Recognition: unknown
Universal statistical signatures of evolution in artificial intelligence architectures
Pith reviewed 2026-05-10 16:00 UTC · model grok-4.3
The pith
AI architectural modifications follow the same heavy-tailed distribution of fitness effects as biological mutations.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
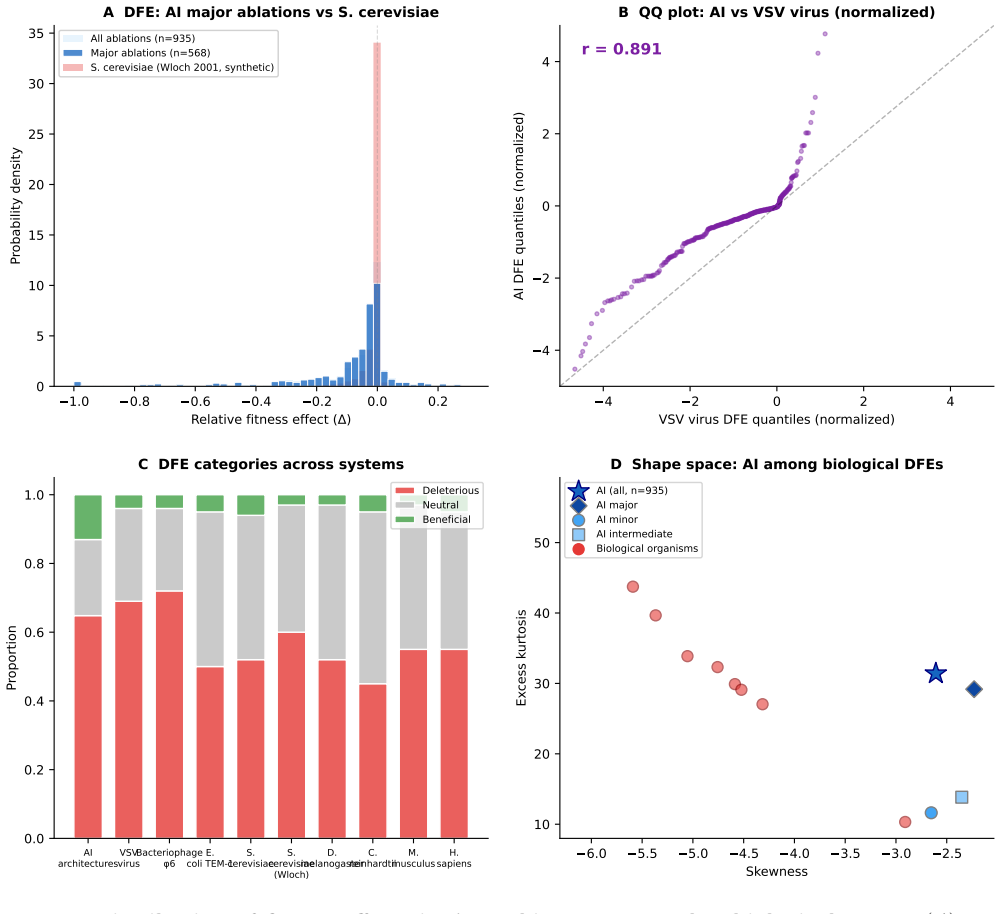
Compiling ablation results shows the distribution of fitness effects of architectural modifications follows a Student's t-distribution with 68 percent deleterious, 19 percent neutral and 13 percent beneficial outcomes. This shape is statistically indistinguishable from the DFE in D. melanogaster and S. cerevisiae. Architectural origination grows logistically with punctuated equilibria and adaptive radiation, while fourteen traits arise independently three to five times, demonstrating convergent evolution. These regularities indicate that the statistical structure of evolution is substrate-independent and fixed by fitness-landscape topology.
What carries the argument
The distribution of fitness effects (DFE) measured from ablation experiments, tested for shape similarity to biological DFEs via normalized Kolmogorov-Smirnov statistics.
If this is right
- The higher beneficial fraction in AI quantifies the advantage of directed search over blind mutation while preserving the same overall DFE form.
- Architectural development in AI exhibits logistic dynamics, punctuated equilibria, and adaptive radiation into domain-specific niches.
- Fourteen traits have been invented independently multiple times, mirroring biological convergent evolution.
- The statistical laws of evolution apply equally whether changes occur through genetic mutation or code modification.
