Recognition: unknown
Dual Triangle Attention: Effective Bidirectional Attention Without Positional Embeddings
Pith reviewed 2026-05-10 16:46 UTC · model grok-4.3
The pith
Dual Triangle Attention splits each head into past and future triangular masks to give bidirectional models implicit positional information without embeddings.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
Dual Triangle Attention separates the query-key subspace of each attention head into two complementary triangular masks—one that attends to past-and-self positions and one that attends to future-and-self positions. This design supplies full bidirectional context while preserving the causal mask's implicit positional inductive bias in both directions. The mechanism is realized as a single compiled kernel call with no added parameters. On an argmax position probe, it learns positional information like causal attention does, unlike standard bidirectional attention. In masked language modeling on natural language and on protein sequences it performs competitively, with the strongest context-exti
What carries the argument
Dual Triangle Attention, which partitions each attention head's query-key subspace into two complementary triangular masks to supply bidirectional context plus positional bias.
If this is right
- Models using Dual Triangle Attention learn positional information in synthetic argmax tasks without explicit embeddings, matching causal attention.
- The mechanism supports masked language modeling on natural language with performance comparable to or better than baselines.
- On protein sequences the same attention produces strong masked language modeling results.
- Pairing Dual Triangle Attention with rotary embeddings yields the best observed context-extension behavior.
- Implementation requires no additional learned parameters beyond ordinary multi-head attention.
Where Pith is reading between the lines
- The same mask-splitting idea could be tested in other biological sequence domains such as DNA or small-molecule strings.
- Removing positional embedding layers might reduce memory use and simplify scaling for very long protein or genomic sequences.
- Performance in open-ended generation rather than masked reconstruction remains an open question for this attention variant.
Load-bearing premise
That dividing the attention subspace into two fixed triangular masks will reliably inject enough positional signal for effective learning in bidirectional masked language modeling tasks.
What would settle it
If Dual Triangle Attention models fail to learn accurate position predictions on the argmax probe while causal-attention models succeed, or if they underperform standard bidirectional attention plus embeddings on masked language modeling, the claim would be falsified.
