Recognition: 2 theorem links
· Lean TheoremGoForth: Language Models for RNA Design under Structure, Sequence, and Coding Constraints
Pith reviewed 2026-05-11 02:11 UTC · model grok-4.3
The pith
Encoder-decoder models trained on real RNA folds generate sequences satisfying mixed structure, sequence, and coding constraints.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
GoForth is a forward-trained RNA language model that conditions on structure, sequence, and coding targets. The basic object is a conditional law over RNA sequences given a user-specified condition, with full inverse folding as a special case. We train encoder-decoder models on witnessed folds rather than on outputs from an inverse-design teacher and validate our methodology on full inverse-folding benchmarks, as well as tasks involving constraints on structure, sequence, and coding. The resulting models achieve fast and high-quality candidate generation for mixed RNA design specifications. Moreover they furnish useful semantic embeddings of design tasks and a robust learned notion of design
What carries the argument
The GoForth conditional encoder-decoder model, trained on witnessed folds to learn distributions over sequences under combined structure, sequence, and coding constraints.
If this is right
- Fast and high-quality candidate generation for mixed RNA design specifications.
- Useful semantic embeddings of design tasks.
- A robust learned notion of designability.
- Effective performance on full inverse-folding benchmarks and constrained tasks.
Where Pith is reading between the lines
- The method could scale to larger RNA molecules or other biopolymers by expanding the training data.
- Learned embeddings might support automated design space exploration or transfer learning across different constraint types.
- Integration with experimental validation loops could use the designability score to guide iterations.
Load-bearing premise
That models trained on observed RNA folds generalize to generate valid sequences for any arbitrary combination of structure, sequence, and coding constraints without additional filtering or retraining.
What would settle it
Finding that on a benchmark of mixed constraints, the generated sequences violate the specified conditions at rates comparable to random sampling or require extensive filtering to achieve high quality.
Figures
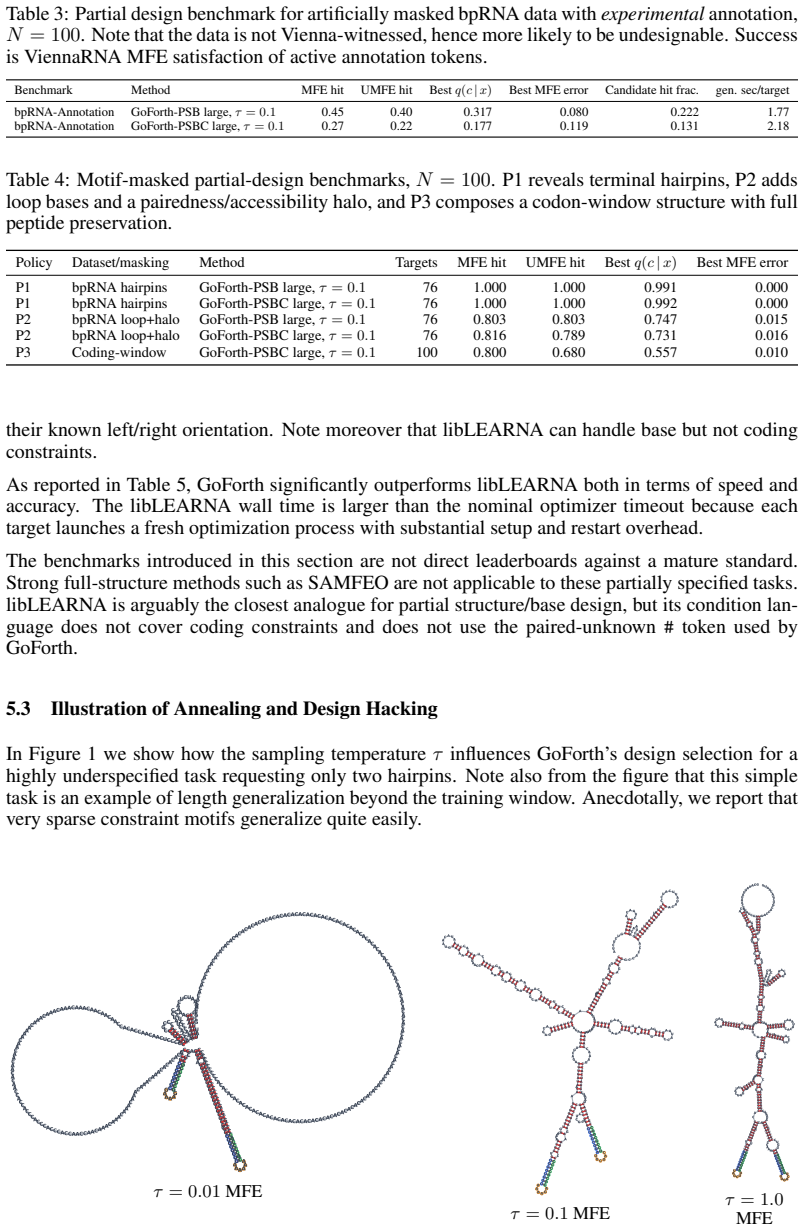




read the original abstract
RNA inverse sequence design has broad biological and engineering applications, but computational methods for practical design queries remain limited. Such queries may impose several constraints at once, including target folds or motifs, fixed bases, and coding restrictions, while leaving arbitrary sequence and structure in unspecified regions. Because these constraints may permit many acceptable sequences, we study RNA design as a conditional generative modeling problem. The basic object is a conditional law over RNA sequences given a user-specified condition, with full inverse folding as a special case. We introduce GoForth, a forward-trained RNA language model that conditions on structure, sequence, and coding targets. The formulation separates three ingredients that are often entangled in RNA design: a sequence prior, a forward folding sampler, and a reward or likelihood oracle. We train encoder-decoder models on witnessed folds rather than on outputs from an inverse-design teacher and validate our methodology on full inverse-folding benchmarks, as well as tasks involving constraints on structure, sequence, and coding. The resulting models achieve fast and high-quality candidate generation for mixed RNA design specifications. Moreover they furnish useful semantic embeddings of design tasks and a robust learned notion of designability.