelle M Li, et al. Procyon: A multimodal foundation model for protein phenotypes.BioRxiv, pages 2024–12, 2025
work page 2024
-
[62]
Aaron Grattafiori, Abhimanyu Dubey, Abhinav Jauhri, Abhinav Pandey, Abhishek Kadian, Ahmad Al- Dahle, Aiesha Letman, Akhil Mathur, Alan Schelten, Alex Vaughan, et al. The llama 3 herd of models. arXiv preprint arXiv:2407.21783, 2024
work page internal anchor Pith review Pith/arXiv arXiv 2024
-
[63]
arXiv preprint arXiv:2203.06125 , year=
Zuobai Zhang, Minghao Xu, Arian Jamasb, Vijil Chenthamarakshan, Aurelie Lozano, Payel Das, and Jian Tang. Protein representation learning by geometric structure pretraining.arXiv preprint arXiv:2203.06125, 2022
-
[64]
Multisensory contributions to low-level,‘unisensory’processing
Charles E Schroeder and John Foxe. Multisensory contributions to low-level,‘unisensory’processing. Current opinion in neurobiology, 15(4):454–458, 2005
work page 2005
-
[65]
E Budinger, P Heil, A Hess, and H Scheich. Multisensory processing via early cortical stages: connections of the primary auditory cortical field with other sensory systems.Neuroscience, 143(4):1065–1083, 2006. 13 Appendix Contents Tokenizer Scaling Analysis 15 Architecture Analysis . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . 15...
work page 2006
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.