Recognition: no theorem link
NexOP: Joint Optimization of NEX-Aware k-space Sampling and Image Reconstruction for Low-Field MRI
Pith reviewed 2026-05-13 01:47 UTC · model grok-4.3
The pith
NexOP jointly optimizes sampling across k-space and repetitions with reconstruction to raise SNR in low-field MRI.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
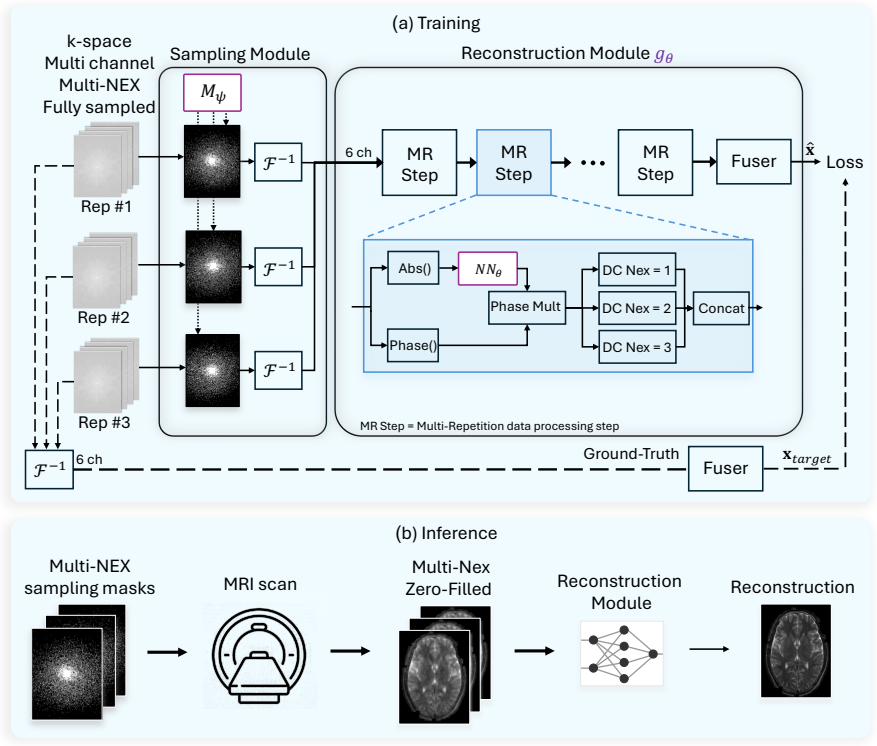
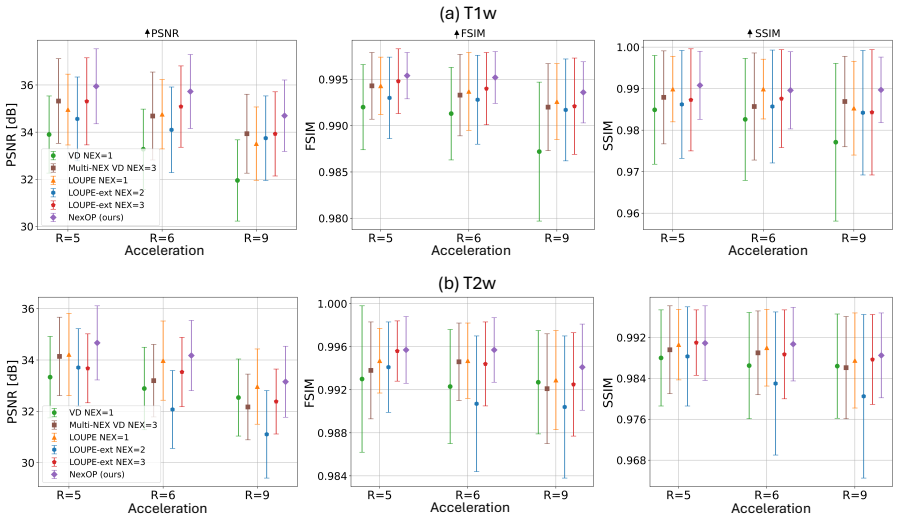
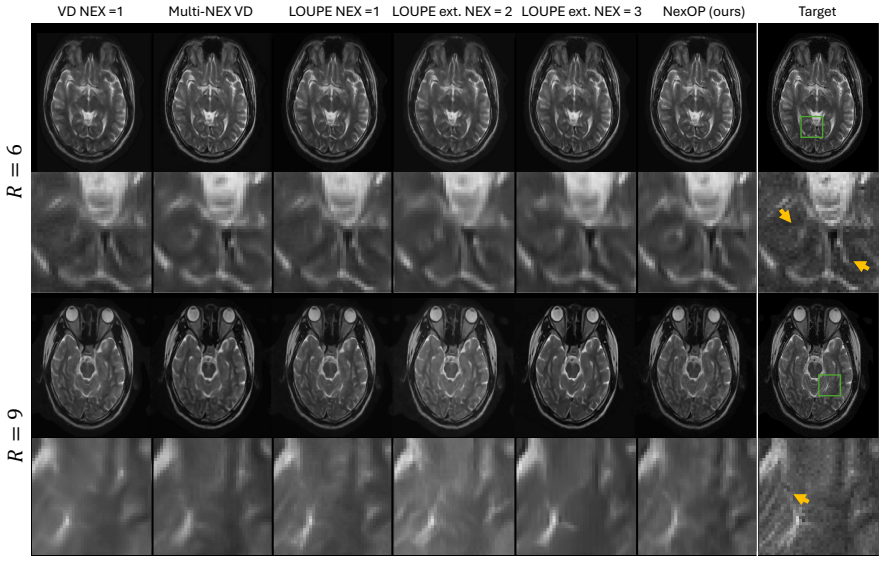
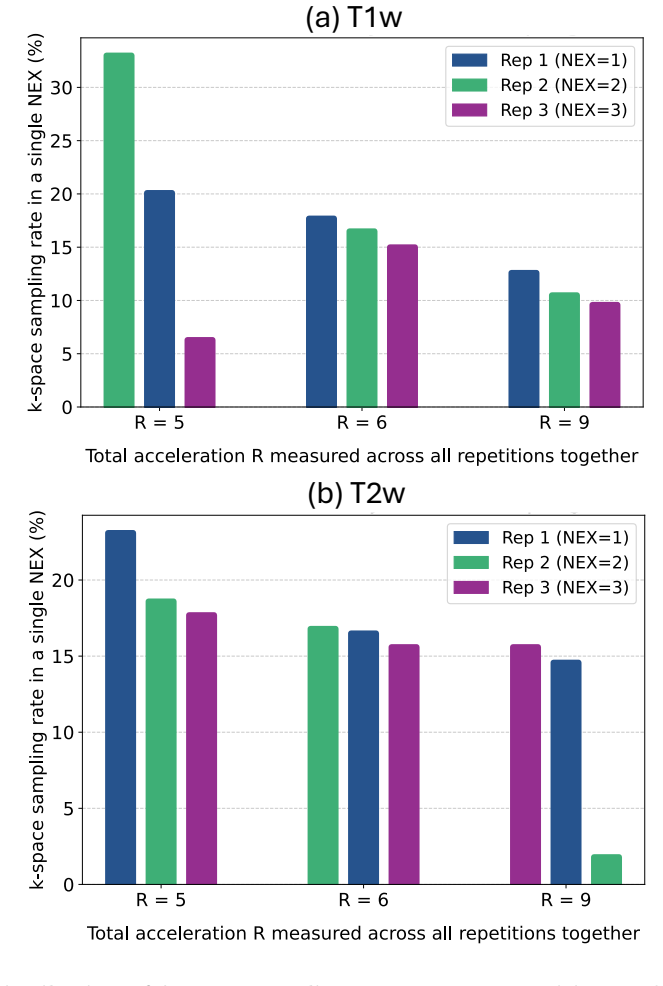
NexOP optimizes sampling density probabilities across the extended k-space-NEX domain under a fixed sampling-budget constraint and introduces a deep-learning architecture for reconstructing a single high-SNR image from multiple low-SNR measurements. Experiments with raw 0.3 T brain data show consistent quantitative and qualitative gains over competing methods across acceleration factors and contrasts. The learned patterns are non-uniform, with sampling density decreasing across repetitions, and a theoretical analysis supports this behavior.
What carries the argument
NexOP framework that jointly learns NEX-aware k-space sampling densities under a fixed budget and reconstructs from multiple low-SNR acquisitions via a dedicated deep neural network.
If this is right
- Learned sampling densities decrease across successive repetitions instead of staying uniform.
- Image quality exceeds that of pipelines that optimize sampling and reconstruction independently.
- Higher SNR is achieved within the same total acquisition budget.
- The approach is directly applicable to portable low-cost MRI hardware.
Where Pith is reading between the lines
- The same joint-optimization idea could be tested on other low-SNR modalities that use repeated acquisitions.
- Pre-computing the learned sampling masks offline would allow real-time use without retraining per patient.
- Pairing the method with scanner-specific noise models might further improve performance across hardware variants.
Load-bearing premise
An end-to-end neural network can discover sampling and reconstruction strategies that generalize beyond the 0.3 T brain data used for training.
What would settle it
Apply a trained NexOP model to brain data acquired at a different field strength or with tissue contrasts absent from training and check whether the resulting images have lower quality metrics than those from uniform NEX sampling.