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The manuscript introduces GoForth, a forward-trained encoder-decoder RNA language model for conditional sequence generation under user-specified combinations of structure, sequence, and coding constraints. It frames RNA inverse design as conditional generative modeling, separates the sequence prior, forward folding sampler, and reward/likelihood oracle, trains exclusively on witnessed natural folds rather than inverse-design teacher outputs, and reports validation on standard inverse-folding benchmarks plus mixed-constraint tasks, claiming fast high-quality candidate generation together with useful semantic embeddings of design tasks and a learned notion of designability.
Significance. If the central claims hold, the separation of prior/sampler/oracle components and training on witnessed folds would provide a clean, extensible framework for RNA design that avoids entanglement common in prior methods and supplies independent grounding. The resulting embeddings and designability measure could support downstream biological applications, while the fast generation under mixed constraints addresses a practical gap in handling arbitrary user-specified specifications.
major comments (2)
- [Abstract] Abstract and results sections: the central claim that encoder-decoder models trained only on witnessed folds directly produce valid sequences for arbitrary mixed constraints (without post-hoc filtering or retraining) is load-bearing yet unsupported by any reported quantitative metrics, ablation studies, or explicit verification that all constraints are satisfied simultaneously on out-of-distribution combinations; the abstract states validation occurs but provides no numbers, error analysis, or evidence isolating generalization from implicit rejection sampling.
- [Methods] Methods and evaluation: the separation into prior, sampler, and oracle is asserted as enabling the result, but no explicit description or experiment demonstrates that the learned conditional law covers user-specified constraint combinations outside the training distribution without requiring additional filtering steps or oracle calls at inference time.
minor comments (2)
- [Abstract] The abstract mentions 'full inverse-folding benchmarks' and 'tasks involving constraints on structure, sequence, and coding' but does not name the specific benchmarks or datasets used, which would aid reproducibility.
- [Introduction] Notation for the conditional law and the three separated ingredients (prior, sampler, oracle) is introduced but not formalized with equations or pseudocode, making the claimed separation harder to follow.
Simulated Author's Rebuttal
We thank the referee for their constructive and detailed comments, which have identified important opportunities to strengthen the clarity and evidentiary support in our manuscript. We address each major comment point by point below, indicating the revisions we will undertake.
read point-by-point responses
-
Referee: [Abstract] Abstract and results sections: the central claim that encoder-decoder models trained only on witnessed folds directly produce valid sequences for arbitrary mixed constraints (without post-hoc filtering or retraining) is load-bearing yet unsupported by any reported quantitative metrics, ablation studies, or explicit verification that all constraints are satisfied simultaneously on out-of-distribution combinations; the abstract states validation occurs but provides no numbers, error analysis, or evidence isolating generalization from implicit rejection sampling.
