Recognition: 2 theorem links
· Lean TheoremPushing Biomolecular Utility-Diversity Frontiers with Supergroup Relative Policy Optimization
Pith reviewed 2026-05-12 01:06 UTC · model grok-4.3
The pith
Supergroup Relative Policy Optimization expands the utility-diversity Pareto frontier for biomolecular generators by directly rewarding set-level diversity.
A machine-rendered reading of the paper's core claim, the machinery that carries it, and where it could break.
Core claim
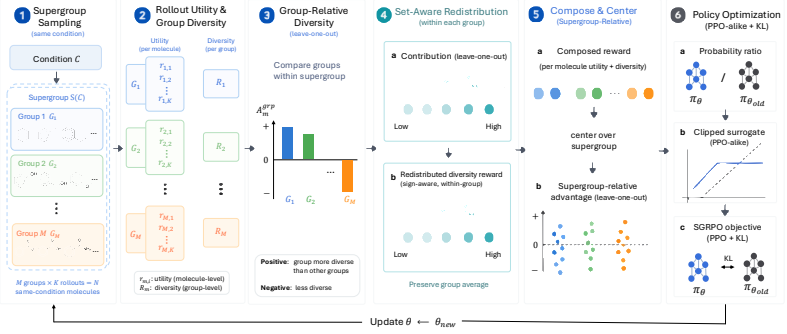
SGRPO constructs rewards from set-level diversity by sampling a supergroup of candidate sets under the same condition, comparing their diversity, and redistributing the group diversity reward to individual rollouts through leave-one-out diversity contributions before combining it with rollout-level utility. This design decouples the method from any particular generator, utility reward, or diversity metric and allows instantiation with different GRPO-style approaches.
What carries the argument
Supergroup sampling with leave-one-out diversity contributions, which converts a set-level diversity score into per-rollout rewards that can be used directly in policy optimization.
If this is right
- SGRPO achieves the best frontier-level metrics across decoding sweeps relative to pretrained generators, GRPO, and memory-assisted GRPO.
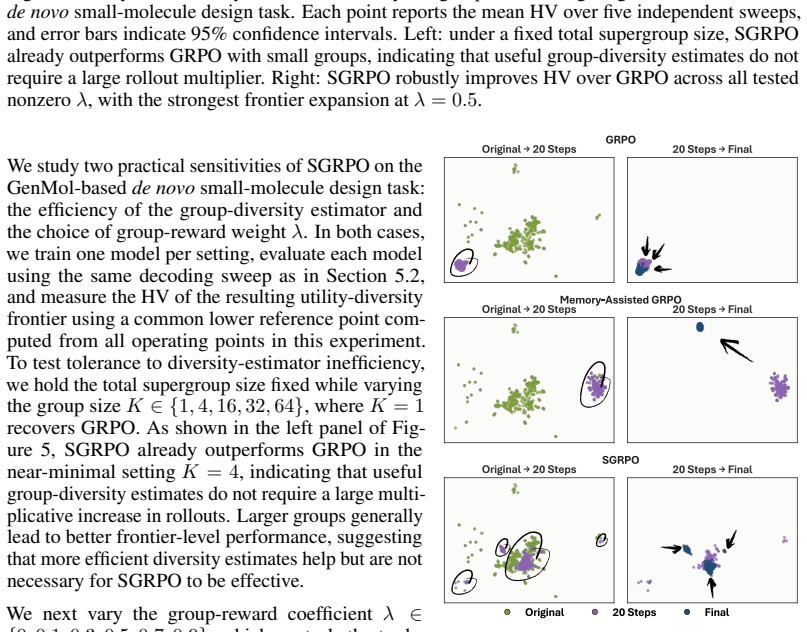
- Direct set-level diversity rewards remain effective even with small supergroup sizes.
- The approach helps preserve broader coverage of the generation distribution during post-training.
Where Pith is reading between the lines
- The same supergroup construction could be tested on non-biomolecular generators where diversity is also a set-level property.
- Optimal supergroup size may vary with task and generator type and could be tuned empirically.
- Combining SGRPO with other set-level objectives beyond diversity would be a direct next step.
Load-bearing premise
That comparing diversity across supergroups of candidate sets and redistributing via leave-one-out contributions gives a stable, unbiased signal that does not introduce artifacts or depend on the particular diversity metric chosen.