Figures




read the original abstract
Modern low-field magnetic resonance imaging (MRI) technology offers a compelling alternative to standard high-field MRI, with portable, low-cost systems. However, its clinical utility is limited by a low Signal-to-Noise Ratio (SNR), which hampers diagnostic image quality. A common approach to increase SNR is through repetitive signal acquisitions, known as NEX, but this results in excessively long scan durations. Although recent work has introduced methods to accelerate MRI scans through k-space sampling optimization, the NEX dimension remains unexploited; typically, a single sampling mask is used across all repetitions. Here we introduce NexOP, a deep-learning framework for joint optimization of the sampling and reconstruction in multi-NEX acquisitions, tailored for low-SNR settings. NexOP enables optimizing the sampling density probabilities across the extended k-space-NEX domain, under a fixed sampling-budget constraint, and introduces a new deep-learning architecture for reconstructing a single high-SNR image from multiple low-SNR measurements. Experiments with raw low-field (0.3T) brain data demonstrate that NexOP consistently outperforms competing methods, both quantitatively and qualitatively, across diverse acceleration factors and tissue contrasts. The results also demonstrate that NexOP yields non-uniform sampling strategies, with progressively decreasing sampling across repetitions, hence exploiting the NEX dimension efficiently. Moreover, we present a theoretical analysis supporting these numerical observations. Overall, this work proposes a sampling-reconstruction optimization framework highly suitable for low-field MRI, which can enable faster, higher-quality imaging with low-cost systems and contribute to advancing affordable and accessible healthcare.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper introduces NexOP, a deep-learning framework for joint optimization of NEX-aware k-space sampling and image reconstruction in low-field MRI. It optimizes sampling density probabilities over the extended k-space-NEX domain under a fixed sampling-budget constraint, proposes a new architecture to reconstruct a single high-SNR image from multiple low-SNR multi-NEX measurements, and reports consistent outperformance on raw 0.3 T brain data across acceleration factors and tissue contrasts. The work also presents non-uniform learned sampling patterns (decreasing density across repetitions) and supporting theoretical analysis.
Significance. If the central claims hold, NexOP would represent a practical advance for low-field MRI by exploiting the NEX dimension to improve SNR without extending scan time, potentially increasing the clinical utility of portable low-cost systems. Strengths include the use of real raw 0.3 T data, the joint end-to-end formulation, and the theoretical analysis that aligns with the observed non-uniform sampling strategies.
major comments (2)
- [Methods] Methods section on the sampling optimization: the parameterization of NEX-aware sampling probabilities and the exact mechanism enforcing the fixed budget constraint during end-to-end training are not specified with sufficient mathematical detail; without this, it is difficult to verify that the learned non-uniform patterns are not an artifact of the constraint implementation.
- [Results] Results section and tables: while consistent outperformance is claimed, the manuscript lacks reported quantitative metrics (e.g., PSNR, SSIM values with error bars), ablation studies isolating the joint optimization benefit, and statistical significance tests across the diverse acceleration factors and contrasts; these omissions weaken the ability to assess the practical magnitude of the reported gains.
minor comments (3)
- [Abstract] The abstract and introduction would benefit from a brief explicit statement of the loss function used for joint training.
- [Figures] Figure captions for the sampling pattern visualizations should include the exact acceleration factors and NEX values corresponding to each panel.
- [Introduction] Notation for the extended k-space-NEX domain could be introduced earlier and used consistently to improve readability.
Simulated Author's Rebuttal
We thank the referee for the positive assessment and recommendation for minor revision. The comments are constructive and will help strengthen the manuscript. We address each major comment below.
read point-by-point responses
-
Referee: [Methods] Methods section on the sampling optimization: the parameterization of NEX-aware sampling probabilities and the exact mechanism enforcing the fixed budget constraint during end-to-end training are not specified with sufficient mathematical detail; without this, it is difficult to verify that the learned non-uniform patterns are not an artifact of the constraint implementation.
Authors: We agree that greater mathematical detail is needed for reproducibility and to rule out implementation artifacts. In the revised manuscript we will expand the Methods section to explicitly define the NEX-aware sampling probabilities as a learnable tensor P(k,n) over the joint k-space–NEX domain, with normalization performed via a temperature-scaled softmax to guarantee valid probabilities. We will also describe the exact budget-enforcement mechanism: a differentiable projection operator applied after each gradient step that rescales the probabilities so that their sum exactly equals the prescribed sampling budget B. This formulation makes clear that the observed non-uniform patterns (decreasing density across repetitions) arise from the joint optimization objective rather than from the constraint itself. revision: yes
-
Referee: [Results] Results section and tables: while consistent outperformance is claimed, the manuscript lacks reported quantitative metrics (e.g., PSNR, SSIM values with error bars), ablation studies isolating the joint optimization benefit, and statistical significance tests across the diverse acceleration factors and contrasts; these omissions weaken the ability to assess the practical magnitude of the reported gains.
Authors: We acknowledge that more granular quantitative reporting would strengthen the results. Although the manuscript already presents quantitative comparisons, we will revise the Results section and tables to include PSNR and SSIM values reported as mean ± standard deviation across the test subjects for every acceleration factor and contrast. We will add ablation studies that isolate the benefit of joint optimization (e.g., fixed versus learned sampling with the identical reconstruction network) and will include statistical significance tests (paired t-tests or Wilcoxon signed-rank tests with p-values) for the observed improvements. These additions will allow readers to better gauge the practical magnitude of the gains. revision: yes
Circularity Check
No significant circularity; derivation is self-contained
full rationale
The paper introduces a new end-to-end deep-learning framework (NexOP) that jointly optimizes NEX-aware k-space sampling probabilities and a reconstruction network under a fixed budget constraint, trained directly on raw 0.3T brain data. No load-bearing step reduces by the paper's own equations or self-citations to a prior fitted quantity; the non-uniform sampling patterns and SNR gains are reported as empirical outcomes of the optimization on held-out acquisitions, not as algebraic identities or renamings of inputs. The theoretical analysis is presented as supporting numerical observations rather than as a uniqueness theorem imported from the authors' prior work. The central claim therefore rests on experimental validation rather than definitional closure.