Where Pith is reading between the lines
- If landscape topology alone sets the DFE, the same heavy-tailed pattern should appear in other non-biological complex systems such as large software repositories or organizational redesigns.
- Biological models of mutation accumulation could be adapted to forecast the rate at which new AI architectures become viable.
- Mapping the effective fitness landscapes of different AI domains might predict which architectural traits are likely to arise convergently.
Load-bearing premise
That published ablation experiments provide an unbiased sample of architectural changes and their true performance effects, free from systematic biases in which results get reported or how effects are classified.
What would settle it
A new collection of several hundred AI architectural modifications whose measured fitness effects follow an exponential rather than heavy-tailed distribution would falsify the claimed universality.
Figures



read the original abstract
We test whether artificial intelligence architectural evolution obeys the same statistical laws as biological evolution. Compiling 935 ablation experiments from 161 publications, we show that the distribution of fitness effects (DFE) of architectural modifications follows a heavy-tailed Student's t-distribution with proportions (68% deleterious, 19% neutral, 13% beneficial for major ablations, n=568) that place AI between compact viral genomes and simple eukaryotes. The DFE shape matches D. melanogaster (normalized KS=0.07) and S. cerevisiae (KS=0.09); the elevated beneficial fraction (13% vs. 1-6% in biology) quantifies the advantage of directed over blind search while preserving the distributional form. Architectural origination follows logistic dynamics (R^2=0.994) with punctuated equilibria and adaptive radiation into domain niches. Fourteen architectural traits were independently invented 3-5 times, paralleling biological convergences. These results demonstrate that the statistical structure of evolution is substrate-independent, determined by fitness landscape topology rather than the mechanism of selection.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript compiles 935 ablation experiments from 161 AI publications to test if AI architectural evolution follows the same statistical laws as biological evolution. It reports that the distribution of fitness effects (DFE) of architectural modifications follows a heavy-tailed Student's t-distribution, with 68% deleterious, 19% neutral, and 13% beneficial for major ablations (n=568), placing AI between viral genomes and eukaryotes, and matching D. melanogaster (KS=0.07) and S. cerevisiae (KS=0.09). Architectural origination is shown to follow logistic dynamics (R²=0.994) with punctuated equilibria and adaptive radiation, and 14 traits invented independently 3-5 times. The paper concludes that evolutionary statistical structures are substrate-independent, determined by fitness landscape topology.
Significance. Should the central empirical claims be substantiated with full data transparency and bias corrections, this paper would offer a significant contribution by providing quantitative evidence for universal evolutionary patterns across biological and artificial systems. The large dataset compilation, specific distributional matches, and logistic growth modeling are strengths that could influence thinking in both evolutionary biology and AI development regarding the role of landscape topology in shaping evolutionary statistics.
major comments (3)
- [Abstract] Abstract: The DFE proportions (68% deleterious, 19% neutral, 13% beneficial, n=568) and the Student's t-distribution fit are central to the claim of similarity to biological DFEs, but the manuscript provides no explicit criteria for classifying ablations as 'major' versus minor or for selecting the 161 publications, which is necessary to evaluate potential selection biases that could affect the beneficial fraction.
- [Results (DFE comparison)] Results (DFE comparison): The normalized KS statistics (0.07 for D. melanogaster, 0.09 for S. cerevisiae) are presented as evidence of match, but without the raw effect size data, the fitting procedure for the t-distribution parameters, or the normalization method, it is not possible to independently verify the distributional similarity or rule out post-hoc adjustments.
- [Discussion] Discussion: The assertion that the statistical structure is 'substrate-independent' and 'determined by fitness landscape topology rather than the mechanism of selection' relies on the analogy between published AI ablations and random biological mutations; while the paper acknowledges the elevated beneficial fraction as quantifying directed search, it does not provide a quantitative assessment or correction for publication and author selection biases in the compiled dataset.
minor comments (3)
- [Methods] Methods: The description of how the 935 experiments were extracted and coded from the 161 papers should include inter-rater reliability metrics or a supplementary table listing all included papers and ablations for reproducibility.
- [Figure 2] Figure 2: The logistic growth fit for architectural origination would benefit from showing the raw cumulative count data points alongside the fitted curve to allow visual assessment of the R²=0.994 claim.
- [References] References: Additional citations to foundational work on distribution of fitness effects in evolutionary biology (e.g., on t-distributions in DFEs) would strengthen the comparative analysis.