Figures




read the original abstract
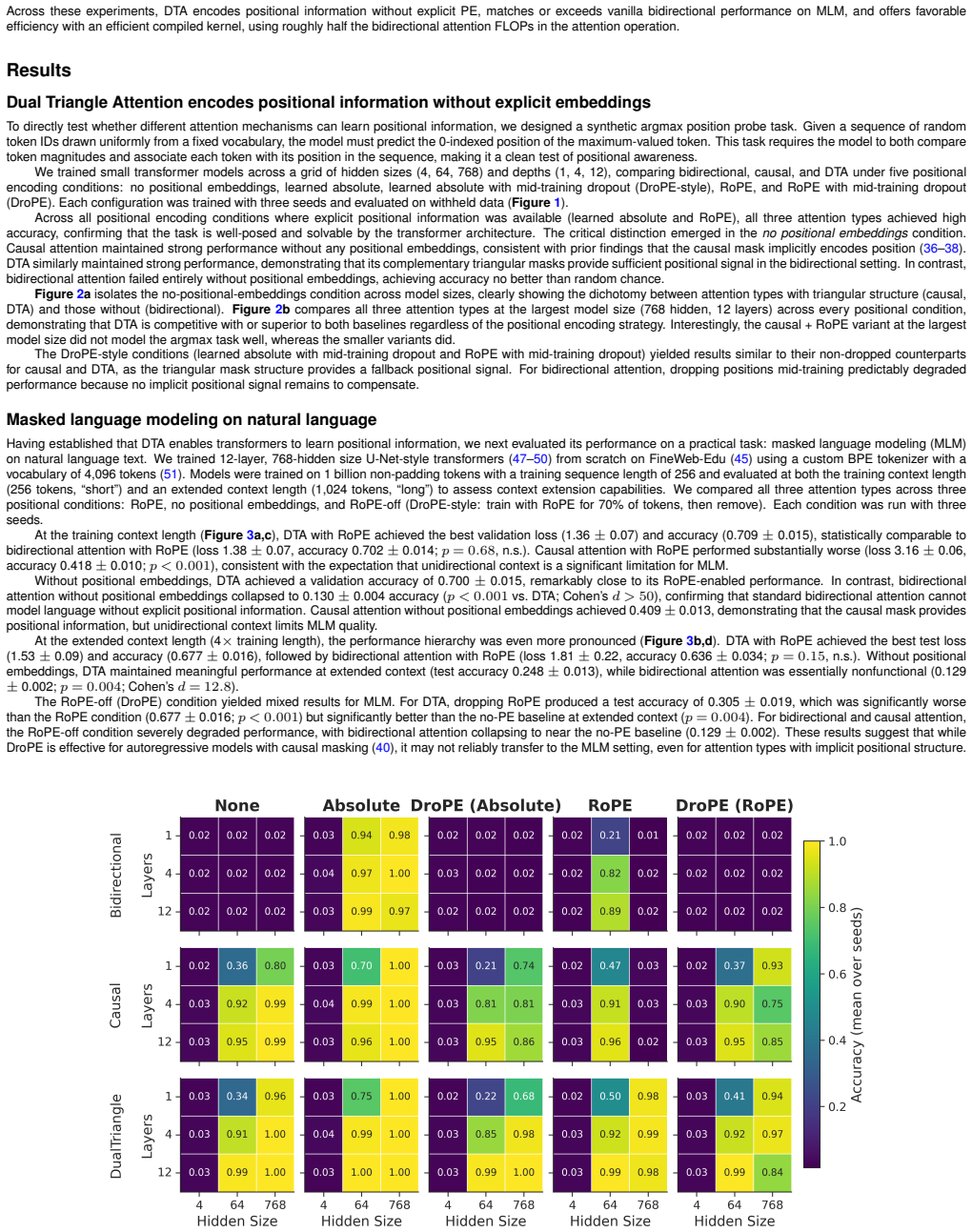
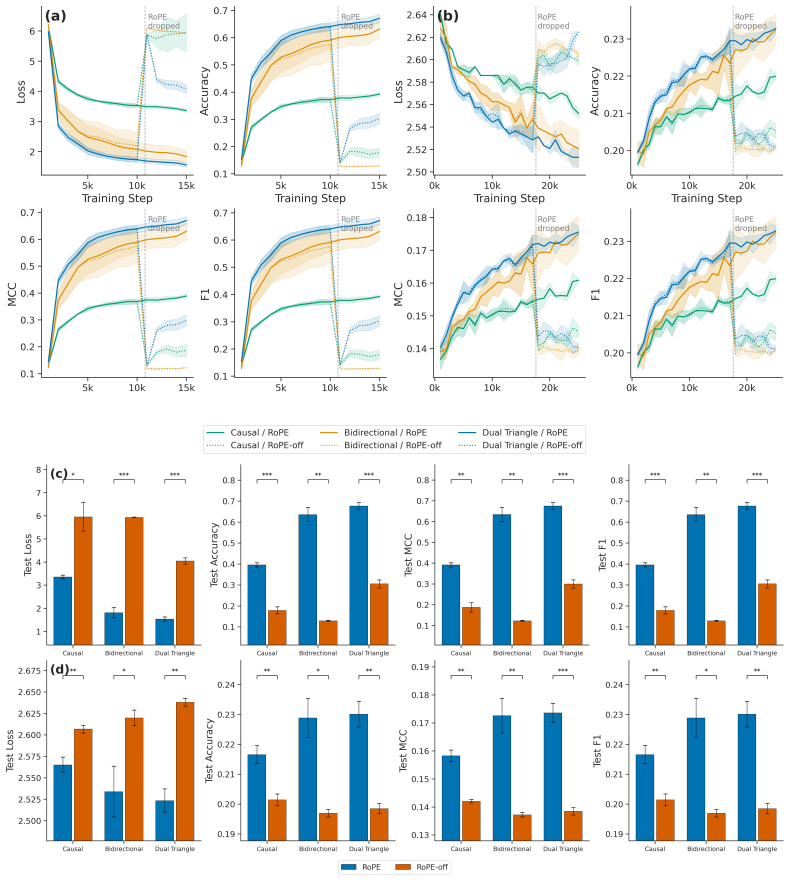
Bidirectional transformers are the foundation of many sequence modeling tasks across natural, biological, and chemical language domains, but they are permutation-invariant without explicit positional embeddings. In contrast, unidirectional attention inherently encodes positional information through its triangular mask, enabling models to operate without positional embeddings altogether. Here, we introduce Dual Triangle Attention, a novel bidirectional attention mechanism that separates the query-key subspace of each attention head into two complementary triangular masks: one that attends to past-and-self positions and one that attends to future-and-self positions. This design provides bidirectional context while maintaining the causal mask's implicit positional inductive bias in both directions. Using PyTorch's flex_attention, Dual Triangle Attention is implemented as a single compiled kernel call with no additional parameters beyond standard multi-head attention. We evaluated Dual Triangle Attention across three settings: (1) a synthetic argmax position probe, (2) masked language modeling (MLM) on natural language, and (3) MLM on protein sequences. In the argmax task, both Dual Triangle Attention and causal attention learn positional information without explicit positional embeddings, whereas standard bidirectional attention cannot. In the MLM experiments, Dual Triangle Attention with Rotary Positional Embeddings (RoPE) achieved the best context extension performance and strong performance across the board. These findings suggest that Dual Triangle Attention is a viable attention mechanism for bidirectional transformers, with or without positional embeddings.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper proposes Dual Triangle Attention, a bidirectional attention variant that partitions each head's query-key subspace into two complementary triangular masks (past-and-self and future-and-self) to supply bidirectional context while retaining the implicit positional inductive bias of causal attention. This allows bidirectional transformers to operate without explicit positional embeddings. The mechanism is implemented as a single flex_attention kernel call with no added parameters. Evaluation consists of a synthetic argmax position probe (where Dual Triangle and causal attention succeed but standard bidirectional fails) plus masked language modeling on natural-language and protein sequences (where the Dual Triangle + RoPE variant performs best on context extension).