Axiom & Free-Parameter Ledger
invented entities (1)
-
NexOP deep-learning architecture
no independent evidence
Reference graph
Works this paper leans on
-
[1]
Sairam Geethanath and John Thomas Vaughan Jr. Accessible magnetic resonance imaging: A review.Journal of Magnetic Resonance Imaging, 49(7):e65–e77, 2019
work page 2019
-
[2]
Low-cost high-performance MRI.Scientific Reports, 5(1): 15177, 2015
Mathieu Sarracanie, Cristen D LaPierre, Najat Salameh, David EJ Waddington, Thomas Witzel, and Matthew S Rosen. Low-cost high-performance MRI.Scientific Reports, 5(1): 15177, 2015
work page 2015
-
[3]
Jos´ e P Marques, Frank FJ Simonis, and Andrew G Webb. Low-field MRI: An MR physics perspective.Journal of Magnetic Resonance Imaging, 49(6):1528–1542, 2019
work page 2019
-
[4]
Adrienne E Campbell-Washburn, Kathryn E Keenan, Peng Hu, John P Mugler III, Kr- ishna S Nayak, Andrew G Webb, Johnes Obungoloch, Kevin N Sheth, J¨ urgen Hennig, Matthew S Rosen, et al. Low-field MRI: a report on the 2022 ISMRM workshop.Magnetic Resonance in Medicine, 90(4):1682–1694, 2023. 16
work page 2022
-
[5]
W. Taylor Kimberly, Annabel J. Sorby-Adams, Andrew G. Webb, Ed X. Wu, Rachel Beekman, Ritvij Bowry, Steven J. Schiff, Adam de Havenon, Francis X. Shen, Gordon Sze, Pamela Schaefer, Juan Eugenio Iglesias, Matthew S. Rosen, and Kevin N. Sheth. Brain imaging with portable low-field MRI.Nature Reviews Bioengineering, 1, 2023
work page 2023
-
[6]
Teresa Guallart-Naval, Jos´ e M Algar´ ın, Rub´ en Pellicer-Guridi, Fernando Galve, Yolanda Vives-Gilabert, Rub´ en Bosch, Eduardo Pall´ as, Jos´ e M Gonz´ alez, Juan P Rigla, Pablo Mart´ ınez, et al. Portable magnetic resonance imaging of patients indoors, outdoors and at home.Scientific Reports, 12(1):13147, 2022
work page 2022
-
[7]
Whole-body magnetic resonance imaging at 0.05 Tesla.Science, 384(6696), 2024
Yujiao Zhao, Ye Ding, Vick Lau, Christopher Man, Shi Su, Linfang Xiao, Alex TL Leong, and Ed X Wu. Whole-body magnetic resonance imaging at 0.05 Tesla.Science, 384(6696), 2024
work page 2024
-
[8]
MRI at low field: a review of software solutions for improving SNR.NMR in Biomedicine, 38(1), 2025
Reina Ayde, Marc Vornehm, Yujiao Zhao, Florian Knoll, Ed X Wu, and Mathieu Sar- racanie. MRI at low field: a review of software solutions for improving SNR.NMR in Biomedicine, 38(1), 2025
work page 2025
-
[9]
Accelerating Low-field MRI: Compressed Sensing and AI for fast noise-robust imaging
Efrat Shimron, Shanshan Shan, James Grover, Neha Koonjoo, Sheng Shen, Thomas Boele, Annabel J Sorby-Adams, John E Kirsch, Matthew S Rosen, and David EJ Waddington. Accelerating Low-field MRI: Compressed Sensing and AI for fast noise-robust imaging. arXiv preprint arXiv:2411.06704, 2024. URLhttps://arxiv.org/html/2411.06704v1
-
[10]
Tal Oved, Beatrice Lena, Chlo ˜AˇS F Najac, Sheng Shen, Matthew S Rosen, Andrew Webb, and Efrat Shimron. Deep learning of personalized priors from past MRI scans enables fast, quality-enhanced point-of-care MRI with low-cost systems.arXiv preprint arXiv:2505.02470, 2025
-
[11]
David I Hoult and RE Richards. The signal-to-noise ratio of the nuclear magnetic resonance experiment.Journal of Magnetic Resonance (1969), 24(1):71–85, 1976
work page 1969
-
[12]
Progress toward a deployable SQUID-based ultra-low field MRI system for anatomical imaging
Michelle A Espy, Per E Magnelind, Andrei N Matlashov, Shaun G Newman, Henrik J Sandin, Larry J Schultz, Robert Sedillo, Algis V Urbaitis, and Petr L Volegov. Progress toward a deployable SQUID-based ultra-low field MRI system for anatomical imaging. IEEE Transactions on Applied Superconductivity, 25(3):1–5, 2014
work page 2014
-
[13]
Mark A Griswold, Peter M Jakob, Robin M Heidemann, Mathias Nittka, Vladimir Jellus, Jianmin Wang, Berthold Kiefer, and Axel Haase. Generalized autocalibrating partially parallel acquisitions (GRAPPA).Magnetic Resonance in Medicine: An Official Journal of the International Society for Magnetic Resonance in Medicine, 47(6):1202–1210, 2002