What would settle it
An experiment that replaces the leave-one-out redistribution with uniform random assignment of the group diversity score and finds that the Pareto-frontier improvement over GRPO baselines disappears.
Figures



read the original abstract
Biomolecular generators are often adapted with reward feedback to improve task-specific utility, but pushing utility alone can concentrate generation on a narrow family of candidates. Maintaining diversity is difficult because sample diversity is a set-level property. We introduce Supergroup Relative Policy Optimization (SGRPO), a flexible GRPO-style framework that directly constructs rewards from set-level diversity. For each condition, SGRPO samples a supergroup of candidate sets, compares their diversity under the same condition, and redistributes the group diversity reward to individual rollouts through leave-one-out diversity contributions before combining it with rollout-level utility. This design decouples SGRPO from a particular generator, utility reward, or diversity metric, and allows instantiation with different GRPO-style approaches. We evaluate SGRPO on de novo small-molecule design, pocket-based small-molecule design, and de novo protein design, instantiating it with both GRPO and Coupled-GRPO across autoregressive and discrete diffusion generators. Across decoding sweeps, SGRPO expands the utility-diversity Pareto frontier and achieves the best frontier-level metrics relative to pretrained generators, GRPO, and memory-assisted GRPO when applicable. Our analyses further show that direct set-level diversity rewards remain effective with small groups and help preserve broader generation-distribution coverage during post-training. The code is available at https://github.com/IDEA-XL/SGRPO.
Editorial analysis
A structured set of objections, weighed in public.
Referee Report
Summary. The paper introduces Supergroup Relative Policy Optimization (SGRPO), a GRPO-style RL framework for biomolecular generators that addresses set-level diversity by sampling supergroups of candidate sets per condition, computing group-level diversity scores, redistributing them to individual rollouts via leave-one-out contributions, and combining with per-rollout utility rewards. It instantiates the method with GRPO and Coupled-GRPO on autoregressive and discrete diffusion models for de novo small-molecule design, pocket-based design, and de novo protein design. Across decoding sweeps, SGRPO is reported to expand the utility-diversity Pareto frontier and outperform pretrained generators, standard GRPO, and memory-assisted GRPO on frontier-level metrics while preserving broader distributional coverage; the approach is claimed to remain effective with small supergroups and to be decoupled from specific generators, utility functions, or diversity metrics. Code is released.
Significance. If the central claims hold, SGRPO offers a modular, metric-agnostic way to incorporate set-level diversity signals into policy optimization for generative models in computational biology and chemistry. This directly targets the common failure mode where utility-only RL collapses generation to narrow families. The empirical breadth across three tasks and two generator families, plus explicit code release, would make the contribution practically useful for downstream biomolecular design workflows.
major comments (2)
- [Method description and experimental analyses] The leave-one-out redistribution of the supergroup diversity score (full-group diversity minus diversity without the focal rollout) induces strong negative dependence among the per-rollout diversity rewards within each supergroup. This dependence is load-bearing for the combined advantage used in the GRPO-style update, yet the manuscript provides no analysis of the resulting reward variance, gradient stability, or sensitivity to supergroup size and diversity metric choice. The abstract asserts effectiveness with small groups, but without quantitative checks (e.g., variance of the diversity component or ablation on group size), it is unclear whether the reported Pareto improvements are robust or partly artifacts of the correlation structure.
- [Experimental results and figures] The central empirical claim—that SGRPO expands the utility-diversity frontier relative to GRPO and memory-assisted GRPO—rests on decoding sweeps and frontier-level metrics, but the abstract and available text give no details on statistical significance testing, exact baseline implementations, or how post-hoc selection of decoding parameters was avoided. Without these, it is difficult to assess whether the reported gains are reproducible or driven by implementation differences.
minor comments (2)
- [Method] Notation for supergroup sampling and leave-one-out computation should be formalized with explicit equations to make the redistribution step unambiguous.
- [Experiments] The diversity metric(s) used in the reported experiments should be stated explicitly, along with any ablations showing invariance to metric choice.