Significance. If the central claim holds, the work offers a parameter-free architectural route to positional awareness in bidirectional models, which could simplify training in domains such as protein language modeling where explicit positional encodings are often costly or brittle. The efficient single-kernel implementation and the synthetic probe result are concrete strengths that would be valuable if replicated with full quantitative reporting.
major comments (3)
- [Abstract / MLM experiments] Abstract and MLM-experiments paragraph: the claim that Dual Triangle Attention works 'with or without positional embeddings' on practical MLM tasks is unsupported because only the Dual Triangle + RoPE variant is reported as achieving best context-extension performance; no metrics, ablations, or even qualitative statements are given for the no-PE Dual Triangle model on the natural-language or protein MLM benchmarks. This is load-bearing for the central 'viable with or without' assertion.
- [Abstract] Abstract: the positive results on the argmax probe and MLM tasks are stated without any quantitative metrics, standard deviations, ablation tables, or implementation hyperparameters, leaving the magnitude and reliability of the claimed improvements impossible to assess from the provided text.
- [Method / Implementation] Implementation description: although the paper states that Dual Triangle Attention is realized as a single compiled flex_attention kernel, no pseudocode, mask-construction equations, or verification that the two triangular masks are applied to disjoint subspaces of the same QK projection are supplied, making reproducibility and correctness verification difficult.
minor comments (1)
- [Abstract] The abstract refers to 'three settings' but only describes the argmax probe and two MLM tasks; a brief enumeration of the exact datasets or sequence lengths used would improve clarity.
Simulated Author's Rebuttal
We thank the referee for the constructive review and valuable feedback. We have revised the manuscript to address the concerns on clarity, quantitative reporting, and implementation details. Point-by-point responses follow.
read point-by-point responses
-
Referee: The claim that Dual Triangle Attention works 'with or without positional embeddings' on practical MLM tasks is unsupported because only the Dual Triangle + RoPE variant is reported as achieving best context-extension performance; no metrics, ablations, or even qualitative statements are given for the no-PE Dual Triangle model on the natural-language or protein MLM benchmarks.
Authors: We agree the abstract phrasing could be clarified. The MLM results emphasize the RoPE variant for best performance, while no-PE viability is shown via the synthetic probe and mechanism design. We have revised the abstract to qualify the statement accurately and added qualitative discussion plus metrics for the no-PE variant on MLM benchmarks in the experiments section. revision: yes
-
Referee: The positive results on the argmax probe and MLM tasks are stated without any quantitative metrics, standard deviations, ablation tables, or implementation hyperparameters.
Authors: We concur that the abstract benefits from more specifics. We have updated it to include key metrics (e.g., probe accuracy, MLM perplexity), references to full tables with standard deviations, and hyperparameters from the main text and appendix. revision: yes
-
Referee: No pseudocode, mask-construction equations, or verification that the two triangular masks are applied to disjoint subspaces of the same QK projection are supplied.
Authors: We have expanded the Methods section with pseudocode for the single flex_attention kernel, explicit equations for the complementary triangular masks, and verification that they partition the QK subspace disjointly per head. This improves reproducibility. revision: yes
Circularity Check
No circularity: direct architectural definition with empirical support
full rationale
The paper defines Dual Triangle Attention directly as a split of the query-key subspace into complementary triangular masks, implemented via a single flex_attention kernel with no added parameters. No equations, derivations, or performance claims reduce by construction to fitted parameters, self-referential definitions, or a chain of self-citations. The argmax probe and MLM evaluations are presented as independent empirical tests rather than tautological restatements of the mechanism. The central claim of bidirectional context with preserved positional bias is therefore self-contained and does not rely on any of the enumerated circular patterns.