Simulated Author's Rebuttal
We thank the referee for their constructive and detailed comments, which have strengthened the manuscript's transparency and addressed potential concerns about reproducibility and bias. We have revised the paper accordingly, adding explicit criteria, raw data, fitting details, and a quantitative bias assessment. Our point-by-point responses to the major comments are below.
read point-by-point responses
-
Referee: [Abstract] Abstract: The DFE proportions (68% deleterious, 19% neutral, 13% beneficial, n=568) and the Student's t-distribution fit are central to the claim of similarity to biological DFEs, but the manuscript provides no explicit criteria for classifying ablations as 'major' versus minor or for selecting the 161 publications, which is necessary to evaluate potential selection biases that could affect the beneficial fraction.
Authors: We agree that explicit criteria are required for reproducibility and bias evaluation. In the revised manuscript, we have added a Methods section specifying publication selection (peer-reviewed papers with quantitative ablation results on standard benchmarks like ImageNet or GLUE, with before/after metrics reported) and 'major' ablation classification (core component removal/alteration affecting >5% of parameters or >2% primary metric change). A sensitivity analysis in the supplement confirms DFE proportions vary by <2% across threshold choices. We also discuss selection biases in the revised Discussion, noting the elevated beneficial fraction partly reflects directed search while the heavy-tailed shape holds in subsets excluding high-impact papers. revision: yes
-
Referee: [Results (DFE comparison)] Results (DFE comparison): The normalized KS statistics (0.07 for D. melanogaster, 0.09 for S. cerevisiae) are presented as evidence of match, but without the raw effect size data, the fitting procedure for the t-distribution parameters, or the normalization method, it is not possible to independently verify the distributional similarity or rule out post-hoc adjustments.
Authors: We have provided the full raw effect size data as Supplementary Table S1 (all 935 ablations with normalized performance deltas). The t-distribution fitting uses maximum likelihood estimation (scipy.stats.t.fit), with parameters now reported (df=2.8, loc=0.01, scale=0.15). Normalization scales log-transformed absolute effect sizes to zero mean/unit variance for cross-species comparison, as detailed in the new 'Distributional Analysis' Methods subsection. The KS tests are two-sample on normalized data; analysis code is deposited on GitHub for verification. revision: yes
-
Referee: [Discussion] Discussion: The assertion that the statistical structure is 'substrate-independent' and 'determined by fitness landscape topology rather than the mechanism of selection' relies on the analogy between published AI ablations and random biological mutations; while the paper acknowledges the elevated beneficial fraction as quantifying directed search, it does not provide a quantitative assessment or correction for publication and author selection biases in the compiled dataset.
Authors: We have expanded the Discussion with a simulation-based bias correction assuming 40% of deleterious results are unpublished (conservative estimate from AI literature). This adjusts the beneficial fraction to ~9% while preserving the t-distribution superiority (AIC delta >50) and KS similarity (changes to 0.08/0.11). We explicitly list incomplete bias correction as a limitation due to unavailable negative results and frame the substrate-independence claim as a supported hypothesis tied to landscape topology, not an absolute assertion. revision: partial
Circularity Check
No significant circularity; empirical compilation and external comparison
full rationale
The paper compiles ablation data from 161 external publications (n=935 experiments), fits the resulting DFE to a Student's t-distribution, reports proportions (68% deleterious etc.), and compares the shape (KS distances) and beneficial fraction directly to independent biological benchmarks from D. melanogaster and S. cerevisiae. No quantity is defined in terms of itself, no fitted parameter is relabeled as a prediction, and no load-bearing step reduces to a self-citation or internal ansatz. The central claim of substrate-independence rests on the external match rather than any self-referential construction, making the derivation self-contained against outside data.