Simulated Author's Rebuttal
We thank the referee for the constructive and detailed feedback. We address each major comment below and have revised the manuscript to incorporate additional analyses and clarifications where the comments identify opportunities for strengthening the presentation.
read point-by-point responses
-
Referee: [Method description and experimental analyses] The leave-one-out redistribution of the supergroup diversity score (full-group diversity minus diversity without the focal rollout) induces strong negative dependence among the per-rollout diversity rewards within each supergroup. This dependence is load-bearing for the combined advantage used in the GRPO-style update, yet the manuscript provides no analysis of the resulting reward variance, gradient stability, or sensitivity to supergroup size and diversity metric choice. The abstract asserts effectiveness with small groups, but without quantitative checks (e.g., variance of the diversity component or ablation on group size), it is unclear whether the reported Pareto improvements are robust or partly artifacts of the correlation structure.
Authors: We agree that the leave-one-out construction deliberately introduces negative dependence to isolate marginal contributions, and that this structure merits explicit examination. The manuscript does report that set-level rewards remain effective with small supergroups (via the analyses referenced in the abstract and experimental results), but we acknowledge the absence of direct quantification of reward variance or gradient stability. In the revised manuscript we will add (i) an ablation varying supergroup size (2–8) with reported variance of the diversity reward component across multiple seeds, (ii) a short discussion of how the induced correlation affects advantage estimates, and (iii) sensitivity checks across the diversity metrics used. These additions will clarify that the observed Pareto gains are not artifacts of the correlation structure. revision: yes
-
Referee: [Experimental results and figures] The central empirical claim—that SGRPO expands the utility-diversity frontier relative to GRPO and memory-assisted GRPO—rests on decoding sweeps and frontier-level metrics, but the abstract and available text give no details on statistical significance testing, exact baseline implementations, or how post-hoc selection of decoding parameters was avoided. Without these, it is difficult to assess whether the reported gains are reproducible or driven by implementation differences.
Authors: We accept that the current text does not supply sufficient detail on these points. In the revision we will (i) report statistical significance (error bars and, where appropriate, p-values from repeated runs with different random seeds), (ii) provide precise descriptions of baseline implementations including hyperparameter matching and any memory-buffer details, and (iii) explicitly state that the reported frontiers derive from a fixed, exhaustive decoding-parameter grid without post-hoc selection. These clarifications will be added to the experimental section and figure captions. revision: yes
Circularity Check
SGRPO introduces independent supergroup sampling and leave-one-out redistribution without reducing to self-definition or fitted inputs
full rationale
The paper defines SGRPO explicitly as a GRPO-style extension that samples supergroups, computes set-level diversity, and redistributes via leave-one-out contributions before adding utility rewards. This is a constructive algorithmic choice presented in the abstract, not derived from prior equations or self-citations in a load-bearing way. No equations or claims in the provided text equate a 'prediction' to a fitted parameter by construction, invoke uniqueness theorems from the same authors, or rename known results. Empirical evaluations on multiple design tasks compare against baselines without circular reduction. The framework remains self-contained against external benchmarks.
Axiom & Free-Parameter Ledger
axioms (1)
- domain assumption Set-level diversity can be meaningfully quantified and used as a reward signal in policy optimization.
Lean theorems connected to this paper
-
IndisputableMonolith/Cost/FunctionalEquation.leanwashburn_uniqueness_aczel unclearSGRPO samples a supergroup of candidate sets, compares their diversity under the same condition, and redistributes the group diversity reward to individual rollouts through leave-one-out diversity contributions before combining it with rollout-level utility.
-
IndisputableMonolith/Foundation/ArithmeticFromLogic.leanembed_injective unclearWe define the group-relative diversity signal as Agrp_m = Rm − 1/(M−1) Σ_{h≠m} Rh = M/(M−1)(Rm − R̄).