Axiom & Free-Parameter Ledger
axioms (2)
- standard math Standard multi-head attention formulation with masking
- domain assumption Triangular causal masks implicitly encode positional information
invented entities (1)
-
Dual Triangle Attention
no independent evidence
Reference graph
Works this paper leans on
-
[1]
Ashish Vaswani, Noam Shazeer, Niki Parmar, Jakob Uszkoreit, Llion Jones, Aidan N. Gomez, Lukasz Kaiser, and Illia Polosukhin. Attention is all you need. arXiv, 2017. doi:10.48550/arXiv.1706.03762. URLhttp://arxiv.org/abs/1706.03762. Number: arXiv:1706.03762
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.1706.03762 2017
-
[2]
Language Models are Few-Shot Learners
Tom B. Brown, Benjamin Mann, Nick Ryder, Melanie Subbiah, Jared Kaplan, Prafulla Dhariwal, Arvind Neelakantan, Pranav Shyam, et al. Language models are few-shot learners. arXiv, 2020. doi:10.48550/arXiv.2005.14165. URLhttp://arxiv.org/abs/2005.14165
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.2005.14165 2020
-
[3]
Zeming Lin, Halil Akin, Roshan Rao, Brian Hie, Zhongkai Zhu, Wenting Lu, Nikita Smetanin, Robert Verkuil, et al. Evolutionary-scale prediction of atomic-level protein structure with a language model.Science, 379(6637):1123–1130, 2023. doi:10.1126/science.ade2574
-
[4]
Highly accurate protein structure prediction with AlphaFold
John Jumper, Richard Evans, Alexander Pritzel, Tim Green, Michael Figurnov, Olaf Ronneberger, Kathryn Tunyasuvunakool, Russ Bates, et al. Highly accurate protein structure prediction with AlphaFold.Nature, 596(7873):583–589, 2021. ISSN 1476-4687. doi:10.1038/s41586-021-03819-2. Number: 7873
-
[5]
Patrick Wilhelm, Thorsten Wittkopp, and Odej Kao
Benjamin Warner, Antoine Chaffin, Benjamin Clavié, Orion Weller, Oskar Hallström, Said Taghadouini, Alexis Gallagher, Raja Biswas, et al. Smarter, better, faster, longer: A modern bidirectional encoder for fast, memory efficient, and long context finetuning and inference. arXiv, 2024. doi:10.48550/arXiv.2412.13663. URL http://arxiv.org/abs/2412.13663
-
[6]
Logan Hallee, Rohan Kapur, Arjun Patel, Jason P . Gleghorn, and Bohdan B. Khomtchouk. Contrastive learning and mixture of experts enables precise vector embeddings in biological databases.Sci Rep, 15(1):14953, 2025. ISSN 2045-2322. doi:10.1038/s41598-025-98185-8
-
[7]
Bichara, and Jason P
Logan Hallee, Nikolaos Rafailidis, David B. Bichara, and Jason P . Gleghorn. Diffusion sequence models for enhanced protein representation and generation. arXiv,
-
[8]
URLhttp://arxiv.org/abs/2506.08293
doi:10.48550/arXiv.2506.08293. URLhttp://arxiv.org/abs/2506.08293
-
[9]
Josh Abramson, Jonas Adler, Jack Dunger, Richard Evans, Tim Green, Alexander Pritzel, Olaf Ronneberger, Lindsay Willmore, et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3.Nature, 630(8016):493–500, 2024. ISSN 1476-4687. doi:10.1038/s41586-024-07487-w
-
[10]
Logan Hallee and Jason P . Gleghorn. Protein-protein interaction prediction is achievable with large language models. bioRxiv, 2023. doi:10.1101/2023.06.07.544109. URLhttps://www.biorxiv.org/content/10.1101/2023.06.07.544109v1. Pages: 2023.06.07.544109 Section: New Results
-
[11]
Logan Hallee, Tamar Peleg, Nikolaos Rafailidis, and Jason P . Gleghorn. Protein language models are accidental taxonomists. bioRxiv, 2025. doi:10.1101/2025. 10.07.681002. URLhttps://www.biorxiv.org/content/10.1101/2025.10.07.681002v1. ISSN: 2692-8205 Pages: 2025.10.07.681002 Section: New Results
-
[12]
Y oung Su Ko, Jonathan Parkinson, Cong Liu, and Wei Wang. TUnA: an uncertainty-aware transformer model for sequence-based protein–protein interaction prediction. Brief Bioinform, 25(5):bbae359, 2024. ISSN 1477-4054. doi:10.1093/bib/bbae359
-
[13]
