Recognition: 2 theorem links
· Lean TheoremSCOPE: Siamese Contrastive Operon Pair Embeddings for Functional Sequence Representation and Classification
Pith reviewed 2026-05-13 01:40 UTC · model grok-4.3
The pith
Protein language model embeddings support operon pair classification at ROC-AUC 0.71 on the DGEB benchmark.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
By applying a Siamese MLP over fused embeddings from a protein language model, the authors obtain a ROC-AUC of 0.71 for operonic pair classification, which is competitive with state-of-the-art models on the DGEB leaderboard; protein language model embeddings therefore constitute a viable, scalable foundation for operon identification across diverse microbial genomes.
What carries the argument
A Siamese MLP head that receives the concatenated pair of protein language model embeddings and learns a binary classifier, in contrast to the DGEB benchmark's use of unsupervised cosine similarity between independently embedded sequences.
If this is right
- Protein language model embeddings can replace hand-crafted physicochemical features for operon pair classification.
- A learned fusion head is not required for competitive average precision once the embeddings are fixed.
- The same embedding approach supports genome-wide annotation in organisms without experimental operon data.
- Regulatory network reconstruction and functional annotation of unannotated genes become feasible at larger scale.
Where Pith is reading between the lines
- The same Siamese architecture could be tested on other pairwise gene relationship tasks such as predicting protein-protein interactions.
- Performance on genomes from distant phylogenetic groups would test whether the embedding geometry generalizes beyond the DGEB distribution.
- Replacing the fixed pre-trained embeddings with task-specific fine-tuning might close the small gap still present in average precision.
- Extending the method to multi-gene operons rather than pairs would require only a change in input representation.
Load-bearing premise
The DGEB benchmark and the geometry of existing protein language model embeddings together capture the functional relationships that hold in unannotated microbial genomes.
What would settle it
Apply the same trained Siamese MLP to operon pairs drawn from a newly sequenced microbial genome absent from the DGEB training and test sets and measure whether ROC-AUC falls substantially below 0.71.
Figures

read the original abstract
Identifying operons is a fundamental step in understanding prokaryotic gene regulation, as classifying genes into operons supports the reconstruction of regulatory networks, functional annotation of unannotated genes, and drug candidate development. Experimental approaches such as RT-PCR and RNA-seq provide precise evidence of operon structure, but are laborious and largely limited to well-studied model organisms, making scalable computational methods essential for genome-wide operon identification. Prior computational approaches have employed traditional classifiers such as logistic regression and decision trees, motivating our use of these as physicochemical baselines. The DGEB benchmark evaluates operonic pair classification by embedding each sequence independently with a pre-trained protein language model and computing pairwise cosine similarity. In contrast, our Siamese MLP learns a classifier over the fused embedding space, which is theoretically better motivated for binary classification, as cosine similarity can yield meaningless scores depending on the regularization of the embedding model. While protein language model embeddings substantially outperform physicochemical features in ROC-AUC, a learned Siamese MLP head does not significantly improve over unsupervised cosine similarity in Average Precision, suggesting that the geometry of the embedding space already captures the functional relationships needed for this task. Nonetheless, our Siamese MLP achieves a ROC-AUC of 0.71, competitive with state-of-the-art models on the DGEB leaderboard. These findings indicate that protein language model embeddings are a viable, scalable foundation for operonic pair classification across diverse microbial genomes, with implications for automated genome annotation, regulatory network reconstruction, and characterization of organisms lacking experimental operon annotations.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript presents SCOPE, a Siamese MLP trained on fused embeddings from pre-trained protein language models to classify operonic gene pairs. Evaluated on the DGEB benchmark, it reports a ROC-AUC of 0.71 that is competitive with existing models; the authors observe that this learned head yields no significant Average Precision gain over unsupervised cosine similarity on the same embeddings and conclude that PLM embedding geometry already encodes the necessary functional relationships, making such embeddings a viable, scalable foundation for operon classification across diverse microbial genomes.
Significance. If substantiated, the result would offer a computationally lightweight route to operon inference in unannotated prokaryotic genomes, supporting downstream tasks such as regulatory-network reconstruction and functional annotation. The observation that cosine similarity already suffices is potentially useful, as it implies that additional supervised training may be unnecessary for this particular task. The work is limited, however, by the absence of dataset statistics, training protocols, and generalization tests that would allow readers to assess whether the DGEB performance translates to phylogenetically distant or poorly annotated genomes.
major comments (3)
- [Abstract] Abstract: the central performance claim (ROC-AUC = 0.71, competitive with SOTA on DGEB) is presented without any accompanying information on the number of positive/negative pairs, class balance, train/validation/test splits, number of independent runs, or statistical tests; without these quantities the competitiveness statement cannot be evaluated.