Reference graph
Works this paper leans on
-
[2]
Model-based reinforcement learning for biological sequence design
Christof Angermueller, David Dohan, David Belanger, Ramya Deshpande, Kevin Murphy, and Lucy Colwell. Model-based reinforcement learning for biological sequence design. In International conference on learning representations, 2019
work page 2019
-
[3]
Viraj Bagal, Rishal Aggarwal, PK Vinod, and U Deva Priyakumar. Molgpt: molecular generation using a transformer-decoder model.Journal of chemical information and modeling, 62(9): 2064–2076, 2021
work page 2064
-
[4]
Dávid Bajusz, Anita Rácz, and Károly Héberger. Why is tanimoto index an appropriate choice for fingerprint-based similarity calculations?Journal of cheminformatics, 7(1):20, 2015
work page 2015
-
[5]
Quantifying the chemical beauty of drugs.Nature chemistry, 4(2):90–98, 2012
G Richard Bickerton, Gaia V Paolini, Jérémy Besnard, Sorel Muresan, and Andrew L Hopkins. Quantifying the chemical beauty of drugs.Nature chemistry, 4(2):90–98, 2012
work page 2012
-
[6]
Esben Jannik Bjerrum, Christian Margreitter, Thomas Blaschke, Simona Kolarova, and Raquel López-Ríos de Castro. Faster and more diverse de novo molecular optimization with double- loop reinforcement learning using augmented smiles.Journal of Computer-Aided Molecular Design, 37(8):373–394, 2023
work page 2023
-
[7]
Thomas Blaschke, Ola Engkvist, Jürgen Bajorath, and Hongming Chen. Memory-assisted reinforcement learning for diverse molecular de novo design.Journal of cheminformatics, 12 (1):68, 2020
work page 2020
-
[8]
Design by adaptive sampling.arXiv preprint arXiv:1810.03714, 2018
David H Brookes and Jennifer Listgarten. Design by adaptive sampling.arXiv preprint arXiv:1810.03714, 2018
-
[9]
Egbert Castro, Abhinav Godavarthi, Julian Rubinfien, Kevin Givechian, Dhananjay Bhaskar, and Smita Krishnaswamy. Transformer-based protein generation with regularized latent space optimization.Nature Machine Intelligence, 4(10):840–851, 2022
work page 2022
-
[10]
Curiosity as a self- supervised method to improve exploration in de novo drug design
Mohamed-Amine Chadi, Hajar Mousannif, and Ahmed Aamouche. Curiosity as a self- supervised method to improve exploration in de novo drug design. In2023 International Conference on Information Technology Research and Innovation (ICITRI), pages 151–156. IEEE, 2023
work page 2023
-
[11]
Xiwei Cheng, Xiangxin Zhou, Yuwei Yang, Yu Bao, and Quanquan Gu. Decomposed direct preference optimization for structure-based drug design.arXiv preprint arXiv:2407.13981, 2024
-
[12]
Robust deep learning–based protein sequence design using proteinmpnn.Science, 378(6615):49–56, 2022
Justas Dauparas, Ivan Anishchenko, Nathaniel Bennett, Hua Bai, Robert J Ragotte, Lukas F Milles, Basile IM Wicky, Alexis Courbet, Rob J de Haas, Neville Bethel, et al. Robust deep learning–based protein sequence design using proteinmpnn.Science, 378(6615):49–56, 2022
work page 2022
-
[14]
Yasha Ektefaie, Olivia Viessmann, Siddharth Narayanan, Drew Dresser, J Mark Kim, and Armen Mkrtchyan. Reinforcement learning on structure-conditioned categorical diffusion for protein inverse folding.arXiv preprint arXiv:2410.17173, 2024
-
[15]
Peter Ertl and Ansgar Schuffenhauer. Estimation of synthetic accessibility score of drug-like molecules based on molecular complexity and fragment contributions.Journal of cheminfor- matics, 1(1):8, 2009
work page 2009
-
[16]
Vendy Fialková, Jiaxi Zhao, Kostas Papadopoulos, Ola Engkvist, Esben Jannik Bjerrum, Thierry Kogej, and Atanas Patronov. Libinvent: reaction-based generative scaffold decoration for in silico library design.Journal of Chemical Information and Modeling, 62(9):2046–2063, 2021. 11
work page 2046
-
[17]