Logan Hallee, Niko Rafailidis, Colin Horger, David Hong, and Jason P . Gleghorn. Annotation vocabulary (might be) all you need. bioRxiv, 2024. doi:10.1101/2024.07. 30.605924. URLhttps://www.biorxiv.org/content/10.1101/2024.07.30.605924v1. Pages: 2024.07.30.605924 Section: New Results. Halleeet al.| arXiv | April 22, 2026 | 8–12
-
[14]
doi:10.1101/2023.10.01.560349 , abstract =
Jin Su, Chenchen Han, Yuyang Zhou, Junjie Shan, Xibin Zhou, and Fajie Yuan. SaProt: Protein language modeling with structure-aware vocabulary. bioRxiv, 2023. doi:10.1101/2023.10.01.560349. URLhttps://www.biorxiv.org/content/10.1101/2023.10.01.560349v1. Pages: 2023.10.01.560349 Section: New Results
-
[15]
Logan Hallee, Nikolaos Rafailidis, and Jason P . Gleghorn. cdsBERT - extending protein language models with codon awareness. bioRxiv, 2023. doi:10.1101/2023.09. 15.558027. URLhttps://www.biorxiv.org/content/10.1101/2023.09.15.558027v1. Pages: 2023.09.15.558027 Section: New Results
-
[16]
Carlos Outeiral and Charlotte M. Deane. Codon language embeddings provide strong signals for use in protein engineering.Nat Mach Intell, 6(2):170–179, 2024. ISSN 2522-5839. doi:10.1038/s42256-024-00791-0
-
[17]
Diffusion language models are versatile protein learners.arXiv preprint arXiv:2402.18567, 2024
Xinyou Wang, Zaixiang Zheng, Fei Y e, Dongyu Xue, Shujian Huang, and Quanquan Gu. Diffusion language models are versatile protein learners. arXiv, 2024. doi: 10.48550/arXiv.2402.18567. URLhttp://arxiv.org/abs/2402.18567
-
[18]
Durrant, Brian Kang, Dhruva Katrekar, David B
Eric Nguyen, Michael Poli, Matthew G. Durrant, Brian Kang, Dhruva Katrekar, David B. Li, Liam J. Bartie, Armin W. Thomas, et al. Sequence modeling and design from molecular to genome scale with evo.Science, 386(6723):eado9336, 2024. doi:10.1126/science.ado9336
-
[19]
G., Ku, J., Poli, M., Brockman, G., Chang, D., Gonzalez, G
Garyk Brixi, Matthew G. Durrant, Jerome Ku, Michael Poli, Greg Brockman, Daniel Chang, Gabriel A. Gonzalez, Samuel H. King, et al. Genome modeling and design across all domains of life with evo 2. bioRxiv, 2025. doi:10.1101/2025.02.18.638918. URLhttps://www.biorxiv.org/content/10.1101/2025.02.18. 638918v1. Pages: 2025.02.18.638918 Section: New Results
-
[20]
Taylor, Tom Ward, Clare Bycroft, Lauren Nicolaisen, et al
Žiga Avsec, Natasha Latysheva, Jun Cheng, Guido Novati, Kyle R. Taylor, Tom Ward, Clare Bycroft, Lauren Nicolaisen, et al. Advancing regulatory variant effect prediction with AlphaGenome.Nature, 649(8099):1206–1218, 2026. ISSN 1476-4687. doi:10.1038/s41586-025-10014-0
-
[21]
Position information in transformers: An overview
Philipp Dufter, Martin Schmitt, and Hinrich Schütze. Position information in transformers: An overview. arXiv, 2021. doi:10.48550/arXiv.2102.11090. URLhttp: //arxiv.org/abs/2102.11090
-
[22]
BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding
Jacob Devlin, Ming-Wei Chang, Kenton Lee, and Kristina Toutanova. BERT: Pre-training of deep bidirectional transformers for language understanding. arXiv, 2019. doi:10.48550/arXiv.1810.04805. URLhttp://arxiv.org/abs/1810.04805
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.1810.04805 2019
-
[23]
Self-attention with relative position repre- sentations
Peter Shaw, Jakob Uszkoreit, and Ashish Vaswani. Self-attention with relative position representations. arXiv, 2018. doi:10.48550/arXiv.1803.02155. URLhttp: //arxiv.org/abs/1803.02155
-
[24]
Exploring the Limits of Transfer Learning with a Unified Text-to-Text Transformer
Colin Raffel, Noam Shazeer, Adam Roberts, Katherine Lee, Sharan Narang, Michael Matena, Y anqi Zhou, Wei Li, et al. Exploring the limits of transfer learning with a unified text-to-text transformer. arXiv, 2023. doi:10.48550/arXiv.1910.10683. URLhttp://arxiv.org/abs/1910.10683