Authors: We agree that the abstract would be improved by including specific quantitative metrics. In the revised manuscript we will update the abstract to report key figures from our experiments, including success rates for simultaneous satisfaction of structure, sequence, and coding constraints on mixed tasks. The results sections already contain quantitative evaluations on standard inverse-folding benchmarks (sequence recovery and structure accuracy) and mixed-constraint tasks (constraint satisfaction rates verified by external folding and coding checks). We will add an ablation comparing training on witnessed folds versus teacher-generated inverse-design data and expand the error analysis with a breakdown of per-constraint and joint satisfaction rates on held-out combinations that were not observed during training. These held-out sets serve as out-of-distribution test cases, and satisfaction is measured independently of the model likelihood, thereby isolating generalization from any implicit rejection sampling. revision: yes
-
Referee: [Methods] Methods and evaluation: the separation into prior, sampler, and oracle is asserted as enabling the result, but no explicit description or experiment demonstrates that the learned conditional law covers user-specified constraint combinations outside the training distribution without requiring additional filtering steps or oracle calls at inference time.
Authors: Section 3 of the manuscript describes the encoder-decoder architecture that receives the user-specified combination of structure, sequence, and coding constraints as input and is trained to model the conditional distribution over sequences given those constraints. Training uses only natural witnessed folds paired with their sequences; no inverse-design teacher outputs are involved. At inference the model directly samples from this conditional distribution by conditioning on the encoded constraints, with no filtering or oracle calls performed during generation. To address the referee's concern we will add a dedicated subsection in Methods that walks through the inference procedure step by step and will include a new experiment that generates sequences for novel constraint combinations absent from the training distribution, reporting the fraction of outputs that satisfy all constraints simultaneously as verified by independent external tools. This will explicitly demonstrate coverage of out-of-distribution combinations without post-hoc steps. revision: partial
Circularity Check
No significant circularity; training on external witnessed folds supplies independent grounding
full rationale
The paper trains encoder-decoder models directly on witnessed (natural) folds as the data source and evaluates generated candidates on standard inverse-folding benchmarks plus constraint tasks. No equations, parameters, or central claims are shown to reduce by construction to quantities fitted from the target result itself. The separation into prior, sampler, and oracle is presented as a modeling choice with external data grounding rather than a self-referential loop. No self-citations are invoked as load-bearing uniqueness theorems or ansatzes. This is the standard non-circular pattern for a conditional generative modeling paper.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption RNA sequences admit a conditional generative distribution given partial structure, sequence, and coding constraints
Lean theorems connected to this paper
-
IndisputableMonolith/Cost/FunctionalEquation.leanwashburn_uniqueness_aczel unclearWe train encoder-decoder models on witnessed folds rather than on outputs from an inverse-design teacher... The formulation separates three ingredients... a sequence prior, a forward folding sampler, and a reward or likelihood oracle.
-
IndisputableMonolith/Foundation/RealityFromDistinction.leanreality_from_one_distinction unclearGoForth uses a sequence of condition tokens... full inverse folding as one slice rather than the defining case.