Paul G Francoeur, Tomohide Masuda, Jocelyn Sunseri, Andrew Jia, Richard B Iovanisci, Ian Snyder, and David R Koes. Three-dimensional convolutional neural networks and a cross- docked data set for structure-based drug design.Journal of chemical information and modeling, 60(9):4200–4215, 2020
work page 2020
-
[18]
Raj Ghugare, Santiago Miret, Adriana Hugessen, Mariano Phielipp, and Glen Berseth. Search- ing for high-value molecules using reinforcement learning and transformers.arXiv preprint arXiv:2310.02902, 2023
-
[19]
Rafael Gómez-Bombarelli, Jennifer N Wei, David Duvenaud, José Miguel Hernández-Lobato, Benjamín Sánchez-Lengeling, Dennis Sheberla, Jorge Aguilera-Iparraguirre, Timothy D Hirzel, Ryan P Adams, and Alán Aspuru-Guzik. Automatic chemical design using a data-driven continuous representation of molecules.ACS central science, 4(2):268–276, 2018
work page 2018
-
[20]
Shansan Gong, Ruixiang Zhang, Huangjie Zheng, Jiatao Gu, Navdeep Jaitly, Lingpeng Kong, and Yizhe Zhang. Diffucoder: Understanding and improving masked diffusion models for code generation.arXiv preprint arXiv:2506.20639, 2025
-
[21]
Ryan-Rhys Griffiths and José Miguel Hernández-Lobato. Constrained bayesian optimization for automatic chemical design using variational autoencoders.Chemical science, 11(2):577–586, 2020
work page 2020
-
[22]
Utilizing reinforcement learning for de novo drug design.Machine Learning, 113(7):4811–4843, 2024
Hampus Gummesson Svensson, Christian Tyrchan, Ola Engkvist, and Morteza Haghir Chehreghani. Utilizing reinforcement learning for de novo drug design.Machine Learning, 113(7):4811–4843, 2024
work page 2024
-
[23]
Max Hebditch, M Alejandro Carballo-Amador, Spyros Charonis, Robin Curtis, and Jim War- wicker. Protein–sol: a web tool for predicting protein solubility from sequence.Bioinformatics, 33(19):3098–3100, 2017
work page 2017
-
[24]
Learning inverse folding from millions of predicted structures
Chloe Hsu, Robert Verkuil, Jason Liu, Zeming Lin, Brian Hie, Tom Sercu, Adam Lerer, and Alexander Rives. Learning inverse folding from millions of predicted structures. InInternational conference on machine learning, pages 8946–8970. PMLR, 2022
work page 2022
-
[25]
Xiuyuan Hu, Guoqing Liu, Quanming Yao, Yang Zhao, and Hao Zhang. Hamiltonian diver- sity: effectively measuring molecular diversity by shortest hamiltonian circuits.Journal of Cheminformatics, 16(1):94, 2024
work page 2024
-
[26]
Hyosoon Jang, Yunhui Jang, Jaehyung Kim, and Sungsoo Ahn. Can llms generate diverse molecules? towards alignment with structural diversity.arXiv preprint arXiv:2410.03138, 2024
-
[27]
Multi-objective molecule generation using interpretable substructures
Wengong Jin, Regina Barzilay, and Tommi Jaakkola. Multi-objective molecule generation using interpretable substructures. InInternational conference on machine learning, pages 4849–4859. PMLR, 2020
work page 2020
-
[28]
Alexia Jolicoeur-Martineau, Aristide Baratin, Kisoo Kwon, Boris Knyazev, and Yan Zhang. Any-property-conditional molecule generation with self-criticism using spanning trees.arXiv preprint arXiv:2407.09357, 2024
-
[29]
Panagiotis-Christos Kotsias, Josep Arús-Pous, Hongming Chen, Ola Engkvist, Christian Tyr- chan, and Esben Jannik Bjerrum. Direct steering of de novo molecular generation with descriptor conditional recurrent neural networks.Nature Machine Intelligence, 2(5):254–265, 2020
work page 2020
-
[30]
Seul Lee, Karsten Kreis, Srimukh Prasad Veccham, Meng Liu, Danny Reidenbach, Yuxing Peng, Saee Paliwal, Weili Nie, and Arash Vahdat. Genmol: A drug discovery generalist with discrete diffusion.arXiv preprint arXiv:2501.06158, 2025
-
[31]
CAGenMol: Condition-Aware Diffusion Language Model for Goal-Directed Molecular Generation
Yanting Li, Zhuoyang Jiang, Enyan Dai, Lei Wang, Wen-Cai Ye, and Li Liu. Cagenmol: Condition-aware diffusion language model for goal-directed molecular generation.arXiv preprint arXiv:2604.11483, 2026