work page 2002
-
[14]
Klaas P Pruessmann, Markus Weiger, Markus B Scheidegger, and Peter Boesiger. Sense: sensitivity encoding for fast mri.Magnetic Resonance in Medicine: An Official Journal of the International Society for Magnetic Resonance in Medicine, 42(5):952–962, 1999
work page 1999
-
[15]
Michael Lustig and John M Pauly. SPIRiT: iterative self-consistent parallel imaging recon- struction from arbitrary k-space.Magnetic resonance in medicine, 64(2):457–471, 2010
work page 2010
-
[16]
Efrat Shimron, Andrew G Webb, and Haim Azhari. CORE-PI: Non-iterative convolution- based reconstruction for parallel MRI in the wavelet domain.Medical Physics, 46(1): 199–214, 2019
work page 2019
-
[17]
Michael Lustig, David Donoho, and John M. Pauly. Sparse MRI: The application of compressed sensing for rapid MR imaging.Magnetic Resonance in Medicine, 58, 2007. ISSN 07403194. 17
work page 2007
-
[18]
Compressed sensing MRI: a review.Critical Reviews in Biomedical Engineering, 41(3), 2013
Sairam Geethanath, Rashmi Reddy, Amaresha Shridhar Konar, Shaikh Imam, Rajagopalan Sundaresan, Ramesh Babu DR, and Ramesh Venkatesan. Compressed sensing MRI: a review.Critical Reviews in Biomedical Engineering, 41(3), 2013
work page 2013
-
[19]
Reinhard Heckel, Mathews Jacob, Akshay Chaudhari, Or Perlman, and Efrat Shimron. Deep learning for accelerated and robust MRI reconstruction.Magnetic Resonance Mate- rials in Physics, Biology and Medicine 2024 37:3, 37:335–368, 7 2024. ISSN 1352-8661
work page 2024
-
[20]
Kerstin Hammernik, Thomas K¨ ustner, Burhaneddin Yaman, Zhengnan Huang, Daniel Rueckert, Florian Knoll, and Mehmet Ak¸ cakaya. Physics-driven deep learning for computa- tional magnetic resonance imaging: Combining physics and machine learning for improved medical imaging.IEEE signal processing magazine, 40(1):98–114, 2023
work page 2023
-
[21]
Kunal Aggarwal, Marina Manso Jimeno, Keerthi Sravan Ravi, Gilberto Gonzalez, and Sairam Geethanath. Developing and deploying deep learning models in brain magnetic resonance imaging: A review.NMR in Biomedicine, 36, 12 2023. ISSN 1099-1492. URL https://pubmed.ncbi.nlm.nih.gov/37539775/
-
[22]
Bo Zhou, Jo Schlemper, Neel Dey, Seyed Sadegh Mohseni Salehi, Kevin Sheth, Chi Liu, James S Duncan, and Michal Sofka. Dual-domain self-supervised learning for accelerated non-Cartesian MRI reconstruction.Medical Image Analysis, 81:102538, 2022
work page 2022
-
[23]
Deep learning enabled fast 3D brain MRI at 0.055 tesla.Science Advances, 9(38), 2023
Christopher Man, Vick Lau, Shi Su, Yujiao Zhao, Linfang Xiao, Ye Ding, Gilberto KK Leung, Alex TL Leong, and Ed X Wu. Deep learning enabled fast 3D brain MRI at 0.055 tesla.Science Advances, 9(38), 2023
work page 2023
-
[24]
N. Koonjoo, B. Zhu, G. Cody Bagnall, D. Bhutto, and M. S. Rosen. Boosting the signal- to-noise of low-field MRI with deep learning image reconstruction.Scientific Reports, 11: 8248, 2021
work page 2021
-
[25]
Available: https://arxiv.org/abs/2509.11790
Efe Ilıcak, Chinmay Rao, Chlo´ e Najac, Beatrice Lena, Baris Imre, Fernando Galve, Joseba Alonso, Andrew Webb, and Marius Staring. Physics-Informed Deep Unrolled Network for Portable MR Image Reconstruction.arXiv preprint arXiv:2509.11790, 2025
-
[26]
Efrat Shimron and Or Perlman. AI in MRI: computational frameworks for a faster, opti- mized, and automated imaging workflow.Bioengineering, 10(4):492, 2023
work page 2023
-
[27]
Cagla D Bahadir, Alan Q Wang, Adrian V Dalca, and Mert R Sabuncu. Deep-learning- based optimization of the under-sampling pattern in MRI.IEEE Transactions on Compu- tational Imaging, 6:1139–1152, 2020
work page 2020
-
[28]
GR Chaithya, Pierre Weiss, Guillaume Daval-Fr´ erot, Aur´ elien Massire, Alexandre Vignaud, and Philippe Ciuciu. Optimizing full 3d sparkling trajectories for high-resolution magnetic resonance imaging.IEEE Transactions on Medical Imaging, 41(8):2105–2117, 2022
work page 2022
-
[29]