Reference graph
Works this paper leans on
- [1]
-
[2]
Advances in Neural Information Processing Systems , volume =
Attention Is All You Need , author =. Advances in Neural Information Processing Systems , volume =. 2017 , url =
work page 2017
-
[3]
Shao, Zhihong and Wang, Peiyi and Zhu, Qihao and Xu, Runxin and Song, Junxiao and Bi, Xiao and Zhang, Haowei and Zhang, Mingchuan and Li, Y. K. and Wu, Y. and Guo, Daya , year =. 2402.03300 , archivePrefix =
work page internal anchor Pith review Pith/arXiv arXiv
-
[4]
Adamczyk, Bartosz and Antczak, Maciej and Szachniuk, Marta , journal =. 2022 , doi =
work page 2022
-
[5]
Danaee, Padideh and Rouches, Mason and Wiley, Michelle and Deng, Dezhong and Huang, Liang and Hendrix, David , journal =. 2018 , doi =
work page 2018
-
[6]
Kalvari, Ioanna and Nawrocki, Eric P. and Ontiveros-Palacios, Nancy and Argasinska, Joanna and Lamkiewicz, Kevin and Marz, Manja and Griffiths-Jones, Sam and Toffano-Nioche, Claire and Gautheret, Daniel and Weinberg, Zasha and Rivas, Elena and Eddy, Sean R. and Finn, Robert D. and Bateman, Alex and Petrov, Anton I. , journal =. 2021 , doi =
work page 2021
-
[7]
Gao, Letian and Lu, Zhi John , year =. 2407.19838 , archivePrefix =
-
[8]
Cole, Jack and Li, Fan and Wu, Liwen and Li, Ke , booktitle =. 2024 , url =
work page 2024
-
[9]
and Honer zu Siederdissen, Christian and Tafer, Hakim and Flamm, Christoph and Stadler, Peter F
Lorenz, Ronny and Bernhart, Stephan H. and Honer zu Siederdissen, Christian and Tafer, Hakim and Flamm, Christoph and Stadler, Peter F. and Hofacker, Ivo L. , journal =. 2011 , doi =
work page 2011
-
[10]
Zadeh, Joseph N. and Steenberg, Conrad D. and Bois, Justin S. and Wolfe, Brian R. and Pierce, Marshall B. and Khan, Amardeep R. and Dirks, Robert M. and Pierce, Niles A. , journal =. 2011 , doi =
work page 2011
-
[11]
Eastman, Peter and Shi, Jade and Ramsundar, Bharath and Pande, Vijay S. , journal =. Solving the. 2018 , doi =
work page 2018
-
[12]
Runge, Frederic and Stoll, Danny and Falkner, Stefan and Hutter, Frank , booktitle =. Learning to Design. 2019 , url =
work page 2019
-
[13]
and Keep, Benjamin and Coppess, Katherine R
Koodli, Rohan V. and Keep, Benjamin and Coppess, Katherine R. and Portela, Fernando and. PLOS Computational Biology , volume =. 2019 , doi =
work page 2019
-
[14]
and Mechery, Vinodh and Meyer, Michelle M
Dotu, Ivan and Garcia-Martin, Juan Antonio and Slinger, Betty L. and Mechery, Vinodh and Meyer, Michelle M. and Clote, Peter , journal =. Complete. 2014 , doi =
work page 2014
-
[15]
Huang, Han and Lin, Ziqian and He, Dongchen and Hong, Liang and Li, Yu , journal =. 2024 , doi =
work page 2024
-
[16]
Kleinkauf, Robert and Mann, Martin and Backofen, Rolf , journal =. 2015 , doi =
work page 2015
-
[17]
and Huang, Liang , booktitle =
Zhou, Tianshuo and Tang, Wei Yu and Mathews, David H. and Huang, Liang , booktitle =. Undesignable. 2024 , publisher =
work page 2024
-
[18]
Zhou, Tianshuo and Mathews, David H. and Huang, Liang , year =. Fast and Versatile. 2603.02283 , archivePrefix =
-
[19]
Chen, Jiayang and Hu, Zhihang and Sun, Siqi and Tan, Qingxiong and Wang, Yixuan and Yu, Qinze and Zong, Licheng and Hong, Liang and Xiao, Jin and Shen, Tao and King, Irwin and Li, Yu , year =. Interpretable. 2204.00300 , archivePrefix =
-
[20]
Nature Communications , volume =
Peni. Nature Communications , volume =. 2025 , doi =
work page 2025
- [21]
- [22]
-
[23]
Journal of Computational Chemistry , volume =
Nucleic Acid Sequence Design via Efficient Ensemble Defect Optimization , author =. Journal of Computational Chemistry , volume =. 2011 , doi =
work page 2011
-
[24]
Journal of the American Chemical Society , volume =
Constrained Multistate Sequence Design for Nucleic Acid Reaction Pathway Engineering , author =. Journal of the American Chemical Society , volume =. 2017 , doi =
work page 2017
-
[25]
and Hutter, Frank and Hoos, Holger H
Andronescu, Mirela and Fejes, Anthony P. and Hutter, Frank and Hoos, Holger H. and Condon, Anne , journal =. A New Algorithm for. 2004 , doi =
work page 2004
- [26]
-
[27]
Garcia-Martin, Juan Antonio and Clote, Peter and Dotu, Ivan , journal =. 2013 , doi =
work page 2013
- [28]
- [29]
-
[30]
A Weighted Sampling Algorithm for the Design of