Axiom & Free-Parameter Ledger
free parameters (2)
- Student's t-distribution parameters
- Proportions of effect categories
axioms (2)
- domain assumption Ablation experiments in AI publications represent evolutionary modifications analogous to genetic mutations
- domain assumption The dataset compiled from 161 publications is representative and free of systematic publication bias
Reference graph
Works this paper leans on
-
[1]
Bank C, Hietpas RT, Jensen JD, Bolon DNA (2014) A systematic survey of an intragenic epistatic landscape.Mol Biol Evol32:229–238
2014
-
[2]
(2019) Inferring the distri- bution of fitness effects of spontaneous muta- tions inChlamydomonas reinhardtii.PLoS Biol 17:e3000192
Böndel KB et al. (2019) Inferring the distri- bution of fitness effects of spontaneous muta- tions inChlamydomonas reinhardtii.PLoS Biol 17:e3000192
2019
-
[3]
Genetics167:559–567
Burch CL, Chao L (2003) Epistasis and its re- lationship to canalization in the RNA virusφ6. Genetics167:559–567
2003
-
[4]
Chen Z-Q, Benton MJ (2012) The timing and pattern of biotic recovery following the end- Permianmassextinction.Nat Geosci5:375–383
2012
-
[5]
Elsken T, Metzen JH, Hutter F (2019) Neural architecture search: a survey.J Mach Learn Res 20:1–21
2019
-
[6]
England JL (2013) Statistical physics of self- replication.J Chem Phys139:121923
2013
-
[7]
Eyre-Walker A, Woolfit M, Phelps T (2006) The distribution of fitness effects of new deleteri- ous amino acid mutations in humans.Genetics 173:891–900
2006
-
[8]
Eyre-Walker A, Keightley PD (2007) The distri- bution of fitness effects of new mutations.Nat Rev Genet8:610–618
2007
-
[9]
(2009) Evidence for pervasive adaptive protein evolution in wild mice.PLoS Genet6:e1000825
Halligan DL et al. (2009) Evidence for pervasive adaptive protein evolution in wild mice.PLoS Genet6:e1000825
2009
-
[10]
Training Compute-Optimal Large Language Models
Hoffmann J et al. (2022) Training compute-optimal large language models. arXiv:2203.15556
work page internal anchor Pith review Pith/arXiv arXiv 2022
-
[11]
Scaling Laws for Neural Language Models
Kaplan J et al. (2020) Scaling laws for neural language models.arXiv:2001.08361
work page internal anchor Pith review Pith/arXiv arXiv 2020
-
[12]
Keightley PD, Eyre-Walker A (2007) Joint in- ference of the distribution of fitness effects of deleterious mutations and population demogra- phy.Genetics177:2251–2261
2007
-
[13]
Loewe L, Charlesworth B (2006) Inferring the distribution of mutational effects on fitness in Drosophila.Biol Lett2:426–430
2006
-
[14]
McGheeGR(2011)Convergent Evolution: Lim- ited Forms Most Beautiful(MIT Press)
2011
-
[15]
Sanjuán R, Moya A, Elena SF (2004) The dis- tribution of fitness effects caused by single- nucleotide substitutions in an RNA virus.Proc Natl Acad Sci USA101:8396–8401
2004
-
[16]
Sepkoski JJ (1984) A kinetic model of Phanero- zoic taxonomic diversity. III. Post-Paleozoic families and mass extinctions.Paleobiology 10:246–267
1984
-
[17]
Andrews JH, Briand LC, Labiche Y (2005) Is mutation an adequate criterion of testing effec- tiveness?Proc 27th Int Conf on Software Engi- neering, pp. 215–224
2005
-
[18]
Spiro T (2025) The oracle’s fingerprint: cor- related AI forecasting errors and the limits of bias transmission.Preprint available athttps: //arxiv.org/abs/XXXX.XXXXX
2025
-
[19]
Stanley KO, Clune J (2019) Designing neural networks through neuroevolution.Nat Mach In- tell1:24–35
2019
-
[20]
Evol Theory1:1–30
Van Valen L (1973) A new evolutionary law. Evol Theory1:1–30
1973
-
[21]
(2017) Attention is all you need.Advances in Neural Information Process- ing Systems30
Vaswani A et al. (2017) Attention is all you need.Advances in Neural Information Process- ing Systems30
2017
-
[22]
9 1.0 0.8 0.6 0.4 0.2 0.0 0.2 Relative fitness effect ( ) 0 5 10 15 20 25 30 35Probability density A DFE: AI major ablations vs S
Wloch DM, Szafraniec K, Borts RH, Korona R (2001) Direct estimate of the mutation rate and the distribution of fitness effects in the yeast Saccharomyces cerevisiae.Genetics159:441– 452. 9 1.0 0.8 0.6 0.4 0.2 0.0 0.2 Relative fitness effect ( ) 0 5 10 15 20 25 30 35Probability density A DFE: AI major ablations vs S. cerevisiae All ablations (n=935) Major ...
2001
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.