Hemant Kumar Aggarwal and Mathews Jacob. J-MoDL: Joint Model-Based Deep Learning for Optimized Sampling and Reconstruction.IEEE Journal on Selected Topics in Signal Processing, 14:1151–1162, 7 2020
work page 2020
-
[30]
Tomer Weiss, Ortal Senouf, Sanketh Vedula, Oleg Michailovich, Michael Zibulevsky, and Alex Bronstein. PILOT: Physics-Informed Learned Optimized Trajectories for Accelerated MRI.Machine Learning for Biomedical Imaging, 1:1–23, 4 2021
work page 2021
-
[31]
Op- timizing sampling patterns for compressed sensing MRI with diffusion generative models
Sriram Ravula, Brett Levac, Ajil Jalal, Jonathan I Tamir, and Alexandros G Dimakis. Op- timizing sampling patterns for compressed sensing MRI with diffusion generative models. arXiv preprint arXiv:2306.03284, 2023. 18
-
[32]
Chaithya Giliyar Radhakrishna and Philippe Ciuciu. Jointly learning non-cartesian k-space trajectories and reconstruction networks for 2d and 3d mr imaging through projection. Bioengineering, 10(2):158, 2023
work page 2023
-
[33]
Scan-Adaptive MRI Undersampling Using Neighbor-based Optimization (SUNO)
Siddhant Gautam, Student Member, Angqi Li, Nicole Seiberlich, Jeffrey A Fessler, Saiprasad Ravishankar, and Senior Member. Scan-Adaptive MRI Undersampling Using Neighbor-based Optimization (SUNO). 6 2025
work page 2025
-
[34]
Cagan Alkan, Morteza Mardani, Shreyas S Vasanawala, and John M Pauly. Learning to Sample MRI via Variational Information Maximization.In NeurIPS 2020 Workshop on Deep Learning and Inverse Problems, 2020
work page 2020
-
[35]
Florian Knoll, Christian Clason, Clemens Diwoky, and Rudolf Stollberger. Adapted random sampling patterns for accelerated mri.Magnetic resonance materials in physics, biology and medicine, 24(1):43–50, 2011
work page 2011
-
[36]
Nicolas Chauffert, Philippe Ciuciu, Jonas Kahn, and Pierre Weiss. Variable density sam- pling with continuous trajectories.SIAM Journal on Imaging Sciences, 7(4):1962–1992, 2014
work page 1962
-
[37]
Learning the sampling density in 2d sparkling mri acquisition for optimized image reconstruction
GR Chaithya, Zaccharie Ramzi, and Philippe Ciuciu. Learning the sampling density in 2d sparkling mri acquisition for optimized image reconstruction. In2021 29th European Signal Processing Conference (EUSIPCO), pages 960–964. IEEE, 2021
work page 2021
-
[38]
Jasper Schoormans, Gustav J. Strijkers, Anders C. Hansen, Aart J. Nederveen, and Bram F. Coolen. Compressed sensing MRI with variable density averaging (CS-VDA) outperforms full sampling at low SNR.Physics in Medicine & Biology, 65:045004, 2 2020. ISSN 0031-9155
work page 2020
-
[39]
Matt A Bernstein, Kevin F King, and Xiaohong Joe Zhou.Handbook of MRI pulse se- quences. Elsevier, 2004
work page 2004
-
[40]
Parallel MR imaging.Journal of Magnetic Resonance Imaging, 36(1):55–72, 2012
Anagha Deshmane, Vikas Gulani, Mark A Griswold, and Nicole Seiberlich. Parallel MR imaging.Journal of Magnetic Resonance Imaging, 36(1):55–72, 2012
work page 2012
-
[41]
Kieren Grant Hollingsworth. Reducing acquisition time in clinical MRI by data under- sampling and compressed sensing reconstruction.Physics in Medicine & Biology, 60(21): R297–R322, 2015
work page 2015
-
[42]
Chi-Ming Tsai and Dwight G Nishimura. Reduced aliasing artifacts using variable-density k-space sampling trajectories.Magnetic Resonance in Medicine: An Official Journal of the International Society for Magnetic Resonance in Medicine, 43(3):452–458, 2000
work page 2000
-
[43]
Variable density compressed sensing in MRI
Nicolas Chauffert, Philippe Ciuciu, and Pierre Weiss. Variable density compressed sensing in MRI. Theoretical vs heuristic sampling strategies. In2013 IEEE 10th International Symposium on Biomedical Imaging, pages 298–301. IEEE, 2013
work page 2013
-
[44]
Image reconstruction by domain-transform manifold learning.Nature, 555(7697):487–492, 2018
Bo Zhu, Jeremiah Z Liu, Stephen F Cauley, Bruce R Rosen, and Matthew S Rosen. Image reconstruction by domain-transform manifold learning.Nature, 555(7697):487–492, 2018