work page internal anchor Pith review doi:10.48550/arxiv.1910.10683 2023
-
[25]
URLhttps://doi.org/10.48550/arXiv
Pengcheng He, Xiaodong Liu, Jianfeng Gao, and Weizhu Chen. DeBERTa: Decoding-enhanced BERT with disentangled attention. arXiv, 2021. doi:10.48550/arXiv. 2006.03654. URLhttp://arxiv.org/abs/2006.03654
work page internal anchor Pith review doi:10.48550/arxiv 2021
-
[26]
Pengcheng He, Jianfeng Gao, and Weizhu Chen
Pengcheng He, Jianfeng Gao, and Weizhu Chen. DeBERTaV3: Improving DeBERTa using ELECTRA-style pre-training with gradient-disentangled embedding sharing. arXiv, 2023. doi:10.48550/arXiv.2111.09543. URLhttp://arxiv.org/abs/2111.09543
-
[27]
RoFormer: Enhanced Transformer with Rotary Position Embedding
Jianlin Su, Yu Lu, Shengfeng Pan, Ahmed Murtadha, Bo Wen, and Yunfeng Liu. RoFormer: Enhanced transformer with rotary position embedding. arXiv, 2023. URL http://arxiv.org/abs/2104.09864
work page internal anchor Pith review arXiv 2023
-
[28]
Llama 2: Open Foundation and Fine-Tuned Chat Models
Hugo Touvron, Louis Martin, Kevin Stone, Peter Albert, Amjad Almahairi, Y asmine Babaei, Nikolay Bashlykov, Soumya Batra, et al. Llama 2: Open foundation and fine-tuned chat models. arXiv, 2023. doi:10.48550/arXiv.2307.09288. URLhttp://arxiv.org/abs/2307.09288
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.2307.09288 2023
-
[29]
Aaron Grattafiori, Abhimanyu Dubey, Abhinav Jauhri, Abhinav Pandey, Abhishek Kadian, Ahmad Al-Dahle, Aiesha Letman, Akhil Mathur, et al. The llama 3 herd of models. arXiv, 2024. doi:10.48550/arXiv.2407.21783. URLhttp://arxiv.org/abs/2407.21783
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.2407.21783 2024
-
[30]
Albert Q. Jiang, Alexandre Sablayrolles, Arthur Mensch, Chris Bamford, Devendra Singh Chaplot, Diego de las Casas, Florian Bressand, Gianna Lengyel, et al. Mistral 7b. arXiv, 2023. doi:10.48550/arXiv.2310.06825. URLhttp://arxiv.org/abs/2310.06825
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.2310.06825 2023
-
[31]
A length-extrapolatable transformer
Yutao Sun, Li Dong, Barun Patra, Shuming Ma, Shaohan Huang, Alon Benhaim, Vishrav Chaudhary, Xia Song, et al. A length-extrapolatable transformer. arXiv, 2022. doi:10.48550/arXiv.2212.10554. URLhttp://arxiv.org/abs/2212.10554
-
[32]
Contextual position encoding: Learning to count what’s important
Olga Golovneva, Tianlu Wang, Jason Weston, and Sainbayar Sukhbaatar. Contextual position encoding: Learning to count what’s important. arXiv, 2024. doi: 10.48550/arXiv.2405.18719. URLhttp://arxiv.org/abs/2405.18719
-
[33]
Extending Context Window of Large Language Models via Positional Interpolation
Shouyuan Chen, Sherman Wong, Liangjian Chen, and Yuandong Tian. Extending context window of large language models via positional interpolation. arXiv, 2023. doi:10.48550/arXiv.2306.15595. URLhttp://arxiv.org/abs/2306.15595
work page internal anchor Pith review doi:10.48550/arxiv.2306.15595 2023
-
[34]
YaRN: Efficient Context Window Extension of Large Language Models
Bowen Peng, Jeffrey Quesnelle, Honglu Fan, and Enrico Shippole. Y aRN: Efficient context window extension of large language models. arXiv, 2023. doi:10.48550/ arXiv.2309.00071. URLhttp://arxiv.org/abs/2309.00071
work page internal anchor Pith review arXiv 2023
-
[35]
Longrope: Extending llm context window beyond 2 million tokens
Yiran Ding, Li Lyna Zhang, Chengruidong Zhang, Yuanyuan Xu, Ning Shang, Jiahang Xu, Fan Y ang, and Mao Y ang. LongRoPE: Extending LLM context window beyond 2 million tokens. arXiv, 2024. doi:10.48550/arXiv.2402.13753. URLhttp://arxiv.org/abs/2402.13753
-
[36]
Round and round we go! what makes rotary positional encodings useful?