work page internal anchor Pith review Pith/arXiv arXiv 2026
-
[32]
Jaechang Lim, Seongok Ryu, Jin Woo Kim, and Woo Youn Kim. Molecular generative model based on conditional variational autoencoder for de novo molecular design.Journal of cheminformatics, 10(1):31, 2018. 12
work page 2018
-
[33]
Zeming Lin, Halil Akin, Roshan Rao, Brian Hie, Zhongkai Zhu, Wenting Lu, Nikita Smetanin, Robert Verkuil, Ori Kabeli, Yaniv Shmueli, et al. Evolutionary-scale prediction of atomic-level protein structure with a language model.Science, 379(6637):1123–1130, 2023
work page 2023
-
[34]
Xuhan Liu, Kai Ye, Herman WT Van Vlijmen, Adriaan P IJzerman, and Gerard JP Van Westen. An exploration strategy improves the diversity of de novo ligands using deep reinforcement learning: a case for the adenosine a2a receptor.Journal of cheminformatics, 11(1):35, 2019
work page 2019
-
[35]
Xuhan Liu, Kai Ye, Herman WT van Vlijmen, Adriaan P IJzerman, and Gerard JP van Westen. Drugex v3: scaffold-constrained drug design with graph transformer-based reinforcement learning.Journal of Cheminformatics, 15(1):24, 2023
work page 2023
-
[36]
Reinvent 4: Modern ai–driven generative molecule design.Journal of Cheminformatics, 16(1):20, 2024
Hannes H Loeffler, Jiazhen He, Alessandro Tibo, Jon Paul Janet, Alexey V oronov, Lewis H Mervin, and Ola Engkvist. Reinvent 4: Modern ai–driven generative molecule design.Journal of Cheminformatics, 16(1):20, 2024
work page 2024
-
[37]
Progen2: exploring the boundaries of protein language models.Cell systems, 14(11):968–978, 2023
Erik Nijkamp, Jeffrey A Ruffolo, Eli N Weinstein, Nikhil Naik, and Ali Madani. Progen2: exploring the boundaries of protein language models.Cell systems, 14(11):968–978, 2023
work page 2023
-
[38]
Gotta be safe: a new framework for molecular design.Digital Discovery, 3(4):796–804, 2024
Emmanuel Noutahi, Cristian Gabellini, Michael Craig, Jonathan SC Lim, and Prudencio Tossou. Gotta be safe: a new framework for molecular design.Digital Discovery, 3(4):796–804, 2024
work page 2024
-
[39]
Marcus Olivecrona, Thomas Blaschke, Ola Engkvist, and Hongming Chen. Molecular de-novo design through deep reinforcement learning.Journal of cheminformatics, 9(1):48, 2017
work page 2017
-
[40]
Jinyeong Park, Jaegyoon Ahn, Jonghwan Choi, and Jibum Kim. Mol-air: Molecular reinforce- ment learning with adaptive intrinsic rewards for goal-directed molecular generation.Journal of Chemical Information and Modeling, 65(5):2283–2296, 2025
work page 2025
-
[41]
Ryan Park, Darren J Hsu, C Brian Roland, Maria Korshunova, Chen Tessler, Shie Mannor, Olivia Viessmann, and Bruno Trentini. Improving inverse folding for peptide design with diversity-regularized direct preference optimization.arXiv preprint arXiv:2410.19471, 2024
work page internal anchor Pith review arXiv 2024
-
[42]
Tiago Pereira, Maryam Abbasi, Bernardete Ribeiro, and Joel P Arrais. Diversity oriented deep reinforcement learning for targeted molecule generation.Journal of cheminformatics, 13(1):21, 2021
work page 2021
-
[43]
Deep reinforcement learning for de novo drug design.Science advances, 4(7):eaap7885, 2018
Mariya Popova, Olexandr Isayev, and Alexander Tropsha. Deep reinforcement learning for de novo drug design.Science advances, 4(7):eaap7885, 2018
work page 2018
-
[44]
Chiara Rodella, Symela Lazaridi, and Thomas Lemmin. Temberture: advancing protein ther- mostability prediction with deep learning and attention mechanisms.Bioinformatics Advances, 4(1):vbae103, 2024
work page 2024
-
[45]
Silvr: guided diffusion for molecule generation
Nicholas T Runcie and Antonia SJS Mey. Silvr: guided diffusion for molecule generation. Journal of chemical information and modeling, 63(19):5996–6005, 2023
work page 2023
-
[46]