work page 2018
-
[45]
Kerstin Hammernik, Teresa Klatzer, Erich Kobler, Michael P. Recht, Daniel K. Sodickson, Thomas Pock, and Florian Knoll. Learning a variational network for reconstruction of accelerated MRI data.Magnetic Resonance in Medicine, 79, 2017. URLhttps://api. semanticscholar.org/CorpusID:3815411. 19
work page 2017
-
[46]
Hemant K Aggarwal, Merry P Mani, and Mathews Jacob. MoDL: Model-based deep learning architecture for inverse problems.IEEE Transactions on Medical Imaging, 38(2): 394–405, 2018
work page 2018
-
[47]
Rishabh Sharma, Panagiotis Tsiamyrtzis, Andrew G. Webb, Ioannis Seimenis, Constanti- nos Loukas, Ernst Leiss, and Nikolaos V. Tsekos. A Deep Learning Approach to Upscaling “Low-Quality” MR Images: An In Silico Comparison Study Based on the UNet Framework. Applied Sciences 2022, Vol. 12, Page 11758, 12:11758, 11 2022. ISSN 2076-3417
work page 2022
-
[48]
Efrat Shimron, William Grissom, and Haim Azhari. Temporal differences (TED) com- pressed sensing: A method for fast MRgHIFU temperature imaging.NMR in Biomedicine, 33(9), 2020
work page 2020
-
[49]
Hongyi Gu, Burhaneddin Yaman, Steen Moeller, Jutta Ellermann, Kamil Ugurbil, and Mehmet Ak¸ cakaya. Revisitingℓ1-wavelet compressed-sensing mri in the era of deep learn- ing.Proceedings of the National Academy of Sciences, 119(33), 2022
work page 2022
-
[50]
Yilmaz Korkmaz, Mahmut Yurt, Salman Ul Hassan Dar, Muzaffer ¨Ozbey, and Tolga Cukur. Deep MRI Reconstruction with Generative Vision Transformers.Lecture Notes in Com- puter Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics), 12964:54–64, 2021. ISSN 16113349
work page 2021
-
[51]
Bilal Kabas, Fuat Arslan, Valiyeh A. Nezhad, Saban Ozturk, Emine U. Saritas, and Tolga C ¸ ukur. Physics-Driven Autoregressive State Space Models for Medical Image Reconstruc- tion.https://arxiv.org/pdf/2412.09331, 12 2024
-
[52]
Arjun D Desai, Batu M Ozturkler, Christopher M Sandino, Robert Boutin, Marc Willis, Shreyas Vasanawala, Brian A Hargreaves, Christopher R´ e, John M Pauly, and Akshay S Chaudhari. Noise2Recon: Enabling SNR-robust MRI reconstruction with semi-supervised and self-supervised learning.Magnetic Resonance in Medicine, 90(5):2052–2070, 2023
work page 2052
-
[53]
Frederic Wang, Han Qi, Alfredo De Goyeneche, Reinhard Heckel, Michael Lustig, and Efrat Shimron. K-band: self-supervised MRI reconstruction via stochastic gradient descent over k-space subsets.arXiv preprint arXiv:2308.02958, 2023
-
[54]
The NMR phased array.Magnetic resonance in medicine, 16(2):192–225, 1990
Peter B Roemer, William A Edelstein, Cecil E Hayes, Steven P Souza, and Otward M Mueller. The NMR phased array.Magnetic resonance in medicine, 16(2):192–225, 1990
work page 1990
-
[55]
Categorical Reparameterization with Gumbel-Softmax
Eric Jang, Shixiang Gu, and Ben Poole. Categorical Reparameterization with Gumbel- Softmax.5th International Conference on Learning Representations - Conference Track Proceedings, 11 2016. URLhttps://arxiv.org/pdf/1611.01144
work page internal anchor Pith review Pith/arXiv arXiv 2016
-
[56]
Martin Uecker, Peng Lai, Mark J Murphy, Patrick Virtue, Michael Elad, John M Pauly, Shreyas S Vasanawala, and Michael Lustig. ESPIRiT—an eigenvalue approach to autocal- ibrating parallel MRI: where SENSE meets GRAPPA.Magnetic Resonance in Medicine, 71(3):990–1001, 2014
work page 2014
-
[57]
Kerstin Hammernik, Jo Schlemper, Chen Qin, Jinming Duan, Ronald M Summers, and Daniel Rueckert. Systematic evaluation of iterative deep neural networks for fast parallel MRI reconstruction with sensitivity-weighted coil combination.Magnetic Resonance in Medicine, 86(4):1859–1872, 2021
work page 2021
-
[58]
Patricia M Johnson, Dana J Lin, Jure Zbontar, C Lawrence Zitnick, Anuroop Sriram, Matthew Muckley, James S Babb, Mitchell Kline, Gina Ciavarra, Erin Alaia, et al. Deep learning reconstruction enables prospectively accelerated clinical knee MRI.Radiology, 307 (2), 2023. 20
work page 2023
-
[59]