Federico Barbero, Alex Vitvitskyi, Christos Perivolaropoulos, Razvan Pascanu, and Petar Veli ˇckovi´c. Round and round we go! what makes rotary positional encodings useful? arXiv, 2024. URLhttp://arxiv.org/abs/2410.06205
-
[37]
Transformer language models without positional encodings still learn positional information
Adi Haviv, Ori Ram, Ofir Press, Peter Izsak, and Omer Levy. Transformer language models without positional encodings still learn positional information. arXiv, 2022. doi:10.48550/arXiv.2203.16634. URLhttp://arxiv.org/abs/2203.16634
-
[38]
Length generalization of causal transformers without position encoding
Jie Wang, Tao Ji, Yuanbin Wu, Hang Y an, Tao Gui, Qi Zhang, Xuanjing Huang, and Xiaoling Wang. Length generalization of causal transformers without position encoding. arXiv, 2024. doi:10.48550/arXiv.2404.12224. URLhttp://arxiv.org/abs/2404.12224
-
[39]
The impact of positional encoding on length generalization in transformers
Amirhossein Kazemnejad, Inkit Padhi, Karthikeyan Natesan Ramamurthy, Payel Das, and Siva Reddy. The impact of positional encoding on length generalization in transformers. arXiv, 2023. doi:10.48550/arXiv.2305.19466. URLhttp://arxiv.org/abs/2305.19466
-
[40]
Rope to nope and back again: A new hybrid attention strategy.arXiv preprint arXiv:2501.18795, 2025
Bowen Y ang, Bharat Venkitesh, Dwarak Talupuru, Hangyu Lin, David Cairuz, Phil Blunsom, and Acyr Locatelli. Rope to nope and back again: A new hybrid attention strategy. arXiv, 2025. doi:10.48550/arXiv.2501.18795. URLhttp://arxiv.org/abs/2501.18795
-
[41]
Extending the context of pretrained llms by dropping their positional embeddings
Y oav Gelberg, Koshi Eguchi, Takuya Akiba, and Edoardo Cetin. Extending the context of pretrained LLMs by dropping their positional embeddings. arXiv, 2025. doi: 10.48550/arXiv.2512.12167. URLhttp://arxiv.org/abs/2512.12167
-
[42]
Johannes Linder, Divyanshi Srivastava, Han Yuan, Vikram Agarwal, and David R. Kelley. Predicting RNA-seq coverage from DNA sequence as a unifying model of gene regulation.Nat Genet, 57(4):949–961, 2025. ISSN 1546-1718. doi:10.1038/s41588-024-02053-6
-
[43]
Lorbeer, Chandana Rajesh, Tristan Karch, et al
Sam Boshar, Benjamin Evans, Ziqi Tang, Armand Picard, Y anis Adel, Franziska K. Lorbeer, Chandana Rajesh, Tristan Karch, et al. A foundational model for joint sequence-function multi-species modeling at scale for long-range genomic prediction. bioRxiv, 2025. doi:10.64898/2025.12.22.695963. URLhttps: //www.biorxiv.org/content/10.64898/2025.12.22.695963v1. ...
-
[44]
Flex attention: A programming model for generating optimized attention kernels
Juechu Dong, Boyuan Feng, Driss Guessous, Y anbo Liang, and Horace He. Flex attention: A programming model for generating optimized attention kernels. arXiv,
-
[45]
doi:10.48550/arXiv.2412.05496. URLhttp://arxiv.org/abs/2412.05496. version: 1
-
[46]
PyTorch: An Imperative Style, High-Performance Deep Learning Library
Adam Paszke, Sam Gross, Francisco Massa, Adam Lerer, James Bradbury, Gregory Chanan, Trevor Killeen, Zeming Lin, et al. PyTorch: An imperative style, high- performance deep learning library. arXiv, 2019. doi:10.48550/arXiv.1912.01703. URLhttp://arxiv.org/abs/1912.01703
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.1912.01703 2019
-
[47]
The FineWeb Datasets: Decanting the Web for the Finest Text Data at Scale
Guilherme Penedo, Hynek Kydlí ˇcek, Loubna Ben allal, Anton Lozhkov, Margaret Mitchell, Colin Raffel, Leandro Von Werra, and Thomas Wolf. The FineWeb datasets: Decanting the web for the finest text data at scale. arXiv, 2024. doi:10.48550/arXiv.2406.17557. URLhttp://arxiv.org/abs/2406.17557
work page internal anchor Pith review doi:10.48550/arxiv.2406.17557 2024
-
[48]
The OMG dataset: An open MetaGenomic corpus for mixed-modality genomic language modeling
Andre Cornman, Jacob West-Roberts, Antonio Pedro Camargo, Simon Roux, Martin Beracochea, Milot Mirdita, Sergey Ovchinnikov, and Yunha Hwang. The OMG dataset: An open MetaGenomic corpus for mixed-modality genomic language modeling. bioRxiv, 2024. doi:10.1101/2024.08.14.607850. URLhttps://www. biorxiv.org/content/10.1101/2024.08.14.607850v2. Pages: 2024.08....