DeepSeekMath: Pushing the Limits of Mathematical Reasoning in Open Language Models
Zhihong Shao, Peiyi Wang, Qihao Zhu, Runxin Xu, Junxiao Song, Xiao Bi, Haowei Zhang, Mingchuan Zhang, YK Li, Yang Wu, et al. Deepseekmath: Pushing the limits of mathematical reasoning in open language models.arXiv preprint arXiv:2402.03300, 2024
work page internal anchor Pith review Pith/arXiv arXiv 2024
-
[47]
Sequential posterior sampling with diffusion models
Tristan SW Stevens, Oisín Nolan, Jean-Luc Robert, and Ruud JG Van Sloun. Sequential posterior sampling with diffusion models. InICASSP 2025-2025 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), pages 1–5. IEEE, 2025
work page 2025
-
[48]
Diversity-aware reinforcement learning for de novo drug design.arXiv preprint arXiv:2410.10431, 2024
Hampus Gummesson Svensson, Christian Tyrchan, Ola Engkvist, and Morteza Haghir Chehreghani. Diversity-aware reinforcement learning for de novo drug design.arXiv preprint arXiv:2410.10431, 2024
-
[49]
Hampus Gummesson Svensson, Ola Engkvist, Jon Paul Janet, Christian Tyrchan, and Morteza Haghir Chehreghani. Diverse mini-batch selection in reinforcement learning for efficient chemical exploration in de novo drug design.arXiv preprint arXiv:2506.21158, 2025. 13
-
[50]
Morgan Thomas, Noel M O’Boyle, Andreas Bender, and Chris De Graaf. Augmented hill-climb increases reinforcement learning efficiency for language-based de novo molecule generation. Journal of cheminformatics, 14(1):68, 2022
work page 2022
-
[51]
Austin Tripp, Erik Daxberger, and José Miguel Hernández-Lobato. Sample-efficient optimiza- tion in the latent space of deep generative models via weighted retraining.Advances in Neural Information Processing Systems, 33:11259–11272, 2020
work page 2020
-
[52]
Oleg Trott and Arthur J Olson. Autodock vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading.Journal of computational chemistry, 31(2):455–461, 2010
work page 2010
-
[53]
Ziwen Wang, Jiajun Fan, Ruihan Guo, Thao Nguyen, Heng Ji, and Ge Liu. Pro- teinzero: Self-improving protein generation via online reinforcement learning.arXiv preprint arXiv:2506.07459, 2025
-
[54]
Talal Widatalla, Rafael Rafailov, and Brian Hie. Aligning protein generative models with experimental fitness via direct preference optimization.bioRxiv, pages 2024–05, 2024
work page 2024
-
[55]
Junhao Xiong, Ishan Gaur, Maria Lukarska, Hunter Nisonoff, Luke M Oltrogge, David F Savage, and Jennifer Listgarten. Proteinguide: On-the-fly property guidance for protein sequence generative models.arXiv preprint arXiv:2505.04823, 2025
-
[56]
Soojung Yang, Doyeong Hwang, Seul Lee, Seongok Ryu, and Sung Ju Hwang. Hit and lead discovery with explorative rl and fragment-based molecule generation.Advances in Neural Information Processing Systems, 34:7924–7936, 2021
work page 2021
-
[57]
Jiaxuan You, Bowen Liu, Zhitao Ying, Vijay Pande, and Jure Leskovec. Graph convolutional policy network for goal-directed molecular graph generation.Advances in neural information processing systems, 31, 2018
work page 2018
-
[58]
Optimization of molecules via deep reinforcement learning.Scientific reports, 9(1):10752, 2019
Zhenpeng Zhou, Steven Kearnes, Li Li, Richard N Zare, and Patrick Riley. Optimization of molecules via deep reinforcement learning.Scientific reports, 9(1):10752, 2019
work page 2019
-
[59]
Xingzheng Zhu, Zhihong Zhao, and Fei Zhu. Scaffold-driven molecular generation via rein- forced rnn with centroid distance evaluation.Expert Systems with Applications, 292:128606, 2025. 14 A Full Training Procedure of SGRPO This appendix provides the full training procedure of Supergroup Relative Policy Optimization (SGRPO), corresponding to Section 4 in ...
work page 2025
discussion (0)
Sign in with ORCID, Apple, or X to comment. Anyone can read and Pith papers without signing in.