- [Abstract] Abstract: the claim that 'protein language model embeddings are a viable, scalable foundation for operonic pair classification across diverse microbial genomes' rests entirely on DGEB results, yet the manuscript supplies no cross-genome hold-out, phylogenetic-distance analysis, or external validation set; this gap directly affects the scalability conclusion.
- [Abstract] Abstract: the statement that the Siamese MLP is 'theoretically better motivated' than cosine similarity because the latter 'can yield meaningless scores depending on the regularization of the embedding model' is asserted without a supporting derivation, reference, or empirical demonstration that cosine scores are in fact uninformative under the regularization used by the PLMs in the study.
minor comments (1)
- [Abstract] The abstract refers to 'physicochemical baselines' (logistic regression, decision trees) but does not list the specific features or preprocessing steps employed; adding this detail would improve reproducibility.
Simulated Author's Rebuttal
We thank the referee for their constructive feedback, which has improved the clarity of our claims and the transparency of our methods. We address each major comment below, indicating revisions where the manuscript has been updated.
read point-by-point responses
-
Referee: [Abstract] Abstract: the central performance claim (ROC-AUC = 0.71, competitive with SOTA on DGEB) is presented without any accompanying information on the number of positive/negative pairs, class balance, train/validation/test splits, number of independent runs, or statistical tests; without these quantities the competitiveness statement cannot be evaluated.
Authors: We agree that these details are necessary to evaluate the performance claim. The revised manuscript updates the abstract with a concise reference to the evaluation protocol and adds a dedicated subsection in Methods detailing the DGEB pair counts, class balance, cross-validation splits, number of independent runs, and statistical comparisons. A summary table of these quantities has also been added to the Results section. revision: yes
-
Referee: [Abstract] Abstract: the claim that 'protein language model embeddings are a viable, scalable foundation for operonic pair classification across diverse microbial genomes' rests entirely on DGEB results, yet the manuscript supplies no cross-genome hold-out, phylogenetic-distance analysis, or external validation set; this gap directly affects the scalability conclusion.
Authors: We acknowledge that stronger generalization evidence would strengthen the scalability statement. DGEB already aggregates pairs from phylogenetically diverse genomes, providing initial support. In revision we added a cross-genome hold-out experiment (training on one set of genomes and testing on held-out genomes) with results reported in a new subsection. A full phylogenetic-distance analysis is noted as a limitation and future direction, as it requires metadata and computations outside the current benchmark scope. revision: partial
-
Referee: [Abstract] Abstract: the statement that the Siamese MLP is 'theoretically better motivated' than cosine similarity because the latter 'can yield meaningless scores depending on the regularization of the embedding model' is asserted without a supporting derivation, reference, or empirical demonstration that cosine scores are in fact uninformative under the regularization used by the PLMs in the study.
Authors: We accept that the original wording required substantiation. The revised abstract now states that the Siamese MLP offers a flexible learned alternative rather than claiming theoretical superiority. The Introduction and Methods sections now include a short explanation with citations to literature on embedding regularization in protein language models, together with an empirical comparison of cosine-score stability across PLM checkpoints in the supplementary material. revision: yes
Circularity Check
No significant circularity; derivation is self-contained empirical evaluation
full rationale
The paper describes training a Siamese MLP on labeled operonic pairs using pre-trained protein language model embeddings and evaluating it with standard metrics (ROC-AUC, Average Precision) against the external DGEB benchmark and physicochemical baselines. No equations, self-definitional loops, fitted parameters renamed as predictions, or load-bearing self-citations appear in the provided text; the central claim follows directly from benchmark performance without reducing to its own inputs by construction. The geometry-capture observation is a post-hoc empirical note, not a definitional equivalence.