Ke Wang, Jonathan I Tamir, Alfredo De Goyeneche, Uri Wollner, Rafi Brada, Stella X Yu, and Michael Lustig. High fidelity deep learning-based MRI reconstruction with instance- wise discriminative feature matching loss.Magnetic Resonance in Medicine, 88(1):476–491, 2022
work page 2022
-
[60]
Aniket Pramanik, Sampada Bhave, Saurav Sajib, Samir D Sharma, and Mathews Jacob. Adapting model-based deep learning to multiple acquisition conditions: Ada-MoDL.Mag- netic resonance in medicine, 90(5):2033–2051, 2023
work page 2033
-
[61]
Mengye Lyu, Lifeng Mei, Shoujin Huang, Sixing Liu, Yi Li, Kexin Yang, Yilong Liu, Yu Dong, Linzheng Dong, and Ed X Wu. M4Raw: A multi-contrast, multi-repetition, multi-channel MRI k-space dataset for low-field MRI research.Scientific Data, 10(1):264, 2023
work page 2023
-
[62]
Sigpy: a python package for high performance iterative reconstruction
Frank Ong and Michael Lustig. Sigpy: a python package for high performance iterative reconstruction. InProceedings of the ISMRM 27th Annual Meeting, Montreal, Quebec, Canada, volume 4819, 2019
work page 2019
-
[63]
Adam Paszke, Sam Gross, Francisco Massa, Adam Lerer, James Bradbury, Gregory Chanan, Trevor Killeen, Zeming Lin, Natalia Gimelshein, Luca Antiga, et al. Pytorch: An imperative style, high-performance deep learning library.Advances in neural informa- tion processing systems, 32, 2019
work page 2019
-
[64]
Adam: A Method for Stochastic Optimization
Diederik P Kingma and Jimmy Ba. Adam: A method for stochastic optimization.arXiv preprint arXiv:1412.6980, 2014
work page internal anchor Pith review Pith/arXiv arXiv 2014
-
[65]
fastMRI: An open dataset and benchmarks for accelerated mri,
Jure Zbontar, Florian Knoll, Anuroop Sriram, Tullie Murrell, Zhengnan Huang, Matthew J Muckley, Aaron Defazio, Ruben Stern, Patricia Johnson, Mary Bruno, et al. fastMRI: An open dataset and benchmarks for accelerated MRI.arXiv preprint arXiv:1811.08839, 2018
-
[66]
Zhou Wang, Alan Conrad Bovik, Hamid Rahim Sheikh, and Eero P. Simoncelli. Image quality assessment: From error visibility to structural similarity.IEEE Transactions on Image Processing, 13, 2004. ISSN 10577149
work page 2004
-
[67]
Lin Zhang, Lei Zhang, Xuanqin Mou, and David Zhang. FSIM: A Feature Similarity Index for Image Quality Assessment.IEEE Transactions on Image Processing, 20(8):2378–2386, 2011
work page 2011
-
[68]
Allister Mason, James Rioux, Sharon E Clarke, Andreu Costa, Matthias Schmidt, Valerie Keough, Thien Huynh, and Steven Beyea. Comparison of objective image quality metrics to expert radiologists’ scoring of diagnostic quality of MR images.IEEE transactions on medical imaging, 39(4):1064–1072, 2019
work page 2019
-
[69]
Pauli Virtanen, Ralf Gommers, Travis E Oliphant, Matt Haberland, Tyler Reddy, David Cournapeau, Evgeni Burovski, Pearu Peterson, Warren Weckesser, Jonathan Bright, et al. SciPy 1.0: fundamental algorithms for scientific computing in Python.Nature Methods, 17 (3):261–272, 2020
work page 2020
-
[70]
Guanhua Wang, Tianrui Luo, Jon-Fredrik Nielsen, Douglas C Noll, and Jeffrey A Fessler. B- spline parameterized joint optimization of reconstruction and k-space trajectories (BJORK) for accelerated 2D MRI.IEEE Transactions on Medical Imaging, 41(9):2318–2330, 2022
work page 2022
-
[71]
Marcelo VW Zibetti, Gabor T Herman, and Ravinder R Regatte. Fast data-driven learning of parallel MRI sampling patterns for large scale problems.Scientific Reports, 11(1):19312, 2021. 21
work page 2021
-
[72]
Acceler- ating ultra-low field MRI with compressed sensing
D Waddington, Efrat Shimron, Nicholas Hindley, Neha Koonjoo, and M Rosen. Acceler- ating ultra-low field MRI with compressed sensing. InProceedings of the Annual Meeting of ISMRM, page 1818, 2022. A Probability maps analysis To evaluate the spatialk-spacesampling distribution learned by the different methods, we analyzed the learned probability mapsqfor t...
work page 2022
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.