-
[49]
U-Net: Convolutional Networks for Biomedical Image Segmentation
Olaf Ronneberger, Philipp Fischer, and Thomas Brox. U-net: Convolutional networks for biomedical image segmentation. arXiv, 2015. doi:10.48550/arXiv.1505.04597. URLhttp://arxiv.org/abs/1505.04597
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.1505.04597 2015
-
[50]
All are worth words: A ViT backbone for diffusion models
Fan Bao, Shen Nie, Kaiwen Xue, Yue Cao, Chongxuan Li, Hang Su, and Jun Zhu. All are worth words: A ViT backbone for diffusion models. arXiv, 2023. doi: 10.48550/arXiv.2209.12152. URLhttp://arxiv.org/abs/2209.12152
-
[51]
KellerJordan/modded-nanogpt
Keller Jordan. KellerJordan/modded-nanogpt. 2026. URLhttps://github.com/KellerJordan/modded-nanogpt. original-date: 2024-06-01T06:01:50Z
2026
-
[52]
Gleghorn-lab/SpeedrunningPLMs
Hallee, Logan. Gleghorn-lab/SpeedrunningPLMs. Gleghorn Lab, Synthyra, 2026. URLhttps://github.com/Gleghorn-Lab/SpeedrunningPLMs. original- date: 2024-12-20T18:47:28Z
2026
-
[53]
Neural Machine Translation of Rare Words with Subword Units
Rico Sennrich, Barry Haddow, and Alexandra Birch. Neural machine translation of rare words with subword units. arXiv, 2016. doi:10.48550/arXiv.1508.07909. URL http://arxiv.org/abs/1508.07909
work page internal anchor Pith review doi:10.48550/arxiv.1508.07909 2016
-
[54]
Muon: An optimizer for hidden layers in neural networks | keller jordan blog. 2024. URLhttps://kellerjordan.github.io/posts/muon/
2024
-
[55]
Kimi K2: Open Agentic Intelligence
Kimi Team, Yifan Bai, Yiping Bao, Y . Charles, Cheng Chen, Guanduo Chen, Haiting Chen, Huarong Chen, et al. Kimi k2: Open agentic intelligence. arXiv, 2026. doi: 10.48550/arXiv.2507.20534. URLhttp://arxiv.org/abs/2507.20534
work page internal anchor Pith review doi:10.48550/arxiv.2507.20534 2026
-
[56]
Gaussian Error Linear Units (GELUs)
Dan Hendrycks and Kevin Gimpel. Gaussian error linear units (GELUs). arXiv, 2023. doi:10.48550/arXiv.1606.08415. URLhttp://arxiv.org/abs/1606.08415
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.1606.08415 2023
-
[57]
GLU Variants Improve Transformer
Noam Shazeer. GLU variants improve transformer. arXiv, 2020. doi:10.48550/arXiv.2002.05202. URLhttp://arxiv.org/abs/2002.05202. Halleeet al.| arXiv | April 22, 2026 | 9–12
work page internal anchor Pith review doi:10.48550/arxiv.2002.05202 2020
-
[58]
Jimmy Lei Ba, Jamie Ryan Kiros, and Geoffrey E. Hinton. Layer normalization. arXiv, 2016. doi:10.48550/arXiv.1607.06450. URLhttp://arxiv.org/abs/1607. 06450
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.1607.06450 2016
-
[59]
Decoupled Weight Decay Regularization
Ilya Loshchilov and Frank Hutter. Decoupled weight decay regularization. arXiv, 2019. doi:10.48550/arXiv.1711.05101. URLhttp://arxiv.org/abs/1711. 05101
work page internal anchor Pith review Pith/arXiv arXiv doi:10.48550/arxiv.1711.05101 2019
-
[60]
Mixed precision training
Paulius Micikevicius, Sharan Narang, Jonah Alben, Gregory Diamos, Erich Elsen, David Garcia, Boris Ginsburg, Michael Houston, et al. Mixed precision training. arXiv,
-
[61]
doi:10.48550/arXiv.1710.03740. URLhttp://arxiv.org/abs/1710.03740
work page internal anchor Pith review doi:10.48550/arxiv.1710.03740
-
[62]
B. W. Matthews. Comparison of the predicted and observed secondary structure of t4 phage lysozyme.Biochimica et Biophysica Acta (BBA) - Protein Structure, 405 (2):442–451, 1975. ISSN 0005-2795. doi:10.1016/0005-2795(75)90109-9
-
[63]
B. L. Welch. The generalization of ‘Student’s’ problem when several different population variances are involved.Biometrika, 34(1-2):28–35, 1947. ISSN 0006-3444. doi: 10.1093/biomet/34.1-2.28. Halleeet al.| arXiv | April 22, 2026 | 10–12 Supplementary information Dual Triangle Attention pseudocode We provide pseudocode for both a naive implementation and t...
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.