Axiom & Free-Parameter Ledger
free parameters (1)
- Siamese MLP weights
axioms (1)
- domain assumption The DGEB benchmark constitutes a reliable and generalizable test for operonic pair classification across microbial genomes
Lean theorems connected to this paper
-
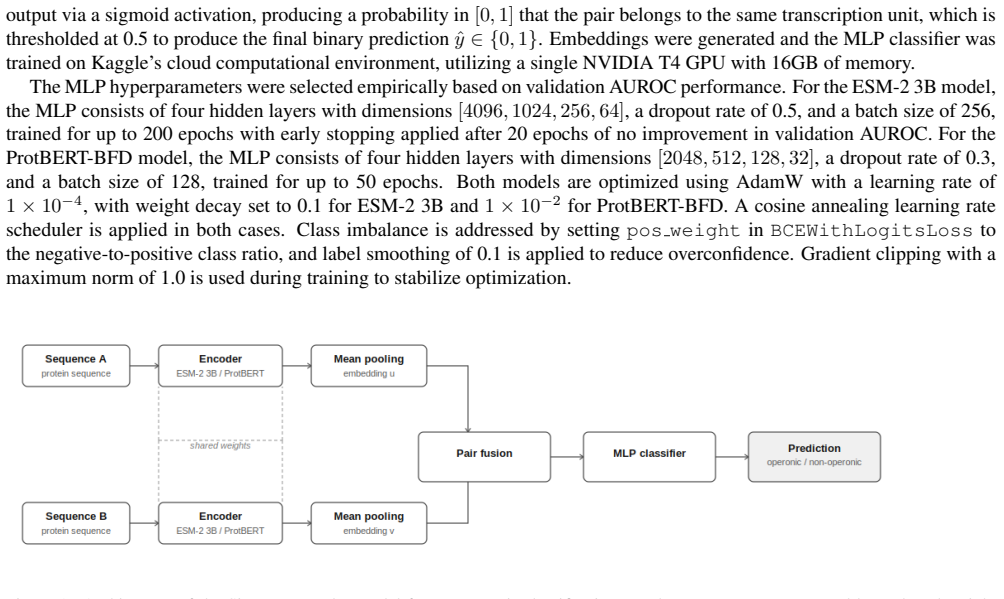
IndisputableMonolith/Cost/FunctionalEquation.leanwashburn_uniqueness_aczel unclearOur encoder-based architecture consists of Siamese transformer models that generate embeddings for each sequence, which are then mean-pooled... fused via concatenation, signed difference, absolute difference, and element-wise product, before being passed to an MLP
-
IndisputableMonolith/Foundation/AlphaCoordinateFixation.leanJ_uniquely_calibrated_via_higher_derivative unclearour Siamese MLP achieves a ROC-AUC of 0.71, competitive with state-of-the-art models on the DGEB leaderboard
Reference graph
Works this paper leans on
-
[1]
Ahmed Elnaggar, Michael Heinzinger, Christian Dallago, Ghalia Rehawi, Yu Wang, Llion Jones, Tom Gibbs, Tamas Feher, Christoph Angerer, Martin Steinegger, Debsindhu Bhowmik, and Burkhard Rost. ProtTrans: Toward understanding the language of life through self-supervised learning.IEEE Transactions on Pattern Analysis and Machine Intelligence, 44(10):7112–712...
work page 2021
-
[2]
Raga Krishnakumar and Anne M. Ruffing. OperonSEQer: A set of machine-learning algorithms with threshold voting for detection of operon pairs using short-read RNA-sequencing data.PLOS Computational Biology, 18(1):e1009731, 2022. 1
work page 2022
-
[3]
Zeming Lin, Halil Akin, Roshan Rao, Brian Hie, Zhongkai Zhu, Wenting Lu, Nikita Smetanin, Robert Verkuil, Ori Kabeli, Yaniv Shmueli, Allan dos Santos Costa, Maryam Fazel-Zarandi, Tom Sercu, Salvatore Candido, and Alexander Rives. Evolutionary-scale prediction of atomic-level protein structure with a language model.Science, 379(6637):1123–1130, 2023. 1, 2
work page 2023
-
[4]
Gabriel Moreno-Hagelsieb. The power of operon rearrangements for predicting functional associations.Computational and Structural Biotechnology Journal, 13:402–406, 2015. Epub 2015 Jul 2. 1, 3
work page 2015
- [5]
-
[6]
Hatice Ulku Osmanbeyoglu and Madhavi K. Ganapathiraju. N-gram analysis of 970 microbial organisms reveals presence of biolog- ical language models.BMC Bioinformatics, 11:131, 2010. 1
work page 2010
-
[7]
Greenbaum, Hannah Carter, Morten Nielsen, and Bjoern Peters
Eve Richardson, Raphael Trevizani, Jason A. Greenbaum, Hannah Carter, Morten Nielsen, and Bjoern Peters. The receiver operating characteristic curve accurately assesses imbalanced datasets.Patterns, 2024. 3
work page 2024
-
[8]
Harald Steck, Chaitanya Ekanadham, and Nathan Kallus. Is cosine-similarity of embeddings really about similarity? InCompanion Proceedings of the ACM Web Conference 2024, 2024. 1
work page 2024
-
[9]
Diverse genomic embedding benchmark for functional evaluation across the tree of life.bioRxiv, 2024
Jacob West-Roberts, Joshua Kravitz, Nishant Jha, Andre Cornman, and Yunha Hwang. Diverse genomic embedding benchmark for functional evaluation across the tree of life.bioRxiv, 2024. 1, 3, 4
work page 2024
-
[10]
Bradley P. Westover, Jeremy D. Buhler, Justin L. Sonnenburg, and Jeffrey I. Gordon. Operon prediction without a training set. Bioinformatics, 21(5):880–888, 2005. 1 5
work page 2005
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.