Reinharz, Vladimir and Ponty, Yann and Waldisp. A Weighted Sampling Algorithm for the Design of. Bioinformatics , volume =. 2013 , doi =
work page 2013
-
[31]
Yang, Xiufeng and Yoshizoe, Kazuki and Taneda, Akito and Tsuda, Koji , journal =. 2017 , doi =
work page 2017
-
[32]
An Unexpectedly Effective Monte Carlo Technique for the
Portela, Fernando , year =. An Unexpectedly Effective Monte Carlo Technique for the
-
[33]
Rubio-Largo,. Solving the. Applied Soft Computing , volume =. 2023 , doi =
work page 2023
-
[34]
and Lach, Grzegorz and Badepally, Nagendar Goud and Moafinejad, S
Wirecki, Tomasz K. and Lach, Grzegorz and Badepally, Nagendar Goud and Moafinejad, S. Naeim and Jaryani, Farhang and Klaudel, Gaja and Nec, Kalina and Baulin, Eugene F. and Bujnicki, Janusz M. , journal =. 2025 , doi =
work page 2025
-
[35]
Zhou, Tianshuo and Dai, Ning and Li, Sizhen and Ward, Max and Mathews, David H. and Huang, Liang , journal =. 2023 , doi =
work page 2023
-
[36]
Tang, Wei Yu and Dai, Ning and Zhou, Tianshuo and Mathews, David H. and Huang, Liang , journal =. 2026 , doi =
work page 2026
-
[37]
Anderson-Lee, Jeff and Fisker, Eli and Kosaraju, Vineet and Wu, Michelle and Kong, Justin and Lee, Jeehyung and Lee, Minjae and Zada, Mathew and Treuille, Adrien and Das, Rhiju , journal =. Principles for Predicting. 2016 , doi =
work page 2016
- [38]
-
[39]
Ward, Max and Courtney, Eliot and Rivas, Elena , journal =. Fitness Functions for. 2023 , doi =
work page 2023
-
[40]
Zhou, Tianshuo and Tang, Wei Yu and Malik, Apoorv and Mathews, David H. and Huang, Liang , year =. Scalable and Interpretable Identification of Minimal Undesignable. 2402.17206 , archivePrefix =
-
[41]
Probabilistic RNA Designability via Interpretable Ensemble Approximation and Dynamic Decomposition
Zhou, Tianshuo and Mathews, David H. and Huang, Liang , year =. Probabilistic. 2602.13610 , archivePrefix =
work page internal anchor Pith review Pith/arXiv arXiv
-
[42]
and Rudolfs, Boris and Romano, Jonathan and Wayment-Steele, Hannah K
Koodli, Rohan V. and Rudolfs, Boris and Romano, Jonathan and Wayment-Steele, Hannah K. and Dunlap, William A. and. Redesigning the. 2021 , doi =
work page 2021
-
[43]
Runge, Frederic and Farid, Karim and Franke, J. 2024 , publisher =
work page 2024
-
[44]
Runge, Frederic and Franke, J. Partial. Bioinformatics , volume =. 2024 , doi =
work page 2024
-
[45]
Hammer, Stefan and Tschiatschek, Birgit and Flamm, Christoph and Hofacker, Ivo L. and Stadler, Peter F. and Will, Sebastian , journal =. 2017 , doi =
work page 2017
-
[46]
Fixed-Parameter Tractable Sampling for
Hammer, Stefan and Wang, Wei and Will, Sebastian and Ponty, Yann , journal =. Fixed-Parameter Tractable Sampling for. 2019 , doi =
work page 2019
- [47]
- [48]
- [49]
-
[50]
Akiyama, Manato and Sakakibara, Yasubumi , journal =. Informative. 2022 , doi =
work page 2022
-
[51]
Multiple Sequence Alignment-Based
Zhang, Yiqun and Lang, Mingyang and Jiang, Wenjun and others , journal =. Multiple Sequence Alignment-Based. 2024 , doi =
work page 2024
-
[52]
Sumi, Shunsuke and Hamada, Michiaki and Saito, Hirohide , journal =. Deep Generative Design of. 2024 , doi =
work page 2024
-
[53]
Zhao, Yuwen and Oono, Kentaro and Takizawa, Hiroaki and Kotera, Masaaki , journal =. 2024 , doi =
work page 2024
-
[54]
Ozden, Furkan and Barazandeh, Sina and Akboga, Dogus and Tabrizi, Sobhan Shokoueian and Seker, Urartu Ozgur Safak and Cicek, A. Ercument , year =
-
[55]
Shukueian Tabrizi, Sobhan and Barazandeh, Sina and Hashemi Aghdam, Hamed and Cicek, A. Ercument , journal =. 2025 , doi =
work page 2025
-
[56]
Klypa, Roman and Bietti, Alberto and Grudinin, Sergei , booktitle =. 2025 , publisher =
work page 2025
-
[57]
Wang, Zhen and Liu, Ziqi and Zhang, Wei and Li, Yanjun and Feng, Yizhen and Lv, Shaokang and Diao, Han and Luo, Zhaofeng and Yan, Pengju and He, Min and Li, Xiaolin , journal =. 2024 , doi =
work page 2024
-
[58]
Wong, Felix and He, Dongchen and Krishnan, Aarti and Hong, Liang and Wang, Alexander Z. and Wang, Jiuming and Hu, Zhihang and Omori, Satotaka and Li, Alicia and Rao, Jiahua and Yu, Qinze and Jin, Wengong and Zhang, Tianqing and Ilia, Katherine and Chen, Jack X. and Zheng, Shuangjia and King, Irwin and Li, Yu and Collins, James J. , journal =. Deep Generat...
work page 2024
-
[59]
and Zhang, Yujian and Huang, Liang , journal =
Zhang, He and Zhang, Liang and Lin, Ang and Xu, Congcong and Li, Ziyu and Liu, Kaibo and Liu, Boxiang and Ma, Xiaopin and Zhao, Fanfan and Jiang, Huiling and Chen, Chunxiu and Shen, Haifa and Li, Hangwen and Mathews, David H. and Zhang, Yujian and Huang, Liang , journal =. Algorithm for Optimized. 2023 , doi =
work page 2023
-
[60]
Li, Sizhen and Moayedpour, Saeed and Li, Ruijiang and Bailey, Michael and Riahi, Saleh and Miladi, Milad and Miner, Jacob and Zheng, Dinghai and Wang, Jun and Balsubramani, Akshay and others , booktitle =. 2023 , url =
work page 2023
-
[61]
Dai, Ning and Tang, Wei Yu and Zhou, Tianshuo and Mathews, David H. and Huang, Liang , year =. Messenger. 2401.00037 , archivePrefix =
-
[62]
Dai, Ning and Zhou, Tianshuo and Tang, Wei Yu and Mathews, David H. and Huang, Liang , journal =. 2025 , doi =
work page 2025
-
[63]
Fornace, Mark and Wang, Christina Wuyan and Lindsey, Michael , year =. Direct. 2604.19718 , archivePrefix =
work page internal anchor Pith review Pith/arXiv arXiv
-
[64]
Joint Design of 5' Untranslated Region and Coding Sequence of
Liu, Yang and Gao, Jie and Zhang, Xiaonan and Fang, Xiaomin , year =. Joint Design of 5' Untranslated Region and Coding Sequence of. 2410.20781 , archivePrefix =
-
[65]
Zhang, He and Liu, Hailong and Xu, Yushan and Huang, Haoran and Liu, Yiming and Wang, Jia and Qin, Yan and Wang, Haiyan and Ma, Lili and Xun, Zhiyuan and Hou, Xuzhuang and Lu, Timothy K. and Cao, Jicong , journal =. Deep Generative Models Design. 2025 , doi =
work page 2025
-
[66]
Yue, Feipeng and Dai, Ning and Tang, Wei Yu and Zhou, Tianshuo and Mathews, David H. and Huang, Liang , year =. Sampling-Based Continuous Optimization for Messenger. 2603.06559 , archivePrefix =
-
[67]
Nori, Divya and Jin, Wengong , year =. 2405.18768 , archivePrefix =
-
[68]
Hu, Tianmeng and Cui, Yongzheng and Luo, Biao and Li, Ke , booktitle =. 2026 , url =
work page 2026
-
[69]
Sun, Minghao and Cao, Hanqun and Zhang, Zhou and Wei, Chen and Wang, Liang and Jia, Tianrui and Liu, Zhiyuan and Fu, Tianfan and Tang, Xiangru and Choi, Yejin and Heng, Pheng-Ann and Wu, Fang and Zhang, Yang , year =. 2510.21161 , archivePrefix =
-
[70]
Patil, Vaibhav and others , year =
-
[71]
Manzourolajdad, Amirhossein and Mohebbi, Mohammad , journal =. Secondary-Structure-Informed. 2025 , doi =
work page 2025
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.